Journal of Creation 30(2):76–82, August 2016
Browse our latest digital issue Subscribe
Origins of pathogenic microbes: part 1—bacteria
The emergence of pathogenic bacteria interests creationists because it impacts on concepts of God’s character. The ecosystem operated in a stable fashion at creation and the basic microbe types participated in maintaining a dynamic homeostatic state. Sin brought a change to features of DNA repair and gene regulation in living systems, and stress was introduced into the equation. This resulted in mutations and other errors giving rise to cellular malfunctions. Mechanisms permitting re-assortment and exchange of genetic material within precise limits were disrupted. Most of these mechanisms are well known, but additional ones have been described recently and more may be discovered. Reasonable hypotheses are generated to account for the emergence of the main groups of pathogenic bacteria by unguided events involving genetic material. These involve events in the microbe, in the host, and changes in the environment. Subsequent to the Fall, changes to living organisms and the ecosystem usually are not considered dependent on the direct activity of God. However, other external agencies could have been involved.

The biblical account of origins indicates that creation was very good. The origin of pathogens causing suffering, disease, and death is a challenging question. A number of possibilities have been suggested as follows:
1) microorganisms (including viruses) were not disease-causing initially. They devolved after creation as a result of mutation and gene shuffling among themselves. As a consequence a small number of pathogenic variants arose,1 2) God created pathogens in the lower orders of life to ensure population control, but this did not extend to microbes inhabiting humans. Rather, mutations gave rise to pathogens among populations of usually beneficial microbes inhabiting the human body and similar mutations in animal pathogens enabled them to jump the species barrier and infect humans,2 3) the genomes of all creatures were designed so that they could adapt rapidly to the environment. These variant-producing DNA segments have become compromised, giving rise to RNA viruses and perhaps retroviruses, ostensibly on account of the removal of the ‘regenerating healing power’ of God,3 4) diversity can be accounted for by postulating that transposable elements were originally designed to produce positive altruistic outcomes, but subsequently caused near neutral, or even deleterious, mutations,4 and 5) microbes were not initially disease-causing in creatures with pain sensations but as a consequence of various changes they appeared, perhaps assisted by exterior agencies.
Change after the Fall conceivably involved shifts in ecosystem balance so that the nature and behaviour of organisms was altered. In an extension of the suggestion about agencies, some have postulated that an evil agent(s) altered or added novel genetic information to the genome of existing organisms or an entirely new line of microbes was allowed to emerge (God’s Curse).5
To answer these proposals, several assumptions are made. Undoubtedly, some of these will be challenged as knowledge increases. The most significant is that the existence of beneficial phenomena in the natural world that are widely expressed will be taken to indicate their essential continuity from the beginning (parsimonious approach), except where biblical information dictates otherwise.
Events promoting the rise of pathogens?
The vast majority of microbes do not inflict damage on their hosts. They are non-pathogenic. The majority of microbes inhabiting humans are classically harmless commensals (one organism benefits, the other is not harmed or benefited).6 Microbes are present in large numbers in and on the human body and ensure our survival. They aid digestion and provide nutrients necessary for our well-being. In herbivores, they are absolutely vital for cellulose digestion and the provision of some of the protein needs. Microbes that cause damage are called pathogens. However, disease does not depend alone on the microbe acquiring special abilities such as toxin genes. Three factors contribute to the expression of disease—microbe, host, and environment.7
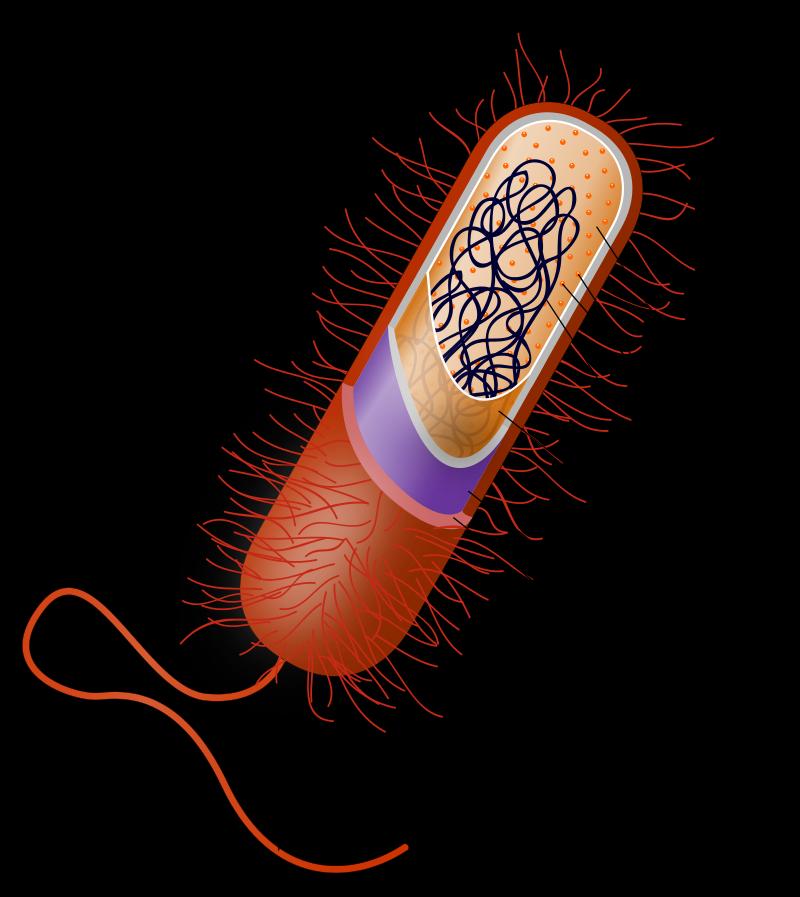
