Weird synthetic proteins are no help to evolution
Changing the genetic code points to design

Introduction
Scientists have actually succeeded in changing the genetic code! This experiment has some interesting implications for theories of origins. We will explain why, but first we all need to understand how cells turn DNA into proteins.
Amino acids are some of the most basic molecular building blocks of life. Living things use 20 different amino acids to make proteins, the macromolecules which carry out almost all functions in the cell. Proteins function as enzymes, ion channels, structural elements, receptors, and molecular machines of all sorts. Each amino acid has a nitrogen-carbon-carbon (N-C-C) backbone with a functional side chain that is attached to the central carbon (figure 1). The side chain is a group of atoms which give each amino acid its distinctive characteristics. For example, they can be electrically charged or neutral, contain special atoms, or be long or short, or big and bulky. These properties are combined to give each protein a unique shape and function. The 20 amino acids can be seen in figure 2.
Unlike the 20-letter protein alphabet, DNA is made up of only four letters called ‘bases’: adenine (A), cytosine (C), guanine (G), and thymine (T). RNA is a similar molecule, with uracil (U) replacing T everywhere it occurs. Messenger RNA (mRNA) is an incredibly important molecule. Before proteins are made, the stretch of DNA (a ‘gene’) that codes for the protein is first transcribed (written) into RNA. This then migrates from the nucleus to the cytoplasm. Importantly, each set of three DNA bases codes for a single amino acid in the resulting protein. Each set of three DNA letters is called a ‘codon’. To get from DNA to protein, molecules called ‘transfer RNAs’ (tRNAs) are matched to each codon. At one end, they carry an amino acid.

The mRNA is translated into proteins in a molecular factory called the ribosome (figure 3), a highly complex, multi-subunit macromolecular machine made of 120 different proteins and RNA molecules. The ribosome has two main sections, the 30S and the 50S subunits. The smaller (30S) subunit is where the codons are matched to individual tRNAs. The larger (50S) subunit is where the amino acids are brought close to one another and formed into an amino acid chain, which makes the protein.
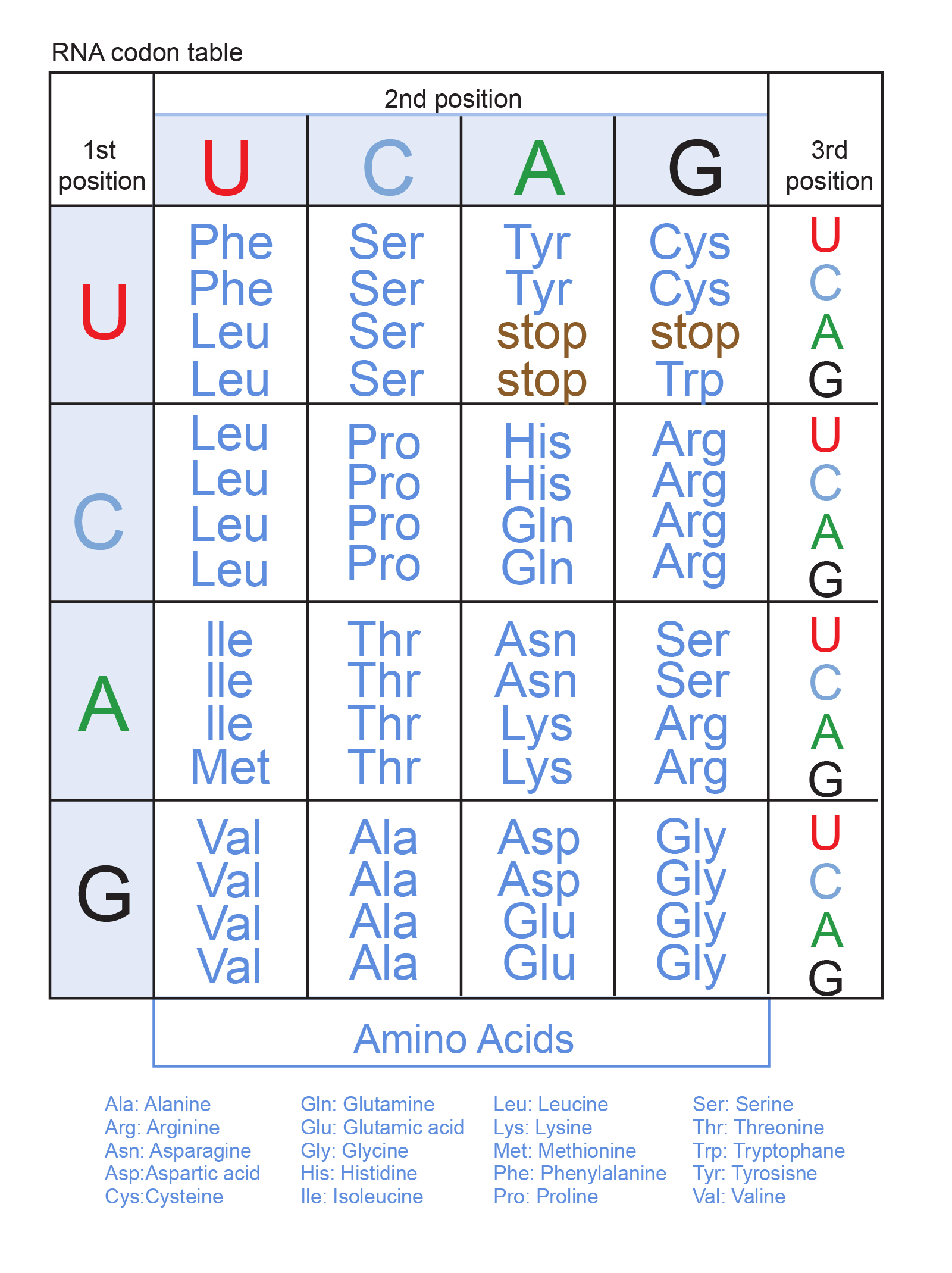
Since there are four possible RNA letters, and three RNA letters in each codon, this makes 4 x 4 x 4 = 64 possible codons (AAA, AAC, AAG, AAU, etc.). But there are only 20 amino acids, meaning that some of them are coded for by multiple codons. Thus, the genetic code is redundant. For example, the amino acid methionine has only one codon, ATG, but leucine has six (figure 4). Three of the 64 codons are used as a ‘stop’ signal to let the cell know that the protein sequence is at its end. This is where the interesting stuff comes in.
Synthetic amino acids
There are many different possible amino acids, but life only uses 20. Wouldn’t it be great if we could assign every single codon to a single amino acid? There would be 64 possible amino acids, which would allow an even larger diversity of proteins.
