Journal of Creation 22(3):18–20, December 2008
Browse our latest digital issue Subscribe
The immunoglobulin heavy chain gene family and gene duplication
The immunoglobulin heavy chain gene family is an example of a gene family with irreducibly complex regulation. Therefore the family could not have been produced by past gene duplication followed by diversification. However, the tandem array of homologous genes within the locus predisposes the chromosomal region to unequal crossing-over and polymerase slip, causing copy number polymorphism in the human population. Duplications early in history may account for some of the paralogues fixed in the human race, but functional complementation between paralogues and deliberate regulation mechanisms both point to intelligent design.
Gene duplication is a specific type of mutation that increases the size of an organism’s genome. Conversely, most genes of ‘advanced’ organisms have nonallelic homologues (paralogues) within the same genome, forming gene families. The degree of sequence homology and functional similarity between paralogous genes vary from family to family. This paper focuses on distinguishing between historical duplication and common design at the time of creation. This review examines the immunoglobulin heavy chain gene family to illustrate which members of this family may be products of past gene duplication, and which are evidently not.
A family with irreducibly complex regulation cannot be produced by gene duplication
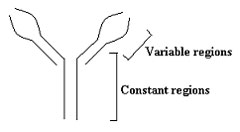
Immunoglobulins, also called antibodies, are proteins consisting of light chains and heavy chains, each having a variable region and a constant region (figure 1). They are classified according to the constant region of the heavy chains. In humans, there are five types (isotypes) of immunoglobulins (IgA, IgG, IgD, IgE, and IgM) corresponding to the five types of heavy chains (α, γ, δ, ε and μ). Each antibody molecule is encoded by a tandem array of repetitive elements on chromosome 14 (figure 2a, top). The highly repetitive V, D and J genes encode the variable region.1 One each of the V, D and J genes joins one of the downstream constant region genes (designated in figure 2 by the lowercase Greek letters). IgM and IgD—whose heavy chains are encoded by the μ and δ genes, respectively—are expressed on the surface of naïve B cells before any immune response. An immune response first causes the µ gene to be expressed as secreted IgM. Later the µ gene is skipped and one of the downstream g, e or α genes are expressed through a process called class switch recombination. The process occurs on the DNA level in antibody-producing cells (B lymphocytes).
Class switch recombination requires transcription and is accomplished with a unique set of cis-acting elements and trans-acting factors. In human and mammals, it requires the simultaneous presence of multiple enhancers (E elements) within the locus of the family, as well as an immunoglobulin (I) promoter and a switch (S) region upstream of each heavy chain gene (figure 2a, top). Vertebrates lacking the S regions (e.g. catfish) do not undergo class switch recombination, even though they may possess linked heavy chain genes.2 Deletion of one of these cis-elements in mice resulted in impaired class switch or no class switch.3-5 The organization of the I promoters, the S regions, and the structural genes suggests that a unit of I promoter S region structural gene was duplicated multiple times to produce the heavy chain gene family. However, the irreducibly complex nature of the family precludes its origination as a single unit. It requires at least two I-S-s units to achieve class switch recombination (figure 2a and b). Lacking a function, the cis-regulatory elements would have degenerated before gene duplication could ever occur.
Of the trans-acting factors involved in class switch, the key enzyme, activation-induced deaminase (AID), is B-cell specific and, therefore, serves only to produce functional antibodies. Although it is also used in another aspect of antibody synthesis (somatic hypermutation of V regions), class switch requires a functional domain of the enzyme that is not required for hypermutation.6,7 There exists strong evidence that other class switch-specific enzymes/cofactors are required to guide μ—α, μ—γ and μ—ε switches to achieve the appropriate immune response.7,8 Therefore multiple genes must be simultaneously added onto the genome outside of the heavy chain gene family for class switch recombination to occur, further underlining the irreducible complexity of the system.
