Journal of Creation 22(3):68–76, December 2008
Browse our latest digital issue Subscribe
Evidence for the design of life: part 2—Baranomes
The major difference between the evolution and creation paradigms is that the evolutionist believes that the natural variation found in populations can explain microbe-to-man evolution via natural selection (Darwinism), while the creationist believes it cannot. This is because the evolutionary, naturalistic framework requires something creationists hold impossible: a continuous addition of novel genetic information unrelated to that already existing. In the creation paradigm neither variation nor selection is denied; what is rejected is that the two add up to explain the origin of species. In part 1, I discussed genetic redundancy and how redundant genes are not associated with genetic duplications and do not mutate faster than essential genes. These observations are sufficient to completely overturn the current evolutionary paradigm and could form the basis for a novel creationist framework to help us understand genomes, variation and speciation. In this second part, I argue and provide biological evidence that life on Earth thrived due to frontloaded baranomes—pluripotent, undifferentiated genomes with an intrinsic ability for rapid adaptation and speciation.
Where redundancy leads
The canonical view is that most variation in organisms is the result of different versions of genes (alleles) and genetic losses. The variation Mendel studied in peas and that led him to discover several basic inheritance laws, was the result of different alleles. At least, so it is taught. One of the seven traits Mendel described in peas was what he called the I locus—it referred to the colour of the seeds. In Mendel’s jargon, ‘I’ stood for dominance (yellow), whereas ‘i’ meant recessive (green). Plants carrying I had yellow seeds, plants lacking I had green seeds. Mendel shed scientific light on inheritance.

Now, 140 years after Mendel’s findings, we know how the yellow-green system works at the molecular level. The colour is determined by the stay-green gene (abbreviated: STG) that codes for a protein involved in the re-absorption of green pigments during senescence.1 The recessive trait i is the mutated form of the STG gene; an inactive variant that cannot re-absorb pigments, so the seeds keep their green colour.
Is the STG gene essential for survival? Most likely it is not. Molecular biology shows Mendel studied the effects of non-essential and redundant genes. Dominance means at least one redundant or non-essential gene is functional; recessive means both copies of redundant and non-essential genes are defunct. In Bacillus subtilis only 270 of the 4,100 genes are essential,2 and in Escherichia coli this is a meagre 303 out of a total of almost 4,300 genes.3 Genetic redundancy is present everywhere,4 and this lead me to believe that biology is quite different from what Darwinians think it is. Namely, organisms are full of genetic tools that are handy but not essential for survival, and selection cannot be involved in shaping these genes. Apparently, genomes are loaded with genetic elements that reside in the genome without selective constraints. This makes sense in the creation paradigm, because the genomes we observe today are remnants of the original genomes in the created kinds. And, apparently, they were created as pluripotent,5 undifferentiated genomes with an intrinsic ability for rapid adaptation and speciation. I have called the undifferentiated, uncommitted, multipurpose genome of a created kind a baranome.6 Baranomes explain genetic redundancy: there is no association with gene duplication, and redundant genes do not mutate faster than essential genes.4

The multiple genomes of Arabidopsis
In 2007, Science reported on the genome of Arabidopsis thaliana, a flowering plant of the mustard family with a small genome that is suitable for extensive genetic studies.7 This report was of particular interest because it showed the genomes of 19 individual plants collected from 19 different stands, ranging from sub-arctic regions to the tropics. According to a commentary summarizing the results of this painstaking analysis
‘ … about four percent of the reference genome either looks very different in the wild varieties, or cannot be found at all. Almost every tenth gene was so defective that it could not fulfill its normal function anymore!
‘Results such as these raise fundamental questions. For one, they qualify the value of the model genomes sequenced so far. “There isn’t such a thing as the genome of a species,” says Weigel. He adds “The insight that the DNA sequence of a single individual is by far not sufficient to understand the genetic potential of a species also fuels current efforts in human genetics.”