Microbes that exhibit polysaccharide capsules (figure 1), toxin- and melanin- forming features, tissue-invading (enzyme) capabilities, and other characteristics frequently are pathogenic. These may have arisen as a consequence of a variety of events. First, elements of the gene regulatory network, particularly the non-coding region of the chromosome, may have been altered. As a consequence, feedback processes, dominant metabolic pathways, and other aspects of cellular function were altered in the host or microbe.8 Second, microbes in mutualistic (organism and host benefit from association) and endosymbiotic associations (living inside another organism) may have been modified through degenerative and other changes. Third, saprophytic equivalents (free-living on dead material) to the present-day pathogens may have been created and have subsequently acquired pathogenic abilities. Fourth, genes may have been lost that enabled the expression of pathogenic ability. Fifth, changes may have occurred in the host, allowing opportunistic organisms to become pathogenic. Sixth, organisms may have adapted under environmental stress or changes in nutrition, facilitating the appearance of pathogens.9 Seventh, intelligent agents may have manipulated the genetic material.10 Finally, there may have been a combination of the above events.
In environments containing a diversity of organic materials rich in DNA, and where microbial growth and decay activities are intense, it is more likely that organisms exchange genetic information.11 Mechanisms known today, including recombination during sexual reproduction, transduction, transformation, and non-random transposition are considered to have been present from creation. Mutations were probably also known then. There are different categories of mutations.12 Those commonly encountered today arise frequently through exposure of the organism to stress (chemical, nutritional, and physical factors). DNA is damaged and the repair mechanisms are unable to rectify the faults, meaning that imprecision in nucleic acid copying occurs. The entire operational strategy set in motion at creation changed after sin’s entrance so that future outcomes were foreseen, but are not attributable to the Planner.
Genesis and expression of pathogenic capabilities
Virulence (relative ability to cause disease) is not strictly an intrinsic microbial characteristic but includes both microbial and host factors. This conclusion is reached on account of commensals and opportunists being able to cause disease in the immunocompromised. In retrospect, the factors classically related to virulence, such as toxicity, aggressiveness, replication and transmission advantages, adherence and attachment, and induction of deleterious immunologic reactions,13 still have relevance.
Changes in colonising status, such as the ability to cause disease, ultimately indicate changes in DNA, gene regulatory network elements and other features. DNA repair mechanisms are variously subject to imperfections so that variant DNA appears. As a result some viable mutants are generated. For example, bacteria may experience mutations on account of damage to their mismatch repair system. These mismatch defects may subsequently be removed, but the mutations generated during the period of high mutation are retained.14 Changes in proteins coded by the genome are expressed through the regulatory elements that involve feedback processes, dominant metabolic pathways, and other features. If the right combination of virulence factors is present in an organism (many of which may be present already in non-pathogenic organisms), then cross talk among virulence factors may result in regulatory changes leading to the actual expression of virulence.15
Changes may also occur in the non-coding region of the chromosome near the promoter (regulator changes). This would cause gene expression levels to be altered in the host or microbe. Sin’s entrance may have altered the repair mechanisms’ efficiency due to the stresses introduced and may also have involved changes in the regulatory elements.
An example involves the well-known bacterium Escherichia coli. Its effectiveness as a gut pathogen is influenced by the DNA modifications (base methylation) which affect the production of an appropriate attachment device (pilus—figure 1). This plays a role in disease development. Methylation levels (represents an epigenetic control mechanism) are governed by the growth conditions of the organism. In other bacteria, virulence is partly lost when methylation fails to occur or is overproduced.16 Theoretically, an organism could be benign, but after the removal of the silencing effects on a gene (influenced by methylation state), it could become pathogenic.17
Microbe changes
Microbes as a group can function in almost all environments and carry out a wide range of activities. Theoretically some microbes could adapt to the pathogenic mode of existence. Selected examples follow.
