Journal of Creation 25(2):14–17, August 2011
Browse our latest digital issue Subscribe
Antifreeze protein evolution: turning wrenches into hammers
Evolutionists have often claimed that gene duplication provides the raw material to produce new functions through subsequent mutation and natural selection. However, finding gene duplications that have produced new functions hasn’t been easy. Most gene duplications studied have been silenced and subjected to deleterious mutations, rendering them useless.1 However, a class of proteins called antifreeze proteins (AFPs) appear to have gone against this trend. AFPs are found in a wide variety of organisms: fish, insects, plants and microbes. They also take as many different forms as there are organisms that have them, and many are believed to have evolved via gene duplication events. These proteins bind to the surface of ice crystals and prevent water molecules from binding to the ice crystals, preventing the ice crystals from growing. This enables organisms to survive in sub-zero temperatures without freezing.
However, postulating that gene duplication and subsequent mutation can result in new functional proteins is not enough. To build a plausible case for neo-Darwinism one needs to identify the source of the new gene and outline the major mutations that actually lead to the change. Researchers have recently posited a detailed evolutionary scenario for the evolution of an antifreeze protein from such a gene duplication event in a species of Antarctic eelpout (ray-finned fish), Lycodichthys dearborni.2 So do these AFPs represent a neo-Darwinian mechanism producing a new protein? And if they are, what does this say for the plausibility of neo-Darwinism forming the mechanistic basis for microbes-to-man evolution?
The edge of eelpout evolution
Lycodichthys dearborni is one of several species of Antarctic eelpout of the family Zoarcidae. It is found in McMurdo Sound, south of the Ross Sea, off the coast of Antarctica. Zoarcids are one of only two families of fish to possess a particular class of AFPs, type III. The origin of type III AFPs has been particularly difficult for evolutionists to trace.3
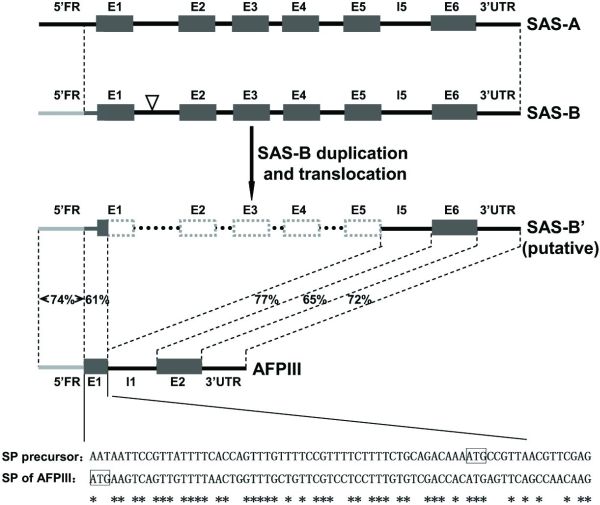
The researchers proposed that the evolutionary model ‘Escape from Adaptive Conflict’ (EAC) fits the evolution of the AFP in the L. dearborni. EAC states that conflict between an old and an emerging new function within a single gene could preserve a gene duplication, which would allow each duplicate to freely optimize one of the functions. They proposed that a type III antifreeze protein gene evolved from a duplicated copy of the sialic acid synthase (SAS) gene called LdSAS-B (Ld stands for Lycodichthys dearborni). They posit that in one duplicate, the N-terminal SAS domain was deleted and replaced with a new signal peptide (figure 1). This removed conflict between SAS and ice-binding functions in the duplicate and allowed rapid optimization of the remaining C-terminal domain to become a secreted AFP.
There appears to be one random mutation that enabled the copy of LdSAS-B to translate a protein that ended up being an effective AFP: a 4.4 kb deletion coding for the SAS N-terminal domain from codon 7 of E1 (exon 1) to the end of E5 (exon 5).4,5 This achieved two fortuitous things that transformed the putative SAS gene into an AFP gene: it stripped it of any SAS function, and enabled a change in reading frame, which altered the signal peptide, enabling the new protein to be secreted.
