Journal of Creation 26(2):88–99, August 2012
Browse our latest digital issue Subscribe
An evaluation of codes more compact than the natural genetic code
Addendum
Design issues in constructing a biological variable-length genetic code

Designing a variable-length genetic code would be error-prone and likely infeasible, which is why the natural genetic code uses a block code,1 with fixed codeword lengths.
We will limit our analysis here by assuming that a non-IDC alternative to the natural genetic code would function in the simplest manner possible, the same way done in nature: through the use of adaptor molecules, like tRNAs.
Non-IDCs have several disadvantages if implemented biologically. We illustrate using a code based on two characters (0,1) and use combinations of these to create the codewords. Figure 5 shows two characters, representing monomers (molecules) and their adaptor molecules on top. A long string of codewords becomes a polymer, which is a series of monomers linked together (figure 6).
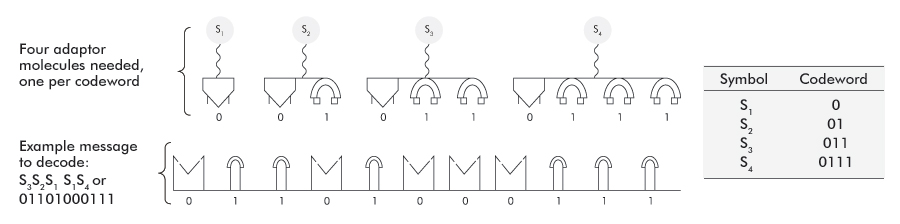
Codewords need to be identified sequentially
To be an IDC, no codeword can contain another valid codeword as an embedded prefix. A coded message is shown in figure 6. The four adaptor molecules (one for each codeword) must not be allowed to interact indiscriminately all along the message-carrying polymer (see bottom of figure 6), for several reasons:
- adaptors on both sides of an unidentified codeword would interfere with its discovery for steric reasons
- codeword positions not being processed would tie up adaptors unnecessarily. This would require a wasteful overabundance of adaptor molecules to be generated
- once an adaptor has identified its cognate codeword, the next codeword must be found. But moving the polymer forward to extract the whole message would be far more difficult and require much more energy if it had to drag along attached adaptors unnecessarily.
Therefore, the biological design should cause each codeword to be extracted sequentially (as done in nature by ribosomes).2
The decoding equipment needs an environment to facilitate identification of each codeword individually (which in our example is of variable length). In nature, adjacent regions to the codon being processed are isolated by the ribosome.
Difficulties to engineer the decoding region
Consider the message in figure 6. Should the decoding equipment focus on one, two, three, or four positions? Suppose we design for one position on the message to be exposed initially. The adaptor for codeword s1 fits perfectly, but is this the right one? The equipment needs to force open another position to find out. If the monomer representing 0 shows up next, then s1 must be right. But if instead a 1 is read, should it conclude the codeword s2 (01) was just read? The system can’t know unless another position is exposed (this is the look-ahead needed by such coding schemes). And even examining the next character is not sufficient; perhaps the correct meaning is s4, which requires creating room at the reading location for yet another check. And each time it is determined that the adaptor currently being examined is the wrong one, it must be flawlessly ejected somehow to permit another one to be tested.3
This coding scheme is a poor solution for biological purposes, leading often to translational errors. For example, if the adaptor for s3 (011) is attached, one cannot expect the system to postpone judgment as to whether s4 (0111) would be a better fit. This is exacerbated by the fact that so many codewords are identical except for a last position (0 vs 01; 01 vs 011; 011 vs 0111).
Let’s consider instead opening the reading location a maximum codeword size (four) initially each time. This would permit the adaptor for s4 to match immediately the last four positions of the message in figure 6. But this strategy permits multiple incorrect codewords to at least temporarily dock where they shouldn’t. In the case of codeword s4 (0111) all the other codewords (0; 01; and 011) could dock in there. Each adaptor molecule competes with the correct codeword4 and the solution would be highly error-prone.
A suggestion would be to produce far more of the larger adaptors, so that they would be more likely to be tested against the four-character region. We could have the adaptors generated for example in the proportion 1000:100:10:1 for codewords s4 :s3 :s2 :s1. This would still lead to far too many translation errors (c. 10% mistranslation per codon would not be tolerable for the genetic code)5 and slow down decoding significantly. Why? Because all rational designs must assign the shortest code word to the symbol most frequently used (to save energy and building material). This means that we’d have a tiny proportion of adaptors for s1 even though it would be present most often. The amino acids used most frequently should be assigned the shortest codeword, contra the solution just offered. Furthermore, it would be very costly biologically to have to generate such an overabundance of adaptors.
Without discussing all the design variants, we can identify some significant drawbacks for all non-IDC design strategies:
- the error rates would be high, with prefix codewords being often interpreted as the intended symbol
- the polymer would have to be forced forward variable distances, ranging between the size of the smallest and the largest codeword. If all energy packages (ATP) had the same value, then energy will be wasted. But creating molecules able to provide different amounts of energy would be difficult to implement and the components costly to maintain. Block codes always move the message-carrying polymer forward the same distance for each codeword.6
Alternatively, if the polymer is fed through at a constant rate, somehow the logic of rejecting embedded codewords and looking ahead would need to be designed reliably with mechanical parts.
References
- Togneri, R. and deSilva, J.S.C., Fundamentals of Information Theory and Coding Design: Discrete Mathematics and Its Applications, Chapman & Hall/CRC, Boca Raton, FL, p. 106, 2003. Return to text.
- Notice the implications for a naturalist origin of the genetic code. Even if DNA, mRNA, the adaptor tRNA molecules and the energy source from ATPs were all available, the genetic code could not work for these reasons. The highly complex ribosomes, consisting of about 100 different proteins and numerous rRNAs must be present ab initio. A ribosome provides the necessary environment to process the codons one at a time. Return to text.
- Although the characters represented by 0 and 1 in figure 5 were drawn as symmetric, they would not be designed as such. Thus, one should not conclude that the adaptor for s3 could match the coded message at positions 1–3 or 2–4 if the adaptor were to be turned 180°. This is true also of tRNA’s, the anticodon for CAT won’t match TAC. Return to text.
- Would removing the rule to limit the number of possible codewords per four character combination to one symbol improve matters? A pattern like 0000 would be immediately recognized as having at least three s1 codewords. However, this design strategy would introduce additional engineering complexities. For example, if pattern 0101 is exposed, then initially two s1 adaptors (pattern 0) could attach (and both be wrong and need to be removed); and s2 in the last two positions has only one chance in three of being correct. To know which is correct, another character position would need to be opened, perhaps by shifting the polymer to the left. If a 1 is exposed, then another position needs to be examined to distinguish between s3 and s4. Incorrect adaptors would need to be removed, and the complexity of ‘knowing’ whether other positions need to be examined would be formidable. Return to text.
- An adaptor like s4 would permit more chemical interactions with the codeword and thus be thermodynamically more stable and remain bonded longer than the simpler adaptors like s1. On the other hand, the much bulkier s4 would be far less mobile and take longer to ‘squeeze’ into the codeword location than the smaller adaptors would. For this reason we could approximate the rate in which each adaptor could be tested by roughly their relative concentrations. Return to text.
- The decoder could be designed to stop the leading monomer represented by character 0 in a specific location, so the amount of energy provided could be designed to always cover the worst case, shifting three positions, but this would be wasteful of a valuable commodity for the case when only one or two position shifts were necessary. Return to text.


Readers’ comments
Comments are automatically closed 14 days after publication.