Mutualistic and endosymbiotic organisms. Altering mutualistic organisms to take on parasitic capabilities is one possibility, as they already possess host entry mechanisms and strategies to acquire nourishment. The rhizobia bacteria are mutualistic with leguminous plants. Special genes (bac) enable them to form intracellular structures, permitting the fixation of atmospheric nitrogen. These genes have a high degree of similarity with those found in Brucella, a well-known intracellular animal pathogen, which allows it to survive in the phagocytic cells of the animal host. In addition, Brucella’s regulatory system for virulence has remarkable similarities to certain genes found in the harmless mutualist (Sinorhizobium meliloti).18 The plant bacterium Agrobacterium, commonly considered a pathogen, could have acquired many of its capabilities from rhizobia.19 How the acquisition occurred is a separate issue.
The bacterium Wolbachia has achieved prominence on account of its apparent ability to transfer information to insect and nematode hosts. For example, the fruit fly (Drosophila ananassae) has the entire genome of the endosymbiotic bacterium inserted into one of its chromosomes.20 These symbionts influence reproductive success and strategy in insects.
Research on the Wolbachia genome has been taken by some to suggest that endosymbionts have acquired this ability through the loss of critical genes needed for growth independent of a host.21 However, introducing Wolbachia into the mosquito is connected to the development of resistance of this insect to transmitting a devastating form of malaria.22 This may indicate that endosymbionts had some protective role from creation and that their present genome imperfectly represents the original.
Saprophytes
Many saprophytic microbes are beneficial to other organisms.23 Normally harmless inhabitants of the soil can become pathogenic under certain favourable environmental conditions. Soil saprophytes also often possess capsules from which the capsule-forming ability of successful pathogens may have been derived.24 These opportunists also become pathogenic when the host defences are breached or compromised25 or when the organism is introduced into the body following prosthesis implantation and the insertion of catheters.26
Bacillus is an example of a saprophyte that has pathogenic representatives. Some members are pathogenic, such as B. anthracis, the causal agent of classical anthrax. It is pathogenic due to its capsule-forming and toxin-generating capacities. Capsule formation is conferred by transfer of extrachromosomal DNA material in plasmids and subsequent modification. Toxin genes are also plasmid borne. An acquisition route for the saprophyte has been indicated by studying a close relative of Bacillus cereus capable of causing anthrax in chimpanzees.
B. cereus appears to be a normal inhabitant of the intestines of soil-dwelling insects and earthworms and may be useful partners with other insects.27 One possibility is that the anthrax-forming B. cereus originally had a well-established role in the soil. Transformation (uptake of naked nucleic acid) or transduction (transfer of genetic information by phage), involving acquisition of plasmids (containing genes that produce toxin) from B. anthracis, apparently gave B. cereus the ability to induce anthrax-like symptoms. Acquisition can occur ostensibly in soil or during co-infection of the host.28
Mutations. Virulence may arise through mutations, which are expressed in a number of ways. Gene loss may lead to the emergence of pathogenic ability. The cadaverine gene (lysine decarboxylase activity) is able to prevent toxin production. The majority of Escherichia coli strains possess this gene, which appears to be connected with their commensal status in the human gut. When cadaverine gene activity is lost in certain strains of Escherichia coli (EIEC) they develop pathogenic ability. The loss of activity is associated with large deletions in the chromosome region around the cadaverine locus. This means that toxin production henceforth can be expressed.29 Again, in Listeria monocytogenes, mutation of a critical gene associated with a regulatory protein may lead to changes whereby the normally saprophytic organism can become constitutively pathogenic.30
Host changes
The best-known example of host changes leading to disease is the process that causes immunocompromisation. These hosts may be predisposed to attack from organisms not normally considered pathogens. Sometimes around half of such individuals experience bacterial infections. For example, the usually saprophytic bacterium Nocardia can be a significant cause of illness.31
Environmental influences facilitate emergence and expression of change
Changes in the environment may be responsible for the emergence and expression of virulence traits. Several examples will be given in each category.
Emergence. Adaptive mutations may occur in response to environmental stresses. Opportunistic pathogens in the Burkholderia cepacia complex affect cystic fibrosis and immunocompromised patients. They may adapt through exposure to antibiotics and other stress conditions. These variants are more suited to colonise the lungs of fibrosis sufferers and contribute to disease progression.32
During stress, mechanisms normally controlling transfer of information horizontally between organisms may be relaxed due to the inactivation of the methyl-directed mismatch repair system. The system can be damaged through exposure to starvation conditions, for instance. Change in this repair system increases the mutation rate substantially, causing hypermutation, and as a result greatly increases the recombination success between Escherichia and Salmonella. This may represent one mechanism responsible for the appearance of virulence genes in the food poisoning bacterium Escherichia coli (0157:H7) from another strain of the same organism.33 However, a number of other possibilities have been suggested in the genesis of the strain.34
Expression. Some microbes are not pathogens when biological balance is maintained in a natural environment. But saprophytic bacteria responding to change may damage potential hosts. For example, Acinetobacter baumanni is significant in wound situations and airway infections in alcoholics. In hospital situations where ethanol (alcohol) is used as a disinfectant, it can function as a nutrient that increases the virulence of the organism by the differential expression of certain genes. The nutrient may also influence the expression of tissue receptors, which would then initiate intracellular signalling, leading to differential gene expression and heightened tissue responses that allow increases in tissue damage by the microorganism.35 The bacterium Agrobacterium tumefaciens (commonly saprophytic in soils) also displays a similar pattern of virulence induction. The organism becomes pathogenic to plants following exposure to phenolic substances and/ or saccharides released. Sensing these products occurs at the membrane level, by sensory proteins, and ultimately to the activation of promoters. The inducing sugars may be released by wounded tissues.36

Mechanisms for transfer of genetic information
There are well-known methods for horizontal transfer of information among bacteria, including transformation (uptake of external DNA through the cell membrane), transduction (transfer by bacterial viruses or phages), and sexual reproduction (referred to as conjugation between closely related types—figure 2). Less well-known is transfer through membrane vesicles and intercellular nanotubes.