Class switch also requires multiple enzymes involved in DNA rearrangement, excision repair and nonhomologous end-joining (NHEJ) pathways (figure 2 legend). Although these enzymes are not class switch-specific, and therefore could theoretically have evolved independently, their expression in B cells and coordination with the class switch-specific factors necessitates careful design. For example, the excision repair enzymes, such as uracil DNA glycosylase (UNG), are activated in B lymphocytes to cause mutations instead of repairing them, both during class switch recombination and during somatic hypermutation.
Moreover, development of a functional class switch mechanism must occur concomitantly with structural and functional diversification of the various immunoglobulin isotypes. The five human immunoglobulin isotypes have different structures, tissue distributions and functions. Switching without diversification is futile, and diversification without switching would fail to express the downstream genes—another aspect of irreducible complexity.

All components of an irreducibly complex system must be created with their designated functions simultaneously. However, Darwinian mechanisms require gene duplication and subsequent diversification, with regulatory elements developed either before or afterwards. Gene families such as the immunoglobulin heavy chain genes are not products of gene duplication because:
- the system could never have functioned as a single-copy gene;
- elements that regulate the entire family (such as the enhancers of the heavy chain locus) do not exist in ‘lower’ organisms;2
- trans-acting factors and family-controlling cis-acting elements cannot be produced by duplication of a structural gene with its linked regulatory elements;
- degeneration of duplicates happens faster and with more certainty than meaningful diversification.9
Copy number polymorphism results from duplication or deletion
Homologous gene clustering predisposes a chromosomal region to unequal crossover or polymerase slip, causing variation in copy numbers within the gene family. Due to the highly repetitive nature of the immunoglobulin heavy chain genes, this locus displays considerable copy number polymorphism among the human population. For example, duplication of the heavy chain genes is 22% in Mongoloids, 10% in Caucasians and 5% in Negroids.10 Most of the duplications encompass multiple genes, such as ψε-γ2, γ1-γ4, α1-ε, etc., and some triplications exist, such as of α1-ε among the Japanese. Deletions, mostly of γ4 or γ1, are much rarer (1.5–3.5% depending on race). Duplications cause no or slight elevation in serum immunoglobulin levels, while deletions cause more significant lowering of the corresponding immunoglobulin levels.11 The lower frequency of deletions in the population is presumably due to negative selection, while the evolutionary significance of duplications is unclear, but presumably not as significant as are deletions.
Beneficial gene repetitions may be produced either by duplication or by design
Figure 2 also demonstrates that the γ-γ-ε-α cluster is repeated once within the locus. There are functional differences between corresponding genes in the two clusters. The products of the γ2 and γ4 genes in the downstream cluster (IgG2 and IgG4) are significantly less potent in activating complement or binding Fc receptors than those of the γ1 and γ3 genes in the upstream cluster (IgG1 and IgG3). While the downstream ε gene encodes all the IgE, the upstream ε gene is a pseudogene (i.e. a gene that does not code for a functional protein).
There exist apparently healthy individuals with homozygous deletions of some heavy chain genes, especially of the genes in the downstream cluster.12,13 It is therefore possible that the two clusters were produced by duplication of one original cluster. Subsequently, the upstream ε gene pseudogenized, while the downstream γ and α genes subfunctionalized (lost parts of their functions due to degenerative mutation). The inability of IgG2 and IgG4 to activate complements offers the human body a noninflammatory type of protection, contrary to IgG1 and IgG3 which cause inflammation through complement activation. Further, IgG2 and IgG4 may have a role in regulating the proinflammatory effects of IgG1 and IgG3. This duplication/subfunctionalization is therefore beneficial to the human race. However, the duplicates could not have fixed in mankind through natural selection because the increase in fitness was conceivably small and the new duplicate would have quickly drifted out of the population.14 On the other hand, the fixation is easier to explain by the founder effect of Noah’s family if the event occurred in Noah’s lineage before the Flood, or by genetic drift if it occurred shortly after the Flood when the population size was still small.