‘Still, it is surprising that Arabidopsis has such a plastic genome. In contrast to the genome of humans or many crop plants such as corn, that of Arabidopsis is very much streamlined, and its size is less than a twentieth of that of humans or corn—even though it has about the same number of genes. In contrast to these other genomes, there are few repeats or seemingly irrelevant filler sequences. “That even in a minimal genome every tenth gene is dispensable has been a great surprise”, admits Weigel [emphases added].’8
Among the 19 stands of Arabidopsis we find dramatic genetic differences. We observe genetic losses as well as genetic novelties. Although the dispensability of genes is easy to understand with respect to genetic redundancy, the observed novelties are much harder to conceive unless we accept that all observed novelties are not novelties at all but genetic tools that have resided in the genome since the day Arabidopsis was created. The genetic ‘novelties’ may simply reflect environmental constraints that have helped preserve these genetic tools.
There is indeed ‘no such thing as the genome of a species’, because what we observe today are rearranged and adapted genomes that were all derived from an original genome that contained all genetic tools we find scattered throughout the population. The ‘great surprise’ is only a great surprise with respect to the Darwinian paradigm. With a pluripotent Arabidopsis genome in mind, the data are not surprising at all. It is in accord with what we might expect from the perspective of a rapid (re)population of the earth.
Modern Arabidopsis genomes look as if they were derived from much larger genomes containing an excess of genetic elements—both coding and non-coding (repetitive) sequences—that can easily be lost, shuffled or duplicated. The ‘dispensable genes’ outlined above can be understood as genetic redundancies originally present in the baranome that over time slowly but steadily fell apart in the 19 individuals, because the environment did not select for them. The study strongly suggests that isolated stands of plants originated as a result of loss of genetic redundancies, duplication and rearrangement of genetic elements. The dispensability of 10% of the genes of Arabidopsis could have been predicted because most of the genes still present in individual genomes are redundant.9 In my opinion, these observations strongly favour the baranome hypothesis.
The law of natural preservation
Genetic redundancy, dispensable genes and disintegrating genomes are scientific novelties revealed to us by modern biology. How can we understand all this? Darwinians hypothesize that life evolved from simple unicellular microbes to multicellular ones via a gradual build-up of biological information, the driving force supposedly being natural selection. According to biology, there is no gradual accumulation of information; biology originated from a ‘big bang’. Sponges, worms, plants and man all have approximately the same genetic content, so the number of genes does not seem to be related to the complexity of organisms.10 In addition, the complex organisms we observe today were not derived from a single or a few simple organisms, but must have derived from a global community of organisms.11 The observations of modern biology pose so many untenable hurdles for naturalistic philosophy that it would be better to simply leave Darwinism for what it is: a set of falsified 19th century hypotheses that do not and cannot explain the origin of species.
The way to understand variation and speciation is through disintegration and rearrangement of primordial baranomes created with an excess of genetic elements. Baranomes initially contained all mechanisms required to quickly respond and adapt to changing environments. They provided organisms with the tools needed to invade many distinct niches, and were ideal for the swift colonization of all corners of the world. Baranomes were multifunctional genomes which can be compared to a Swiss army knife. A Swiss knife contains many functions which are not immediately necessary in a particular environment; but some of them are extremely handy in the mountains, others in the woods, still others are made for opening bottles and cans, or tools for making a fire. Depending on where you are, you may require different sets of functions. Similarly, depending on where the organism lives, it demands different functions (i.e. protein-coding genes and their protein products) from its genome. The environment then determines what part of the non-essential genome is under constraint and it is only this part that will be conserved. In other words, the law of natural preservation—conventionally coined ‘natural selection’—determines the differentiation of the pluripotent genome.12

C3 and C4 plants
From the creation paradigm, we might expect to find more than one carbon-fixation system. For example a system that functions optimally in warm, tropical regions, also systems that operate at sub-arctic temperatures. We find that plants do indeed have two photosystems for fixing carbon fixation; they are known as C3 and C4. The optimum temperature for carbon fixation in C3 plants is between 15–20ºC, whereas the C4 plants have an optimum around 30–40ºC.13 Today many plants are either C3 plants or C4 plants, but we also find plants that have both C3 and C4. There is clear indication for redundancy of the two photo-systems, because many plants have only one of both systems operable; either C3 or C4, plus remnants of the other system.