Communication of information through intercellular nanotubes may develop between the same or different species growing on solid surfaces. These tubes enable acquisition of characteristics in a non-hereditary fashion so that antibiotic resistance becomes a temporary feature of the recipient bacterium. Some plasmids (extrachromosomal DNA material) may be transferred reciprocally.37 Outer membrane vesicles produced by bacteria may also facilitate delivery of proteins and DNA and RNA between bacterial strains.38
Horizontal gene transfer can transform normally harmless bacteria into disease-causing ones. Indeed, it appears that the majority of genes for pathogenicity moving across taxa are acquired through bacterial viruses via transduction.11 These phages may stimulate toxin production, allow cell adherence to host tissues, allow host defences to be evaded, and may stimulate genes to replicate and transduce.39 The classic example of conversion of a non–disease-causing bacterium to a pathogen is Corynebacterium diphtheria.40 Acquiring a phage carrying the structural gene for exotoxin production enables it to become pathogenic.41
Genes for exotoxins can be carried by plasmids and phages. New pathogens may arise following the transduction of a toxin gene from some environmental source to a member of the normal human microbiota. For example, Staphylococcus aureus may have acquired its exotoxin gene from the environmental bacterium Pseudomonas (>95% similar) via a phage.42 Furthermore, genes responsible for pathogenic ability in Corynebacterium (dtx), Shigella (stx), and Vibrio (ctx) are present in environmental samples, and additional sources of the Staphylococcus exotoxin-like gene have been found there as well. Phages isolated from the environment (water, sediment, and soil) would have carried all but the dtx gene. Conceivably, the genes would have functioned in the environmental bacteria to maintain homeostasis. There are indications that complex interactions also can occur among phages. These have enabled wild type and defective non-pathogenic Vibrio cholera to become pathogenic.43 In the soil environment there exists a wealth of phage types with their numbers paralleling trends in bacterial populations.44 Phage-bacterium and interphage interactions occur in the natural environment. This allows the construction of a coherent hypothesis for the emergence of pathogenic organisms.
An example of information transfer from an environmental bacterium is suggested from a study of non-pathogenic Escherichia coli isolates. Some of these carry a suite of genes enabling efficient iron uptake, such as the yersiniabactin system, which confers ecological fitness. When similar genes are present in Yersinia they contribute to pathogenic ability.45 The transfer of the pathogenicity island is thought to be facilitated by a conjugative plasmid which has a wide host range.46
Other horizontal gene-transfer phenomena are known. Few credible suggestions exist to explain the gene acquisition noted among some organisms.47 There are formidable restriction enzymes and other barriers to transformation among some bacteria and cyanobacteria. Some strains may lack these barriers, thus permitting transformation (figure 3).48 In other instances, stress factors override the constraints normally experienced.49 Additional sources of the genetic information transfer include the following:

- Transfer of secretion-system capabilities among bacteria and beyond. Evidence suggests that the type III secretion proteins, widely present among plant and animal Gram negative bacteria, have been acquired through horizontal gene transfer from donor bacteria. Transfer also is postulated to occur in organisms that range from bacteria to protozoa. These secretion proteins facilitate pathogenic and mutualistic modes of existence. Many pathogenic bacteria use such a secretion system, which allows proteins to enter eukaryotic cells. This secretion system has similarities to components required for the biosynthesis of flagella and other operational features noted in mutualist rhizobial bacteria.50 These similarities suggest either transfer of capabilities among organisms or experimental manipulation by intelligent agents.