That is why the laboratory of Jason Chin, at the Medical Research Council’s Laboratory of Molecular Biology in Cambridge decided to engineer a slightly altered genetic code. They reasoned that a small part of the genome could be allocated to code for extra amino acids besides the regular, or “canonical”, amino acids that are so well-known to molecular biologists. By doing so, the Chin lab hopes to create new proteins, which, for example, could be used in the fight against disease.1
The amino acid serine has six codons, and there are also three stop codons. They took the approximately 4-million-letter genome of the well-known laboratory bacterium Escherichia coli and stripped out every occurrence of two serine codons and one of the stop codons (18,214 total positions in the genome).
While the bacterium’s altered genetic code didn’t actually code for any new amino acid, the bacterium is still viable even after its genetic material was changed in a substantial manner. However, the potential now exists for new and interesting things to be done with this bacterium. Nothing has been done, but the potential is there to add novel amino acids. However, this will not be easy.

The entire exercise is an example of intelligent design. For example, before adding a new amino acid to the genetic code, they would have to add a new amino acid manufacturing pathway. But to get this novel amino acid into a protein they would first have to develop a system for attaching the amino acid to a tRNA. The cell usually does this with a class of proteins called aminoacyl-tRNA synthetases. There is a different one for each amino acid, so they will have to get the original to stop recognizing the codon normally associated with that amino acid. They will then have to invent a new system that matches the new amino acid to the newly released codon. Then they will have to reprogram a protein-coding gene to re-include the codon they stripped away at the beginning of the experiment. Can this be done? Possibly. But the fact that it takes so much thought and experimentation, using the best engineering known, to modify the cell only goes to demonstrate that the cell was originally engineered by a very thoughtful Creator.
Chin’s lab was also able to create a new kind of ribosome which can read four bases at a time, as opposed to the usual three. This could allow genes to code up to 4 x 4 x 4 x 4 = 256 amino acids, making possible even more protein variability with even more novel amino acids. However, no four-based codons were actually produced.
In other experiments, more than 100 different unnatural amino acids have been generated and successfully built into mammalian cells. Some contain special atoms, like fluorine (figure 5). Such amino acids could increase the functionality of proteins. However, these novel amino acids are not actually coded for or produced by the cells themselves. Rather, they are chemically attached in the lab to an altered stop codon tRNA, added to the bacterial growth medium, and the bacteria are tricked into taking it into their cells. It can then be built into the growing protein. But if the unnatural amino acid is not present in the culture medium containing the cells, it will not be built into the protein at all, and the protein chain stops growing.2
What does this mean for evolution?