For instance, in modern Yellow tops (Flaveria spp) we not only see functional C3, C4 or the combination of C3 plus C4 photo-systems, but we also observe C4 remnant in C3 plants.14 The presence of remnants of one of the systems qualifies as evidence for a baranome containing both photo-systems, and indicates that the C4 system is not stringently preserved when the C3 system is also present (figure 1). The two ‘frontloaded’ photosystems ensure a rapid colonization of both high and low altitudes, and hot and cold environments. In the tropics, the C4 system, which functions optimally at high temperatures, should be active, whereas the C3 system is redundant. Here, the ‘hot’ system would be under permanent environmental constraint and be conserved. Due to accumulation of debilitating mutations in the genetic elements comprising cold systems, these would rapidly disintegrate. A genetic program designed for tropical regions does not make sense in arctic regions, and vice versa. It is the organism’s environment or habitat that determines whether genetic elements are useful or not. Conforming to the baranome hypothesis, the habitat determines genetic redundancy. There is no biological reason to assume why unused, habitat-induced redundancies should be preserved. The law of natural preservation tells us that unused genes will rapidly degrade.
What baranomes contain
Baranomes are information carriers. They were frontloaded with three classes of DNA elements: essential, non-essential and redundant. When essential elements mutate to change the amino acid sequences, the information carrier as a whole is immediately subject to a severe reproductive disadvantage. In the worst case the mutation is incompatible with life and mutated essential DNA elements will not be present in the gene pool of the next generation. Essential DNA elements can be defined as biological information that is unable to evolve. Non-essential genes are genes that are allowed to mutate and may thus contribute to allelic variations. As they produce non-lethal phenotypes, they contribute to the variation observed in populations. Classic Mendelian genetics is largely due to variation in non-essential genes. Variation in non-essential genes is what geneticists call alleles. Recessive Mendelian traits can usually be attributed to dysfunctional non-essential genetic elements; in particular elements that determine expression of morphogenesis programs, including those that determine length and shape—‘the morphometry’—of the organism. To induce the recessive trait the disrupted (or inactivated) alleles must be inherited from both parents, because an active wild-type gene usually compensates for an inactivated gene. In Mendel’s jargon, this compensation is known as dominance.
The third class of frontloaded genetic elements are correct genes that underlie genetic redundancy. They make up a special class of non-essential genes and have only recently been discovered. That is because their existence cannot be deduced from genetic experiments: they do not contribute to a detectable phenotype. Their peculiarity is that redundant genes may be completely lost from the genome without any effect on reproductive success. That redundant genetic elements make up a major part of the genome of all organisms became evident when biologists interested in gene function developed gene-knockout strategies; with the remarkable observation that many knockouts do not have a phenotype.4 Genetic redundancy is an intrinsic property of pluripotent baranomes. It should be noted, however, that the environment also plays a crucial role in determining whether a genetic algorithm is redundant, non-essential or essential. The pathway for vitamin C synthesis, for instance, is a diet-induced genetic redundancy which is inactive in humans, four primates, guinea pigs and fruit-eating bats as a result of two debilitating mutations.15
The law of natural preservation often dictates the course for the development of baranomes. In addition, baranomes initially contained variation-inducing genetic elements (VIGEs) that helped to induce rapid duplications and rearrangements of genetic information. Modern genomes of all organisms are virtually littered with VIGEs (which are usually referred to as remnants of retroviruses; LINEs, SINEs, Alus, transposons, insertion sequences, etc.) and due to their ability to duplicate and more genetic material they facilitate and induce variation in genomes.16
Speciation from baranomes
Variation in reproducing populations is mostly due to position effects of VIGEs. That is because the presence of VIGEs in or near genes determines the activity of those genes and hence their expression.17 Variation is a result of a change in gene expression. In addition, VIGEs that function as chromosome swappers may also help us understand reproductive barriers. A reproductive barrier between organisms is in fact another term for ‘speciation’, the formation of novel species. ‘Species’ is meant here in the sense of Ernst Mayr’s species concept, which includes intrinsic reproductive isolation.18 Indeed, the gene swapping mechanism present in primordial pluripotent genomes also allowed for intrinsic reproductive isolation. If we want to understand how chromosome-swapping VIGEs are involved in speciation, we first have to look into some details of sexual reproduction.