- Bacterium-to-plant transfer. The bacterium Agrobacterium tumefaciens transfers genes from a plasmid (T-plasmid) to selected dicotyledonous plants, where they integrate naturally with the plant nuclear DNA and subsequently cause tumour formation. Tumours form as a consequence of auxin and cytokine production and the synthesis of supporting nutrients (opines) by the genes transferred.51
- Plant-to-bacterium transfer. Genetic information acquired by bacteria may involve sources outside the prokaryotes. Information may come from decaying plants in the natural environment. For example, marker genes associated with chloroplast DNA present in residues of tobacco plants colonizing by the pathogenic bacterium Ralstonia solanacearum are transmissible to soil bacteria. More significantly, plant chloroplast DNA sequences were shown in 8% of bacteria growing on the residue. In the same environment Acinetobacter and Pseudomonas acquired exogenous plasmid DNA by transformation. Even without the presence of the plant pathogenic Ralstonia, Acinetobacter is able to transform on decaying tissues.52 These studies raise the possibility of significant transfer of genetic information in soil and plant environments. Such transfer of information should not surprise as chloroplast DNA has many features similar to prokaryote DNA.53
Another intriguing illustration is provided by the pathogen Xanthomonas axonopodis. It causes a disease of citrus trees characterized by the appearance of wet lesions. The bacterium contains a gene sequence that encodes a protein able to influence the capacity of the plant to take ions and water into cells. It has a high degree of similarity with a sequence in the plant that encodes a plant natriuretic peptide. This protein maintains homeostasis when the plant is subjected to osmotic stress. Evidence suggests that the gene was acquired by the bacterium through horizontal gene transfer from the plants. This allows the bacterium to induce plants to increase uptake of ions and water into cells, which favours the growth and survival of the bacterium in plant tissues.54 The acquisition of information from the plant may have been via similar mechanisms as discussed in the previous paragraph.
The basic assumption made in many research studies is that if a gene sequence shows greater similarity to those held by distantly related organisms than that by close relatives, it has the possibility of being an acquired gene.55 It may be argued in some circumstances that similarity in DNA signature strongly suggests the existence of a Designer. The complexity of inferences that can be made from data means that a number of lines of evidence need to converge before horizontal transfer can be asserted to have occurred with confidence.56
Conclusions
Several fundamental changes following the Fall are indicated in Genesis and Job. Many of the changes associated with the emergence of pathogenic bacteria from non-pathogenic ones can be accounted for by naturalistic phenomena. However, the possibility remains that intelligent agents may have manipulated the genetic material.
There is evidence for alteration in the gene regulatory network (mutations and other phenomena) in the presence of stress, particularly the non-coding region of the chromosome. As a consequence, feedback processes, dominant metabolic pathways, and other aspects of cellular function can be changed in the host or microbe. Evidence has been presented for the exchange of information among mutualists, endosymbionts, and saprophytic bacteria with the occasional help of bacterial viruses and accession of DNA from other organisms. Gene loss enabling the expression of pathogenic ability has been responsible for the emergence of some pathogens, and it also appears that changes in the host genome have allowed opportunistic organisms to become pathogenic. Environmental stressors have facilitated the appearance of pathogens. Ostensibly, the massive increases in decaying organic debris available following the Fall, with the associated release of DNA into the environment, has played a role in the changes I have sought to explain.
References and notes
- Bergman, J., Did God make pathogenic viruses? J. Creation 13(1):115–125, 1999; Francis, J.W., The organosubstrate of life: a creationist perspective of microbes and viruses, Proceedings of the Fifth International Conference on Creationism, pp. 433–444, 2003, www.answersingenesis.org/articles/aid/v4/n1/organosubstrate-of-life. Return to text.
- Rana, F., Did God create flesh-eating bacteria? A creation model for the origin of human disease, Reasons to Believe, 2010, www.reasons.org/articles/did-god-create-flesh-eating-bacteria-a-creation-model-for-the-origin-of-human-disease. Return to text.
- Borger, P., Evidence for the design of life: part 2—Baranomes, J. Creation 22(3):68–76, 2008; Borger, P., The design of life: part 4—variation-inducing genetic elements and their function, J. Creation 23(1):107–114, 2009. Return to text.
- Wood, T.C., The Ageing process: rapid post-flood intrabaraminic diversification caused by altruistic genetic elements (AGES), Origins 54:5–34, 2002. Return to text.
- Gillen, A., Microbes and the days of creation, Answers Research J. 1:7–10, 2008; Shipton, W.A., Thorns also and thistles, J. Adventist Theological Society 23(1):18–45, 2012. Return to text.
- David, R., Microbiome: pathogens and commensals fight it out, Nature Reviews Microbiology 10:445, 2012 | doi:10.1038/nrmicro2818. Return to text.
- Sokurenko, E.V., Hasty, D.L. and Dykhuizen, D.E., Postadaptive mutations: gene loss and variation in bacterial pathogens, Trends in Microbiology 7(5):191–195, 1999. Return to text.
- MacNeil, L.T. and Walbout, A.J.M., Gene regulatory networks and the role of robustness and stochasticity in the control of gene expression, Genome Research 21(5):645–657, 2011. Return to text.
- Climate change after sin is inferred by reference to Genesis 3:21. Moving from no clothes to animal skin coverings to maintain comfort may suggest this; Arnold, D.L., Jackson, R.W., Waterfield, N.R. et al., Evolution of microbial virulence: the benefits of stress, Trends in Genetics 23(6):293–300, 2007; Beck, M.A., Nutritionally induced oxidative stress: effect on viral disease 1’2’3, American J. Clinical Nutrition 71(6):1676s–1679s, 2000. Return to text.
