Journal of Creation 8(1):5–6, April 1994
Browse our latest digital issue Subscribe
Antibiotic resistance in bacteria
Many people believe that bacterial resistance to antibiotics demonstrates at least a simple form of evolution. There is no doubt that bacteria exposed to antibiotics often develop the ability to live in an environment which is normally poisonous to them. This does not involve the evolution of complex structures like eyes or ears, but many still feel that it is somehow proof of the neoDarwinian fish-to-philosopher scenario.
This misconception may be partly due to the fact that even many science graduates believe that the mechanism of antibiotic resistance involves the acquisition of new DNA information by accidental mutations (copying mistakes in DNA during reproduction). But resistance does not normally arise like this.
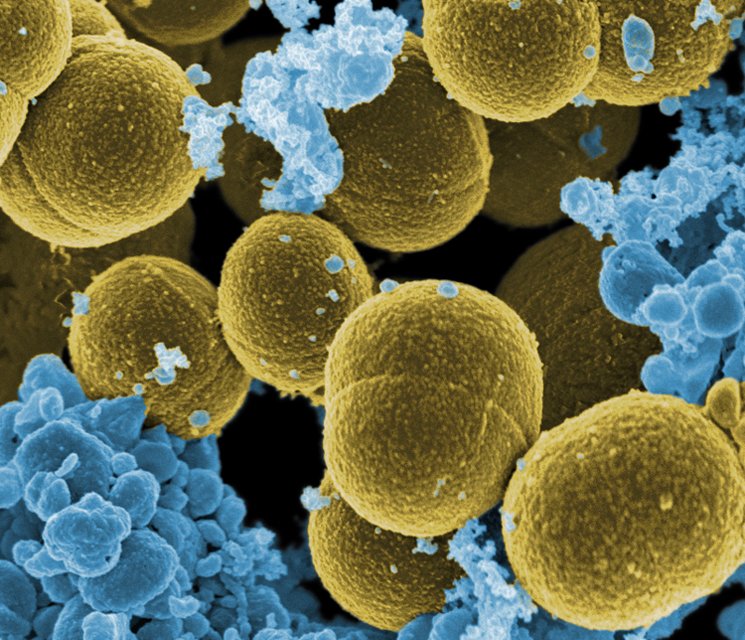
Loss of control over an enzyme’s production can engender antibiotic resistance. Take for instance penicillin resistance in Staphylococcus bacteria. This requires the bacterium to have DNA information coding for production of a complicated enzyme (penicillinase) which specifically destroys penicillin. It is extremely unlikely that such complex information could arise in a single mutational step, and in fact it does not. Mutation can cause the loss of control of its production, so much greater amounts are produced, and a bacterium producing large quantities of penicillinase will survive when placed in a solution containing penicillin, whereas those producing lesser amounts will not. The information for producing this complicated chemical was, however, already present. It took only three years for penicillin resistance to appear after the widespread introduction of penicillin.1
A mutational loss or defect can cause resistance. For instance, Mycobacterium tuberculosis, the cause of TB, has an enzyme which (as well as its other useful functions) changes the antibiotic isoniazid into a form which destroys the bacterium. A mutation causes the loss of that enzyme and helps the pathogen withstand isoniazid.2 To give another example: the 4-quinolone antibiotics attack the enzyme DNA gyrase inside various bacteria.3 An informationally insignificant mutation which results in the substitution of one amino acid by another destroys the enzyme/antibiotic interaction.
More commonly, resistance arises through mutational defects that cause the inactivation of genes which control transport through the cell membrane. If the antibiotic is less efficiently taken up, it does not accumulate as readily to toxic levels.
Antibiotic resistance commonly arises in ways which have nothing to do with mutation. For instance, in some microbes the antibacterial chemical, sulphonamide, works by blocking the ability to synthesize the vitamin folic acid. If the bacterium acquires new DNA which bypasses the block to produce this vitamin, then sulphonamide will not work as well. This pathogen is therefore resistant. The question is, where did the new DNA come from?
It is now known that bacteria can obtain such DNA (which can be outside the chromosome, as a so-called ‘plasmid’) from other bacteria which already have this information. This can happen through infection with bacterial viruses, through direct transfer from another bacterium during conjugation (‘mating’ of bacteria), or even directly through the cell wall. This acquiring of resistance from another source is clinically very important; note that it does not involve the appearance of any new, complex information which was not already present in the world.
Regardless of the form of resistance, when the population is no longer exposed to the poison, the tendency is for the more efficient, non-resistant types to do better. This is scarcely an ‘uphill’ change, of the type necessary to eventually produce complex organs and structures.
Interestingly, in most cases of antibiotic resistance, the resistant types have not somehow ‘evolved’ in response to antibiotics, because the resistant types were there before man began to use the antibiotics. We know this because 5 bacteria survive freezing, and there are cultures which have been thawed out which were frozen before many modern antibiotics were developed. Some of these bacteria are already resistant to one or more antibiotics.
In 1988, researchers did autopsies on three of the Northwest Passage explorers who froze to death in the Arctic in 1845. Bacteria from their colons were carefully cultured, and many were already resistant to the most powerful modern antibiotics.4
Evolutionists may speculate that such resistance factors arose originally by step-by-step mutations, perhaps as they encountered similar chemicals in some past environment. However, as far as observation is concerned, most cases of antibiotic resistance arising are not the result of mutation at all. The information is already there; selection acts by choosing between the various forms. This does not create any new information, so does not tell us anything at all about the evolution of increased complexity.
So-called ‘supergerms’ in hospitals are not ‘super’ at all. What has happened is that the use of antibiotics in modern hospitals has meant that the only ones surviving are those which have all the resistance factors. If a person gets a serious infection with one of these resistant types, the infection is not therefore more aggressive than if it was a non-resistant form of the same bug; it is simply that doctors are powerless to treat it. In fact, it is generally a weaker form of the pathogen.
Many patients are discharged from hospital carrying such ‘supergerms’ on their skin, which resist all attempts to get rid of them in hospital. However, when the patient gets home into a ‘dirtier’ environment, they usually rapidly disappear, because they are now forced to compete with ordinary microbes, which are ‘stronger’. These other microbes could not survive in the artificial environment of the hospital with its arsenal of antibiotics.
Such back-and-forth shifts in bacterial populations in response to the environment therefore tell us nothing about how something like a bacterium may have evolved into a more complex organism. Even though bacteria have been mutating through millions of generations (equivalent to many millions of years in more complex animals) since first discovered, the bacterial types we have today have not significantly altered since first described by Robert Koch last century.
References
- Neu, H. C., The crisis in antibiotic resistance, Science 257:1064–1073, 1992. Return to text.
- Zhang, Y, Heym, B., Allen, B., Young D. and Cole, S., The catalase-peroxidase gene and isoniazid resistance of Mycobacterium tuberculosis, Nature 358:591–593, 1992. (See also the popular report by Beardsley, T., Paradise lost? Microbes mount a comeback as drug resistance spreads, Scientific American 267(5):12–13, 1992.) Return to text.
- Lewin, C. S., Resistance to the 4-quinolones, Journal of Medical Microbiology 36:9–11, 1992. Return to text.
- McGuire, R., Eerie: human Arctic fossils yield resistant bacteria, Medical Tribune, 29 December 1988, pp. 1, 23. Return to text.






Readers’ comments
Comments are automatically closed 14 days after publication.