In all cells of sexually reproducing organisms the chromosomes are present as homologous pairs. One is inherited from the father and the other from the mother. The arrangement of homologous chromosomes allows them to easily pair up. Each parental chromosomes recognizes the other and they easily align. The alignment is necessary for the formation of gametes during meiosis, where the two sets of parent chromosomes are reduced to one set. Differences in chromosome pattern impede the pairing of chromosomes at meiosis, resulting in hybrid sterility. Chromosomal rearrangements may be one of the most common forms of reproductive isolation, allowing rapid adaptive radiation of multipurpose genomes without the need for geographic isolation or natural selection. The activity of chromosome-swapping VIGEs may thus have produced reproductive barriers and hence facilitating speciation.
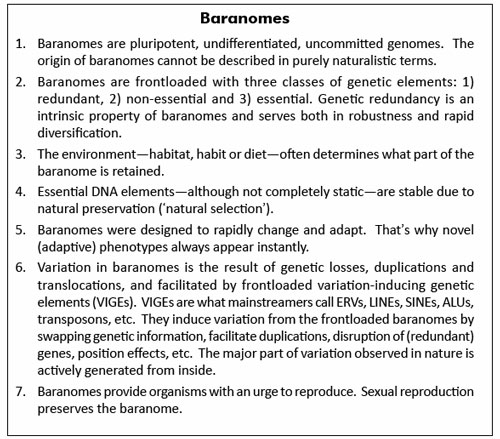
If it is true that chromosomal order determines whether organisms are able to reproduce, speciation can theoretically be reversed by chromosomal adjustments. In other words, we must be able to produce viable offspring from two reproductively isolated species just by rearranging their chromosomes. This may sound like an untestable hypothesis, but experimental evidence demonstrates that it is indeed possible to ‘unspeciate’ distinct, reproductively isolated, species by chromosomal adjustments. Using Mayr’s species definition, yeasts of the genus Saccharomyces comprise six well-defined species, including the well known baker’s yeast.19 The Saccharomyces species will readily mate with one another, indicating that they stem from one single baranome (figure 2), but pairings between distinct species produce sterile hybrids. Three of the six species are characterized by a specific genome rearrangement, known as reciprocal chromosomal translocations. Reciprocal chromosome translocation occurs when the arms of two distinct chromosomes are exchanged. Analysis of the six species revealed that translocations between the chromosomes do not correlate with the group’s sequence-based phylogeny, a finding that has been interpreted as ‘translocations do not drive the process of speciation’. However, a study carried out by the Institute of Food Research in Norwich, United Kingdom, showed that the chromosomal rearrangements in Saccharomyces do indeed induce reproductive isolation between these organisms.19 The reported experiments were designed to engineer the genome of Saccharomyces cerevisiae (baker’s yeast) so as to make it collinear with that of Saccharomyces mikatae, which normally differs from baker’s yeast by one or two translocations. The results showed that the constructed strains with imposed genomic collinearity allow the generation of hybrids that produced a large proportion of spores that were viable. Viable spores were also obtained in crosses between wild-type baker’s yeast and the naturally collinear species Saccharomyces paradoxus, but not in crosses between species with non-collinear chromosomes.