- Plantinga, A., Where The Conflict Really Lies: Science, religion and naturalism, Oxford University Press, Oxford, p. 150, 2011. Of course, Job 1 and 2 suggest this scenario to some modern readers.Return to text.
- Gal-Mor, O. and Finlay, B.B., Pathogenicity islands: molecular toolbox for bacterial virulence, Cellular Microbiology 8(11):1707–1719, 2006. Return to text.
- Somatic hypermutation represents a special category as the process contains highly sophisticated elements and the effects are limited to a single cell—cf. Peled, J.U., Kuang, F.L., Iglesias-Ussel, M.D. et al., The biochemistry of somatic hypermutation, Annual Review of Immunology 26:481–511, 2008. Return to text.
- Casadevall, A. and Pirofski, L., Host–pathogen interactions: redefining the basic concepts of virulence and pathogenicity, Infection and Immunity 67(8):3703–3713, 1999; Casadevall, A. and Pirofski, L., Host–pathogen interactions: the attributes of virulence, J. Infectious Diseases 184(3):337–344, 2001. Return to text.
- Vulić, M., Lenski, R.E. and Radman, M., Mutation, recombination, and incipient speciation of bacteria in the laboratory, Proceedings of the National Academy of Sciences USA 96(13):7348–7351, 1999; Willey, J.M., Sherwood, L.M. and Woolverton, C.J., Prescott’s Microbiology, 8th edition, McGraw Hill, New York, pp. 371–374, 2011. Return to text.
- Schubert, S., Picard, B., Gouriou, S. et al., Yersinia high-pathogenicity island contributes to virulence in Escherichia coli causing extraintestinal infections, Infection and Immunity 70(9):5335–5337, 2002. Return to text.
- Casadesús, J. and Low, D.A., Epigenetic gene regulation in the bacterial world, Microbiology and Molecular Biology Reviews 70(3):830–856, 2006; Low, D.A. and Casadesús, J., Clocks and switches: bacterial gene regulation by DNA adenine methylation, Current Opinion in Microbiology 11(2):106–112, 2008. Return to text.
- Kasuga, T. and Gijzen, M., Epigenetics and the evolution of virulence, Trends in Microbiology 21(11):575–582, 2013. Return to text.
- Gándara, B., Merino, A.L. and Martínez-Romero, E., Limited genetic diversity of Brucella spp., J. Clinical Microbiology 39(1):235–240, 2001; Martínez, M., Ugalde, R. A. and Almirón, M., Dimeric Brucella abortus Irr protein controls its own expression and binds haem, Microbiology 151:3427–3433, 2005. Return to text.
- Freiberg, C., Fellay, R., Bairoch, A. et al., Molecular basis of symbiosis between Rhizobium and legumes, Nature 387(6631):394–401, 1997. Return to text.
- Dunning Hotopp, J.C., Clark, M.E., Oliveira, D.C.S.G. et al., Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes, Science 317(5845):1753–1756, 2007. Return to text.
- Foster, J., Ganatra, M., Kamal, I. et al., The Wolbachia genome of Brugia malayi: endosymbiont evolution within a human pathogenic nematode, PloS Biology 3(4):e121, 2005, doi: 10.1371/journal.pbio.0030121. Return to text.
- Bian, G., Joshi, D., Dong, Y. et al., Wolbachia invades Anopheles stephensi populations and induces refractoriness to Plasmodium infection, Science 340(6133):748–751, 2013. Return to text.
- Gnanamanickam, S.S. (Ed.), Plant-associated Bacteria, Springer, Dordrecht, Netherlands, 2007. Return to text.
- Killham, K., Soil Ecology, University of Cambridge Press, Cambridge, p. 44, 2001. Return to text.
- Shipton, W.A., The Biology of Fungi Impacting Human Health, Trafford Publishing, Singapore, pp. 3, 150–151, 2012. Return to text.
- Irlinger, F., Safety assessment of dairy microorganisms: coagulase-negative staphylococci, International J. Food Microbiology 126(3):302–310, 2008. Return to text.
- Jensen, G.B., Hansen, B.M., Eilenberg, J. et al., The hidden lifestyles of Bacillus cereus and relatives, Environmental Microbiology 5(8):631–640, 2003. Return to text.
- Ezepchuk, Y.V., Biology of pathogenicity (theoretical review), Annual Review and Research in Biology 3(4):805–813, 2013; Klee, S.R., Bruzuszkiewicz, E.B., Nattermann, H. et al., The genome of Bacillus isolate causing anthrax in chimpanzees combines chromosomal properties of B. cereus and B. anthracis virulence plasmids, PloS One, 9 July 2010 | doi: 10:1371/journal.pone.0010986; Luna, V.A., King, D.S., Peak, K.K. et al., Bacillus anthracis virulent plasmid pX02 genes found in large plasmids of two other Bacillus species, J. Clinical Microbiology 44(7):2367–2377, 2006; Makino, S., Uchida, I., Terakado, N. et al., Molecular characterization and protein analysis of the cap region, which is essential for encapsulation in Bacillus anthracis, J. Bacteriology 171(2):722–730, 1989. Return to text.