What would novel, non-canonical amino acids mean for evolution? Would organisms with more than 20 amino acids be able to ‘evolve’ faster?
Hardly.
As mentioned previously, the genetic code is redundant. For example, the codons CCA, CCC, CCG and CCT all code for the amino acid proline (Pro). Thus, we can represent the proline codon with the CCN, where N represents any of the four DNA letters. This redundancy in the triplet code allows for so-called “silent” mutations to occur. Since the last position of the triplet can be anything, any mutation there has no effect. For example, if CCT mutates to CCC, both codons still code for proline. Silent mutations are otherwise known as synonymous mutations, because they “say the same thing”. They don’t cause any damage to the organism and their effect is usually unnoticeable.3

However, if all 64 possible codons would code for separate amino acids, synonymous mutations would be impossible. Any mutation, even in the last position of the codon, would necessarily mutate the amino acid into another one. In this scenario, the previously mentioned triplets CCT and CCC would now code for different amino acids. This would likely damage the protein, because the side chain of the new amino acid would likely not fit in the protein. Coding for more amino acids would actually make the genome more fragile. The fact that the genetic code is redundant allows mutations to occur in the last position of codons without any effect.
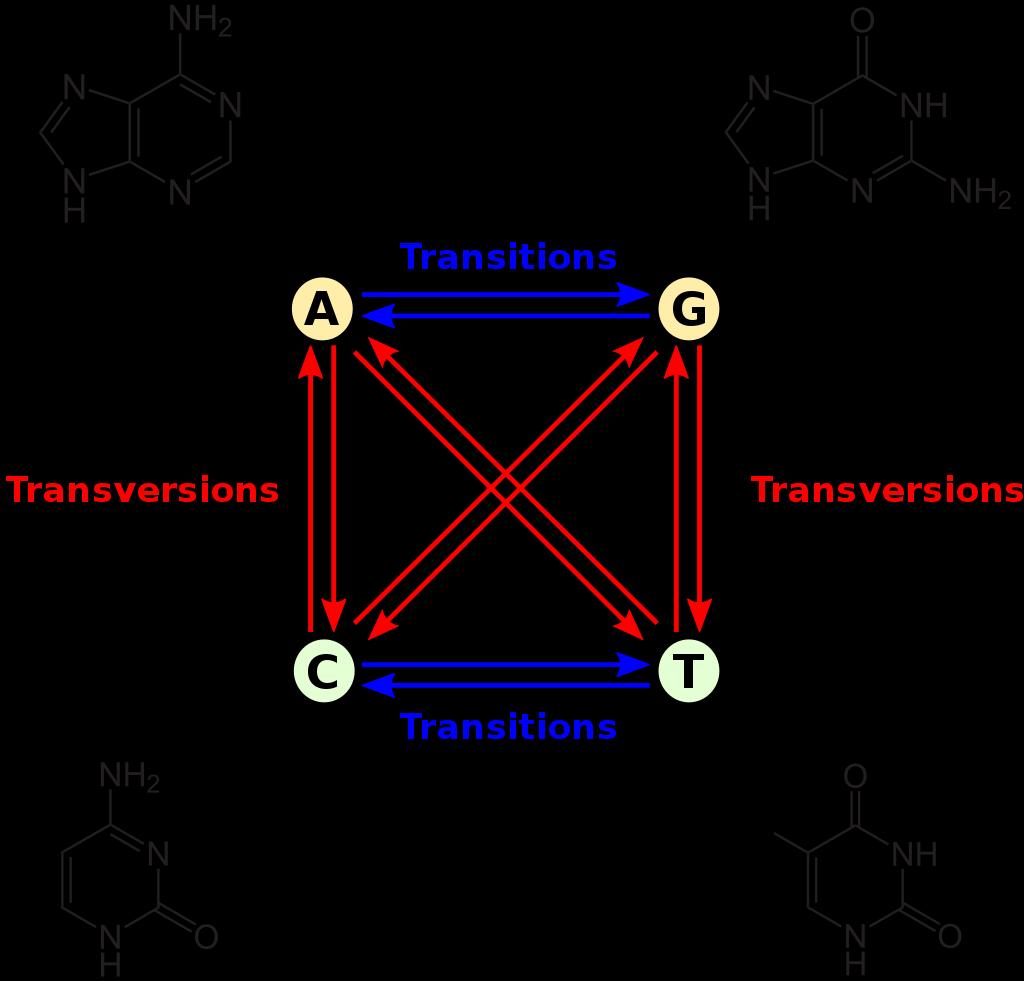
The genome is actually robust to change. It appears to have been designed that way on purpose. First, most mutations are synonymous. This was shown for example, in a study of cancer-related genes.4 Second, most non-synonymous mutations involve swapping amino acids that are similar to one another. This minimizes the effect on the shape and activity of most proteins. For example, a change from leucine to isoleucine involves two amino acids which differ only in a methyl group (–CH3). In a study of 139,477 mutations in 4,582 genes, researchers found that 21 of the 30 most frequent mutations involve changes between amino acids which belong to the same structural group: hydrophilic (soluble in water), neutral or hydrophobic (non-soluble in water).5 Third, most mutations are transitions. A transition is when a purine (A or G) is swapped for a purine or a pyrimidine (C or T) is swapped for a pyrimidine (figure 6). The code is set up so that the most common mutations (transitions) have a smaller potential effect.6 Transversions are more likely to replace an amino acid with one from another functional group. For example, if CAC mutates to CAA (a C to A transversion, which is rare), the charged amino acid histidine changes to the uncharged glutamine.