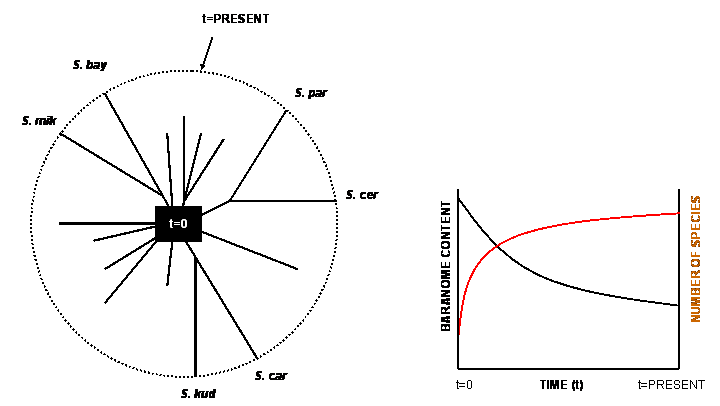
This is empirical proof that a reproductive barrier between species can be reversed just by reconfiguration of their chromosomes. In addition to reciprocal chromosomal translocations, many small-scale genomic rearrangements, involving the amplification and transposition of VIGEs may cause reproductive isolation. VIGEs are thus basic to understanding variation and speciation of baranomes. Modern biology demonstrates that although the six species of Saccharomyces yeasts are all derived from one single baranome, their individual karyotypes20 determine whether they can interbreed and leave offspring.
That the karyotype is an important determinant of reproductive isolation is also observed in deer. Eight species of Asian deer of the genus Muntiacus inhabit an area speading from the high mountains in the Himalay to the low land forests of Laos and Cambodia. Their karyotypes differ dramatically; chromosome number varies from a low of only three pairs to a high of 23.21 The muntjack species demonstrate that individuals that differ substantially by chromosomal reorganizations of otherwise identical genetic material will invariably be sterile. The sterility of muntjack hybrids is exclusively due to the inability of the chromosomes to pair. The chromosomes of distinct species simply cannot form pairs, and formation of viable reproductive cells is impossible. The karyotype accounts for reproductive isolation, and the baranome hypothesis leaves room for speciation events through adaptive radiation.
Identification of baranomes
How do we identify whether organisms descended from one primordial multipurpose genome? Darwinians claim a continuum between genomes of distinct species and view all modern species as transition stages, so this question is not of particular interest. For micro-organisms this may well be true. Between bacteria, the exchange of biological information is common and for this purpose they possess elaborate mechanisms to facilitate the uptake of foreign DNA from the environment. Still, over 5,000 distinct bacteria have been scientifically described, indicating distinctive borders between bacterial ‘species’.22
Likewise, the biological facts in higher organisms these are distinctive borders between genomes; borders determined by reproductive barriers. For instance, humans and chimps both have comparable genomic content, but very distinctive karyotypes, so the species cannot reproduce with each other. Therefore, the question raised above is not easy to answer. Because genomes tend to continuously lose unused genetic information over time, genomic content may not be suitable to identify common descent from the same primordial baranome.
A first indication that two distinct species have descended from the same baranome is this ability to mate. The offspring does not have to be fertile; neither does it have to be viable at birth. Zygote formation is a significant indication that the organisms were derived from the same baranome.
The best tool for baranome identification currently available, however, may be indicator genes. Indicator genes are essential genes with a highly specific marker. In the human baranome (Homo bn) we indeed observe indicator genes, such as FOXP223 and HAR1F.24 Both genes are also present in primates, but in humans they have highly specific characteristics not found in primates, indicating that human genomes stem from a distinct baranome. Specific characteristics typify humans. A comparative analysis of indicator genes in primates is sufficient to discriminate between man and chimpanzee (Pan bn) baranomes, or whether ancient bones belong to the human baranome. Recent research shows that indicator genes may indeed be a promising tool for baranome detection. Analyses of ancient Neandertal DNA, revealed typical human FOXP2 characteristics.25 This observation is compelling evidence that both modern humans and Neandertals originate from one and the same baranome. Further research is required to develope a full range of baranome indicator genes for other organisms.