- Maurelli, A.T., Fernandez, R.E., Bloch, C.A. et al., “Black holes” and bacterial pathogenicity: a large genomic deletion that enhances the virulence of Shigella spp. and enteroinvasive Escherichia coli, Proceedings of the National Academy of Sciences USA 95(7):3943–3948, 1998. Return to text.
- Freitag, N.E., Port, G.C. and Miner, M.D., Listeria monocytogenes—from saprophyte to intracellular pathogen, Nature Reviews Microbiology 7(9):623–628, 2009. Return to text.
- Vento, S., Cainelli, F. and Temesgen, Z., Lung infections after cancer chemotherapy, Lancet Oncology 9(10):982–992, 2008. Return to text.
- Silva, I.N., Tavares, A.C., Ferreira, A.S. et al., Stress conditions triggering mucoid morphotype variation in Burkholderia species and effect on virulence in Gallerina mellonella and biofilm formation in vitro, PloS One, 16 December 2013 | doi:10.1371/journal.pone.0082522. Return to text.
- Cox, E.C., Recombination, mutation and the origin of species, BioEssays 17(9):747–749, 1995; LeClerc, J.E., Li, B., Payne, W.L. et al., High mutation frequencies among Escherichia coli and Salmonella pathogens, Science 274(5290):1208–1211, 1996. Return to text.
- Donnenberg, M.S. and Whittam, T.S., Pathogenesis and evolution of virulence in enteropathogenic and enterohemorrhagic Escherichia coli, J. Clinical Investigation 107(5):539–548, 2001. Return to text.
- Bailey, K.L., Wyatt, T.A., Romberger, D.J. et al., Alcohol functionally upregulates Toll-like receptor 2 in airway epithelial cells, Alcoholism, Clinical and Experimental Research 33(3):499–504, 2009; Cangelosi, G.A., Ankenbauer, R G. and Nester, E.W., Sugars induce the Agrobacterium virulence genes through periplasmic binding protein and a transmembrane signal protein, Proceedings of the National Academy of Sciences USA 87(17):6708-6712, 1990; Fiester, S.E. and Actis, L.A., Stress responses in the opportunistic pathogen Acinetobacter baumannii, Future Microbiology 8(3):353–365, 2013. Return to text.
- Subromoni, S., Nathoo, N., Klimov, E. et al., Agrobacterium tumefaciens responses to plant-derived signalling molecules, Frontiers in Plant Science 5:322, 2014 | doi:10.3389/fpls.2014.00322. Return to text.
- Dubey, G.P. and Ben-Yehuda, S., Intercellular nanotubes mediate bacterial communication, Cell 144(4):590–600, 2011. Return to text.
- Fulsundar, S., Harms, K., Flaten, G.E. et al., Gene transfer potential of outer membrane vesicles of Acinetobacter baylyi and effects of stress on vesiculation, Applied and Environmental Microbiology 80(11):3469–3483, 2014; Hasegawa, Y., Futamata, H. and Tashiro, Y., Complexities of cell-to-cell communication through membrane vesicles: implications for selective interaction of membrane vesicles with microbial cells, Frontiers in Microbiology 6:633, 2015 | doi:10.3389/fmicb.2015.00633; Mashburn-Warren, L.M. and Whiteley, M., Special delivery: vesicle trafficking in prokaryotes, Molecular Microbiology 61(4):839–846, 2006. Return to text.
- Christie, G.E., Allison, H.E., Kuzio, J. et al., Prophage-induced changes in cellular cytochemistry and virulence; in: Hyman, P. and Abedon, S.T. (Eds.), Bacteriophages in Health and Disease, CABI, Wallingford, UK, pp. 33–60, 2012; Kuhl, S., Abedon, S.T. and Hyman, P., Diseases caused by phages, in Bacteriophages in Health and Disease, pp. 21–32, 2012; Plata, K., Rosato, A.E. and Wegrzyn, G., Staphylococcus aureus as an infectious agent: overview of biochemistry and molecular genetics of its pathogenicity, Acta Biochimica Polonica 56(4):597–612, 2009; Wagner, P.L. and Waldor, M.K., Bacteriophage control of bacterial virulence, Infection and Immunity 70(8):3985–3993, 2002; Ziebuhr, W., Ohlsen, K., Karch, H. et al., Evolution of bacterial pathogenesis, Cellular and Molecular Life Sciences 56(9–10):719–728, 1999. Return to text.
- Freeman, V.J., Studies on the virulence of bacteriophage-infected strains of Corynebacterium diphtheria, J. Bacteriology 61(6):675–688, 1951. Return to text.