I suggest the large deletion mutation in the putative LdSAS-B copy produced both the increased ice-binding capabilities and enabled the putative AFP to be secreted out of the cell. The deletion was so large, it rendered the copy gene unrecognizable to the translation machinery as an SAS gene. It also destroyed the integrity of the first exon, which exposed coding for a signal peptide targeting the protein for secretion that was previously latent in the 5’ flanking region of LdSAS-B.
As Cheng et al. found:
“We discovered a precursor signal peptide coding sequence appropriately located in the extant LdSAS-B, starting from 54 nt upstream of the translation start site through the first six codons of LdSAS-B E1 (Fig. 2 and Fig. S2B). An intragenic deletion from the seventh codon of E1 through E5 of LdSAS-B and linkage of the new E1, the old I5 [intron 5], and E6 would complete the formation of the nascent two-exon AFPIII gene encoding the secretory antifreeze protein.”6
This latent coding was therefore mistakenly recognized by the translation machinery as the first exon (which had been truncated at the 3’ end due to the deletion mutation) rather than as part of the 5’ flanking region, and it became the signal peptide needed to signal the AFP (largely coded for on the second exon of AFPIII).
Strong selective pressure in the Antarctic waters would make tandem duplications of AFPIII a likely response to the conditions in order to increase the amount of AFP manufactured, and would kill off any eelpout that didn’t have the gene. Therefore, in short, the answer is yes, random mutation and natural selection is a likely mechanism for how this AFP and many others were produced.
From neo-Darwinism to evolution
However, the story doesn’t end there. The foundational question remains: is this an example of neo-Darwinism evidence for molecules-to-man evolution? There is no doubt that this is a new, functional protein; many readers might be convinced that it answers the creationist’s demand for evidence for naturalistic evolution. But it’s not that simple: to possibly stand as evidence for molecules-to-man evolution, evidence for new proteins has to have at least four characteristics:
- they need to be formed by a plausible naturalistic mechanism,
- they need to be complex,
- they need to be specified, and
- they need to be functionally integrated into the organism’s biochemical processes.
AFPs generally fill criterion 1: there is good evidence that many, if not most, are formed through random mutation and natural selection. Moreover, this particular naturalistic scenario for the Antarctic eelpout AFP is plausible. One may wonder whether designed mechanisms for variation had some role to play, especially given the large number of tandem repeats and the retrotransposon attached to the 5’ end of the gene cluster. The origin of such designed adaptability is hard for evolution to explain, but it is speculative at best for this particular scenario.
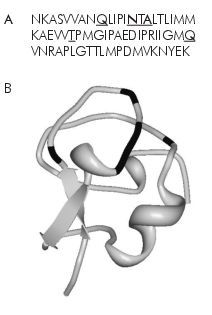
Not all AFPs fulfil criterion two: many are very simple, yet very ordered.7 The Antarctic eelpout’s AFP is only 65 peptides long, but is not arranged in a repetitious sequence.8 Moreover, many short sentences that carry a lot of information are less than 65 characters long. Consider how the original Greek text of John 1:1 would’ve been written:
ΕΝΑΡΧΗΙΗΝΟΛΟΓΟΣΚΑΙΟΛΟΓΟΣΗΝΠΡΟΣΤΟΝΘΕΟΝΚΑΙΘΕΟΣΗΝΟΛΟΓΟΣ
That sentence is only 52 characters long (51 if the subscript iota is left out), and there is much more repetition of characters there than in the AFP. Complexity is therefore not merely a measure of sequence length, but also largely of the lack of repetition in the sequence. So while short, it is of sufficient complexity to satisfy criterion 2.
Criterion three, specificity, is where AFPs in general run into major problems. As stated previously, AFPs are notorious for having as many forms as the creatures that possess them. Consider a hammer and a wrench. Each is designed for a different function: a hammer can’t do all that a wrench can, but many things can be used to do the job of a hammer when a hammer isn’t available, including a wrench. However, everyone knows that a wrench was not designed to be a hammer, and if you try to use it as a hammer too often, it may no longer work as a wrench.
The SAS protein is like the wrench, and AFPs are like hammers. All that AFPs are required to do is bind to ice, and for that they basically need only a hydrophobic end and a hydrophilic end—i.e. they must be amphiphilic. The hydrophilic end binds to the ice crystals, while the hydrophobic end repels water molecules, stopping them from binding to the ice. Such a situation is found in many proteins with completely different structures because of the 20 universal amino acids that comprise proteins, half of which are hydrophilic and half are hydrophobic.9 Being an AFP is a very non-specific job that many different random proteins could perform.