Darwin revisited
From the baranome hypothesis we can begin to understand how Africa’s Rift Valley lakes became populated with hundreds of species of Cichlids within a mere few thousand years. We can also understand the origin of dozens of (sub) species of woodpeckers, crows, finches, ducks and deer. And we begin to see how wings could develop many times over in stick insects.26 We also understand why two distinct sex systems operate in Japanese wrinkled frogs (Rana rugosa),27 and why Dictyostelium has genetic programs for both sexual and asexual reproduction.28 And we begin to see why ancient trilobites radiated so rapidly.29 The required genetic programs (Dictyostelium’s sexual reproduction program make up over 2,000 genes!) did not have to evolve step-by-step under the guidance of natural selection. Rather, these programs were a ‘dormant’ frontloaded part of the baranome and only required a ‘wake-up call’. If Darwin had the knowledge of 21st century biology, I believe his primary conclusion would be similar to what I propose: limited common descent through adaptive radiation from pluripotent and undifferentiated baranomes. The limits to common descent are determined by the elements that have been frontloaded into that baranome. Natural varieties of sexually reproducing organisms can be established by means of differential reproductive success, but reversal to wild types follows as soon as selective constraints are relieved and hybridization between previously isolated populations occurs. Hybridization is, in fact, nothing but reversal to a more original multipurpose genome; the wild type is the more stable (‘robust’) form of the baranome because it contains more redundancies. Reversal to the wild-type was a known principle in Darwin’s day, but Darwin dismissed the obvious and invented his own naturalistic biology:
‘To admit this view [of unstable created species] is, as it seems to me, to reject a real for an unreal, or at least for an unknown, cause. It makes the works of God a mere mockery and deception; I would almost as soon believe with the old and ignorant cosmogonists, that fossil shells had never lived, but had been created in stone so as to mock the shells now living on the sea-shore.’30
Darwin rejected the baranome for theological, or at best for philosophical, reasons. Why would a flexible, highly adaptable, pluripotent genome present in primordial creatures make God’s work ‘mere mockery and deception’? A pluripotent genome with an intrinsic propensity to rapidly respond to changing situations elegantly explains ‘the co-adaptations of organic beings to each other and to their physical conditions.’ Organisms that cannot adapt or, in other words, organisms that lose their ‘evolvability’ are bound to become extinct. The baranome hypothesis with frontloaded VIGEs is sufficient to explain what Darwin observed and there is no need to invoke a gradual and selection-mediated evolution from microbe-to-man, which is non-existing and not in accord with scientific observations anyway. In every generation, VIGE activity generates novel genetic contexts for pre-existing information and hence gives rise to novel variation. VIGEs were an intrinsic property of baranomes, and they are the source of variation and adaptive radiation. It must be emphasized that, because all elements that induce variation are already in the genome, there is no need for the millions of years required for Darwinian evolution.
Conclusion and perspective
The findings of modern biology show that life is quite different from that predicted by the evolutionary paradigm. Although the evolutionary paradigm assumes an increase of genetic information over time, the scientific data show that an excess of biological information is present even in the simplest life forms, and that we instead observe genetic losses. A straightforward conclusion therefore should be that life on Earth thrived due to frontloaded baranomes—pluripotent, undifferentiated genomes with an intrinsic ability for rapid adaptation and speciation. Baranomes are genomes that contained an excess of genes and variation-inducing genetic elements, and the law of natural preservation shaped individual populations of genomes according to what part of the baranome was used in a particular environment.
With so many genomes sequenced and an ever increasing knowledge of molecular biology, we will find more and more evidence to support the baranome hypothesis. We will increasingly recognize traces and hints of frontloaded information still present in the genomes of modern species. We may expect that the genomes of the descendants of the same multipurpose genome independently lost redundant genetic elements. We may expect to find impoverished genomes and also reproductively isolated populations at different latitudes to be highly distinct with respect to their genomic content. We may even be able to piece together the genomic content of the original multipurpose genome of these species simply by adding up all unique genetic elements present in the entire population.
Finally, it will be possible to detect indicator genes, such as the FOXP2, which may become genetic tools for establishing the borders between distinct baranomes. Frontloaded baranomes are an important tool to help us understand biology. I believe there is grandeur in this view of life, where the Great Omnipotent Designer chose to breathe life into a limited number of undifferentiated, uncommitted, pluripotent baranomes; and from these baranomes all of the earth was covered with an almost endless variety of the most beautiful and wonderful creatures.31 Or, as the Bible says, ‘Let the earth bring forth grass, the herb yielding seed, and the fruit tree yielding fruit after his kind, whose seed is in itself, upon the earth: and it was so.’