- Uchida, T., Gill, D.M. and Pappenheimer, A.M., Mutation in the structural gene for diphtheria toxin carried by temperate phage β. Nature, New Biology 233(35):8–11, 1971; Uchida, T., Pappenheimer, A.M. and Greany, R., Diphtheria toxin and related proteins. 1. Isolation and properties of mutant proteins serologically related to diphtheria toxin, J. Biological Chemistry 248(11):3838–3844, 1973. Return to text.
- Casas, V., Magbanua, J., Sobrepeña, G. et al., Reservoir of bacterial exotoxin genes in the environment, International J. Microbiology 2010, article ID 754368 | doi:org/10.1155/2010/754368. Return to text.
- Hassan, F., Kamruzzaman, M., Mekalanos, J J. et al., Satellite phage TLC φ enables toxigenic conversion of CTX phage through dif site alteration, Nature 467(7318):982–985, 2010. Return to text.
- Williamson, K.E., Soil phage ecology: abundance, distribution, and interactions with bacterial hosts; in: Witzany. G. (Ed.), Biocommunication in Soil Microorganisms, Springer, New York, pp. 113–136, 2011. Return to text.
- Hacker, J. and Kaper, J.B., Pathogenicity islands and the evolution of microbes, Annual Review of Microbiology 54:641–679, 2000; Toh, H., Oshima, K., Toyoda, A. et al., Complete genome sequence of the wild-type commensal Escherichia coli strain SE15, belonging to phylogenetic group B2, J. Bacteriology 192(4):1165–1166, 2010. Return to text.
- Antonenka, U., Nölting, C., Heesemann, J. et al., Horizontal transfer of Yersinia high-pathogenicity island by the conjugative RP4 att B target-presenting shuttle plasmid, Molecular Microbiology 57(3):727–734, 2005. Return to text.
- Gilbert, C., Schaack, S., Pace, J.K. et al., A role for host–parasite interactions in the horizontal transfer of DNA transposons across animal phyla, Nature 464(7293):1347–1350, 2010; Yue, J., Hu, X., Sun, H. et al., Widespread impact of horizontal gene transfer on plant colonization of land, Nature Communications 3, article 1152, 2012 | doi:10.1038/ncomms2148. Return to text.
- Corvaglia, A.R., Francois, P., Hernandez, D. et al., A type III-like endonuclease functions as a major barrier to horizontal gene transfer in clinical Staphylococcus aureus strains, Proceedings of the National Academy of Sciences USA 107(26):11954–11958, 2010; Stucken, K., Koch, R. and Dagan, T., Cyanobacterial defense mechanisms against foreign DNA transfer and their impact on genetic engineering, Biological Research 46(4):373–382, 2013. Return to text.
- LeClerc, Li, Payne et al., ref. 33. Return to text.
- Freiberg, C., Fellay, R., Bairoch, A. et al., Molecular basis of symbiosis between Rhizobium and legumes, Nature 387(6631):394–401, 1997; Brown, N.F. and Finlay, B.B., Potential origins and horizontal transfer of type III secretion systems and effectors, Mobile Genetic Elements 1(2):118–121, 2011; Silva, D.C F., Silva, R.C., Ferreira, R.C. et al., Examining marginal sequence similarities between bacterial type III secretion systems and Trypanosoma cruzi surface proteins: horizontal gene transfer of convergent evolution, Quantitative Biology, Populations and Evolution, Cornell University, 2012, www.arxiv.org/abs/1211.6611v1. Return to text.
- de la Riva, G.A., González-Cabrera, J., Vázquez-Padrón, R. et al., Agrobacterium tumefaciens: a natural tool for plant transformation, EJB Electronic J. Biotechnology 1(3):118–133, 1998. Return to text.
- Monier, J-M., Bernillon, D., Kay, E. et al., Detection of potential transgenic plant DNA recipients among soil bacteria, Environmental Biosafety Research 6(1–2):71–83, 2007; Pontiroli, A., Rizzi, A., Simonet, P. et al., Visual evidence of horizontal gene transfer between plants and bacteria in the phytosphere of transplastomic tobacco, Applied and Environmental Microbiology 75(10):3314–3122, 2009. Return to text.
- Hofkin, B.V., Living in a Microbial World, Garland Science, Taylor and Francis Group, LLC, New York, p. 211, 2011. Return to text.
- Nembaware, V., Seoighe, C., Sayed, M. et al., A plant natriuretic peptide-like gene in the bacterial pathogen Xanthomonas axonopodis may induce hyperhydration in the plant host: a hypothesis of molecular mimicry, BMC Evolutionary Biology 4:10, 2004 | doi:10.1186/1471-2148-4-10. Return to text.
- Yue, Hu, Sun et al., ref. 47. Return to text.
- Schaack, S., Gilbert, C. and Feschotte, C., Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution, Trends in Ecology and Evolution 25(9):537–546, 2010. Return to text.




Readers’ comments
Comments are automatically closed 14 days after publication.