AFPs also fail to satisfy the fourth criterion: functional integration into the cell’s biochemical processes. They are not known to interact in any other cellular process other than secretion from the cell and binding to ice crystals.10 ‘Proteins’ such as AFPs effectively slow the efficiency of other functions in the cell by cluttering it with degenerate debris.11 Under normal circumstances, any mutation that enables the translation of either degenerate DNA or non-coding DNA will merely ‘gum up the works’ of the cell and ‘distract’ the transcription and translation machinery from other tasks, slowing cellular processes down.12 But again, fortuitous combinations that were already latent in the LdSAS-B gene meant that the translation of this particular debris was beneficial for survival in a particular environment.
This example may cause some excitement for evolutionists, but it doesn’t contradict what we may expect within a biblical scenario. It is a far cry from demonstrating that mutation and natural selection are necessary and sufficient to explain the history of life. Evolutionists have failed to grasp the evidential burden they have to actually substantiate their just-so stories on how high-information-content biological structures could form naturalistically. AFPs in general, and this example in particular, fail as evidence for molecules-to-man evolution.
Conclusion
AFPs are demonstrably simplistic proteins in their function, more akin to beneficial debris than a new complex and specified protein. Creating an antifreeze protein naturalistically is qualitatively different from creating, for example, the blood-clotting cascade, cellular differentiation programs, the photosynthetic pathway, or a bacterial flagellum. The difference is between incidental (and accidental) function and essential biological structure.
References
- Liu, Y. and Moran, D., Do new functions arise by gene duplication? J. Creation 20(2):82–89, 2006. Return to text.
- Deng, C., Cheng, C.-H.C., Yea, H., Heb, X. and Chen, L., Evolution of an antifreeze protein by neofunctionalization under escape from adaptive conflict, PNAS 107(50):21593–21598, 14 December 2010. Return to text.
- Cheng, C.-H.C., Freezing avoidance in polar fishes; in: Gerday, C. (ed.), Encyclopedia of Life Support Systems (EOLSS)—Theme 6.73 Extremophiles, Developed under the Auspices of the UNESCO, Eolss Publishers, UK, 2003; www.eolss.net. Return to text.
- A second mutation may have also been necessary to achieve a functional AFP. See Deng et al., ref. 2, supporting information, p. 1; Figure S1. A 6-nt deletion near the end of an early AFPIII may have somehow enabled what was a previously silenced duplication to be translated again, becoming a functional AFP. This deletion may have been necessary because the 5’ pseudogene ψAFPIII possesses a 6-nt sequence in its E2 that is present in E6 of LdSAS-B, but not present in E2 of any coding AFPIII gene. This suggests that ψAFPIII is ancestral to AFPIII. However, near the end of the protein-coding sequence of E2 seems a strange place for a mutation that enables a sequence to be translated again. Moreover, the ψAFPIII precursor has been functional, and may simply have become pseudogenized after a tandem repeat was created, and the repeat (an early AFPIII gene) underwent the 6-nt deletion which created a better AFP than the ψAFPIII precursor. The repeat was itself then tandemly repeated. Return to text.
- Deng et al., ref. 2, supporting information, p. 1; Figure S2. Return to text.
- Deng et al., ref. 2, p. 21594. Return to text.
- Davies, P.L. and Hew, C.L., Biochemistry of fish antifreeze proteins, FASEB J. 4:2460–2468, 1990. Return to text.
- Deng et al., ref. 2, supporting information, p. 2; Figure S6. Return to text.
- Williams, A., Florid forensic fable: a review of The Making of the Fittest: DNA and the Ultimate Forensic Record of Evolution by Sean B. Carroll, J. Creation 21(3):27–31, 2007. Return to text.
- Behe, M., The Edge of Evolution, Free Press, New York, p. 81, 2007. Return to text.
- Behe, ref. 10, pp. 80–81. Return to text.
- Williams, A., Mutations: evolution’s engine becomes evolution’s end! J. Creation 22(2):60–66, 2008. Return to text.





Readers’ comments
Comments are automatically closed 14 days after publication.