An increased amino acid repertoire may seem advantageous, but more amino acids need more biochemical pathways to produce the increased number of novel amino acids. This would require more enzymes, more regulatory factors and more transfer RNA molecules to handle each new amino acid. The more unnatural the amino acid, the more different the biochemical processes would have to be in order to generate that amino acid. Indeed, other researchers had to create a new biosynthetic pathway to add a novel amino acid, p-aminophenylalanine, in E. coli (figure 7), something that the Chin lab did not do in their own experiments.7 Doing this increases the energy demand on the cell. An increased number of amino acids has a similar problem as an increased number of DNA bases.

Getting back to the question of using four, instead of three, letters per codon, since all of life is based on codon triplets, all genes in a genome with four-letter codons would have to be rewritten. Unless the scientists wanted to rewrite the entire genome, two parallel but incompatible coding systems would have to exist side-by-side within the same cell. Genes using triplets would not work with a four-letter-based code, and vice-versa. This would be similar to working on a computer which uses both 0, 1, and 2 (ternary) as well as 0 and 1 (binary) operations. The field of quantum computing has been exploring such things, but this involves high technology, not random chance processes. Plus, a four-letter-based codon system is inefficient in mammals.8 There is no advantage to the organism, even though we might be able to capitalize on some new property inherent in such a system.
Conclusion
Life contains a multitude of complex and robust codes. A code presupposes an Intelligent Designer, who created the genetic code to work efficiently in living things. The genetic code works efficiently, and even though it is robust to mutation, random mutations only break it apart. Laboratory experiments only show that expanding the genetic code requires intelligent input. Cells, DNA, and the genetic code all point to the fact that life was created by an Almighty, all-knowing God.
References and notes
- Highfield, R., Weird synthetic proteins could let us build new kinds of life, wired.co.uk/article/synthetic-proteins-new-life, 26 August 2019. Return to text.
- Nödling, A.R., Spear, L.A., Williams, T.L., Luk, L.Y.P., and Tsai, Y.H., Using genetically incorporated unnatural amino acids to control protein functions in mammalian cells. Essays Biochem. 63(2):237–266, 2019. Return to text.
- This is not to say all synonymous mutations are completely neutral. Different ribosome families are designed to optimally work with different gene families, which in turn have different codon preferences. Also, some proteins require a specific timing of translation to fold properly. Changing a common codon to a rare codon might create a delay in translation and a useless protein, even though the amino acid sequence is unaltered. Return to text.
- Chu, D. and Wei, L., Nonsynonymous, synonymous and nonsense mutations in human cancer-related genes undergo stronger purifying selections than expectation. BMC Cancer. 19(1):359, 2019. Return to text.
- Sasidharan, R. and Chothia, C., The selection of acceptable protein mutations, Proc. Natl. Acad. Sci. USA. 104(24):10080–10085, 2007. Return to text.
- Maeshiro, T. and Kimura, M., The role of robustness and changeability on the origin and evolution of genetic codes. Proc. Natl. Acad. Sci. USA 95(9):5088–5093, 1998. Return to text.
- Wang, Q., Parrish, A.R., and Wang, L., Expanding the genetic code for biological studies. Chem. Biol. 16(3):323–336, 2009. Return to text.
- Niu, W., Schultz, P.G., and Guo, J., An expanded genetic code in mammalian cells with a functional quadruplet codon. ACS Chem. Biol. 8:1640–1645, 2013. Return to text.



Readers’ comments
Comments are automatically closed 14 days after publication.