Karyotype rearrangements
In 1970, Neil Todd developed the karyotypic fission hypothesis (KFH)32 to correlate the physical appearance of chromosomes with the evolutionary history of mammals. Todd postulated wholesale fission of all medio-centric chromosomes. Todd’s fast-track, single event, genome rearrangement still is the most parsimonious theory to account for mammalian karyotypes and potentially explains rapid speciation events. Todd’s was rejected mainly because it postulated something opposing the dominant Darwinian paradigm.
In 1999, Robin Kolnicki revived Todd’s KFH. Although her kinetochore reproduction hypothesis33 was largely theoretical, each step had a known cellular or molecular mechanism. During DNA replication, just before meiotic synapsis and sister chromatid segregation, the formation of an extra kinetochore on all chromosomes is facilitated. The kinetochore is the organizing centre that holds the sister chromatids together during meioses and is composed mainly of repetitive DNA sequences. The freshly added kinetochores do not disrupt the distribution of chromosomes to daughter cells during meiosis because tension-sensitive checkpoints operate to prevent errors in chromosome segregation. The result is a new cell with twice the number of telocentric chromosomes.10
The duplication of the kinetochores on many chromosomes at the same time is highly unlikely in a naturalistic model, but the telocentric chromosomes of rhinoceros, rock wallaby and many other species are physical evidence that their genomes were formed instantly.
I postulate that the genomes, as we observe them today, are the result of thousands of years of rearrangements (fission, fusion and duplications) brought about by specific variation-inducing genetic elements (VIGEs). Initially, well controlled rearrangements may have been facilitated by these elements, but over time the control over regulated genome rearrangement deteriorated. VIGEs may be the genetic basis to help us understand wholesale genomic rearrangements from pluripotent baranomes.
In order to rapidly occupy novel niches, a mechanism or ability to create reproductive barriers may have been intrinsic to baranomes. The ability to adapt, including speciation-events, is merely due to neutral rearrangements of chromosomes, and the VIGEs involved may easily become inactive because of the permanent accumulation of debilitating mutations. The remnants of VIGEs can still be found in contemporary genomes; they are known as (retro)(trans)posons, LINEs, SINEs, Alu insertion sequences, etc. Some VIGEs may have started a life of their own and now jump around more or less uncontrolled.
References
- Armstead, I. et al., Cross-species identification of Mendel’s I locus, Science 315:7, 2007. Return to text.
- Kobayashi, K. et al., Essential Bacillus subtilis genes, Proc. Natl Acad. Sci. USA 100:4678–4683, 2003. Return to text.
- Baba, T., et al. Construction of Escherichia coli K12 in-frame, single gene knockout mutants: the Keio collection, Mol. Syst. Biol. 2, 2006, doi:10.1038/msb4100050; <www.nature.com/msb/journal/v2/n1/pdf/msb4100050.pdf>. Return to text.
- Terborg, P., Evidence of the design of life: part 1 Genetic redundancy, J. Creation 22(2):79–84, 2008. Return to text.
- Pluri = more, many. In the context of baranomes, pluripotent means ‘with a capacity to diversify into more than one variety’. Return to text.
- Baranome is a composite derived from baramin and genome. Return to text.
- Clark R.M. et al., Common sequence polymorphisms shaping genetic diversity in Arabidopsis thaliana, Science 317:338–342, 2007. Return to text.
- One species, many genomes, 20 July 2007, <www.eurekalert.org/pub_releases/2007-07/m-osm072007.php>. Return to text.
- Bouche, N. and Bouchez, D., Arabidopsis gene knockout: phenotypes wanted, Curr. Opin. Plant Biol. 4:111–117, 2001. Return to text.
- Koonin, E.V., The Biological Big Bang model for the major transitions in evolution, Biology Direct 2:21, 2007; <www.biology-direct.com/content/2/1/21>. Return to text.
- Whitfield, J., Born in a watery commune, Nature 427:674–676, 2004. Return to text.
- Unlike unused Swiss army knife functions, which will not wear-and-tear, unused biological function will rapidly deteriorate due to lack of ‘selective constraint’. Return to text.