Although a duplication/subfunctionalization scenario can be speculated for the origin of the two clusters, there always remains the possibility that they were created in Adam and Eve to produce subclasses of immunoglobulins with diverse functions. In fact, immunoglobulins encoded by the two heavy chain gene clusters are differentially regulated. While class switch to the upstream cluster is activated by a subset of immune cells called Th1 cells, class switch to the downstream cluster is activated by another subset of T lymphocytes (Th2 cell). There is increasing evidence pointing to deliberate regulation mechanisms that facilitate class switch to the downstream heavy chain genes in situations where a noninflammatory response is more desirable. For example, IgA1-producing B lymphocytes can be induced to secrete IgA2 when they are recruited from the systemic circulation to the mucosa of the colon.15
Likewise, we cannot rule out the hypothesis that the two linked γ genes within each γ-γ-ε-α cluster indicate another duplication event before duplication of the entire cluster, although all four γ genes may have been created together to encode functionally different IgG subclasses.
Conclusion
Just as multiple V, D and J genes are needed to produce the diversity required for adaptive immunity,1 the immunoglobulin heavy chain genes were designed as a family to produce various immunoglobulin isotypes for different functions. The unique regulatory elements of class switch recombination must have been created along with the structural genes. However, the tandem array of homologous DNA sequences in the locus allowed errors in crossover and/or replication, producing duplications and deletions during human history, for good or for bad.
References
- Bergman, J. and O’Sullivan, N., Did immune system antibody diversity evolve? J. Creation 22(2):92–96, 2008. Return to text.
- Stavnezer, J. and Amemiya, C.T., Evolution of isotype switching, Semin. Immunol. 16(4):257–275, 2004. Return to text.
- Pinaud, E., et al. Localization of the 3’ IgH locus elements that effect long-distance regulation of class switch recombination, Immunity 15(2):187–199, 2001. Return to text.
- Shinkura, R., et al. The influence of transcriptional orientation on endogenous switch region function, Nat. Immunol. 4(5):435–441, 2003. Return to text.
- Zhang, J., et al. A selective defect in IgG2b switching as a result of targeted mutation of the I gamma 2b promoter and exon, EMBO J. 12(9):3529–3537, 1993. Return to text.
- Barreto, V., et al. C-terminal deletion of AID uncouples class switch recombination from somatic hypermutation and gene conversion, Mol. Cell 12(2):501–508, 2003. Return to text.
- Ta, V.T., et al, T., AID mutant analyses indicate requirement for class-switch-specific cofactors, Nat. Immunol. 4(9):843–848, 2003. Return to text.
- Ma, L., Wortis, H.H. and Kenter, A.L., Two new isotype-specific switching activities detected for Ig class switching, J. Immunol. 168(6):2835–2846, 2002. Return to text.
- Liu, Y. and Moran, D., Do new functions arise by gene duplication? J. Creation. 20(2):82–89, 2006. Return to text.
- Rabbani, H., et al. Duplications and deletions of the human IGHC locus: evolutionary implications, Immunogenetics 45(2):136–141, 1996. Return to text.
- Rabbani, H., et al, L., The influence of gene deletions and duplications within the IGHC locus on serum immunoglobulin subclass levels, Clin. Immunol. Immunopathol. 76(3 Pt 2):S214–S218, 1995. Return to text.
- Brusco, A., et al. A recurrent breakpoint in the most common deletion of the Ig heavy chain locus (del A1-GP-G2-G4-E), J. Immunol. 163(8):4392–4398, 1999. Return to text.
- Engström, P.E., et al. Salivary IgG subclasses in individuals with and without homozygous IGHG gene deletions, Immunology 89(2):178–182, 1996. Return to text.
- Sanford, J., et al. Using numerical simulation to test the validity of Neo-Darwinian theory; in: Snelling A.A. (Ed.), Proc. Sixth Inter. Conf. Creationism, pp. 165–175, 2008. Return to text.
- He, B., et al. Intestinal bacteria trigger T cell-independent immunoglobulin A(2) class switching by inducing epithelial-cell secretion of the cytokine APRIL, Immunity 26(6):812–826, 2007.Return to text.





Readers’ comments
Comments are automatically closed 14 days after publication.