- Hess, Pflanzenphysiologie, UTB für Wissenschaft, pp. 136, 195–196, 1999. Return to text.
- Kutchera, U. and Niklas K.J., Photosynthesis research on Yellowtops: Macroevolution in progress, Theory Biosci. 125:81–92, 2007. Return to text.
- Truman, R. and Terborg, P., Why the shared mutations in the Hominidae exon X GULO pseudogene are not evidence for common descent. J. Creation 21(3):118–127, 2007. Return to text.
- Variation-inducing genetic elements will be the topic of part 3 of this series of articles. Return to text.
- Biologists used to think of genes as mere segments of DNA that specify a protein (or RNA molecule), easy to recognize in the genome because they know how to read the protein building code. Although this idea is not completely wrong, the new biology shows that it is far more complex. Rather, genes are extended stretches of DNA lacking clear borders that in addition to providing the code for the functional output (protein or RNA molecule) also contain all genetic elements required to control and regulate the production of the output. If genes did not contain information to induce and terminate its own expression, that gene would be useless. A gene is a complete information-kit that tells how and when to build a protein (or RNA molecule). Return to text.
- Queiros, de K., Ernst Mayr and the modern concept of species, Proc. Natl Acad. Sci. USA 102:6600–6607, 2005. Return to text.
- Delneri, D., et al. Engineering evolution to study speciation in yeasts, Nature 422:68–72, 2003. Return to text.
- By karyotype geneticists refer to the total number and configuration of the entire set of chromosomes of an organism. Return to text.
- Levy, H.P., Schultz, R.A. and Cohen, M.M., Comparative gene mapping in the species Muntiacus muntjac, Cytogenet. Cell Genet. 61:276–281, 1992. Return to text.
- Staley, J.T., The bacterial species dilemma and the genomic-phylogentic species concept, Philos. Trans. R. Soc. Lond. B. Sci. 361:1899–1909, 2006. Return to text.
- Terborg, P. and Truman, R., The FOXP2 gene supports Neandertals being fully human, J. Creation 22(2):13–14, 2008. Return to text.
- Terborg, P. and Truman, R., The HAR1F gene: a Darwinian paradox, J. Creation 21(3):55–57, 2007. Return to text.
- Krause, J. et al., The derived FOXP2 variant of modern humans was shared with Neandertals, Curr. Biol. 17:908–912, 2007. Return to text.
- Whiting, M.F., Bradler, S. and Maxwell, T., Loss and recovery of wings in stick insects, Nature 421:264–267, 2003. Return to text.
- Uno, Y., et al., Comparative chromosome mapping of sex-linked genes and identification of sex chromosomal rearrangements in the Japanese wrinkled frog (Rana rugosa, Ranidae) with ZW and XY sex chromosome systems, Chromosome Res. 16:637–647, 2008. Return to text.
- Personal communication with Dr. L. Bosgraaf, University of Groningen, The Netherlands. Return to text.
- Hunt, G., Variation and early evolution, Science 317:459–460, 2007. Return to text.
- Darwin, C., On The Origin of Species, 6th edition, First published by John Murray, London, 1871. Return to text.
- Adapted from the final sentence in: Darwin, ref. 30. Return to text.
- Todd, N.B., Karyotypic fissioning and canid phylogeny. J. Theor. Biol. 26:445–480, 1970. Return to text.
- Kolnicki, R.L., Kinetochore reproduction in animal evolution: cell biological explanation of karyotypic fission theory, Proc. Natl Acad. Sci. USA, 97:9493–9497, 2000. Return to text.
- Cai, J. and Zhao, R. et al., De novo origination of a new protein-coding gene in Saccharomyces cerevisiae, Genetics 179:487–496, 2008. Return to text.
- Cooper, T.F., Rozen, D.E. and Lenski, R.E., Parallel changes in gene expression after 20,000 generations of evolution in Escherichia coli, Proc. Natl Acad. Sci. USA, 100:1072–1077. 2003. Return to text.




Readers’ comments
Comments are automatically closed 14 days after publication.