Journal of Creation 12(1):3–5, April 1998
Browse our latest digital issue Subscribe
Design in living organisms (motors: ATP synthase)
In our everyday experience, we can usually tell whether something has been designed. The main evidence is high information content. The information content of any arrangement is the size, in bits, of the shortest algorithm required to generate that arrangement. This means that repetitive structures, like crystals, have a low information content, because all that is needed is to specify a few positions, then the instructions ‘more of the same’. The difference between a crystal and an enzyme or DNA is like the difference between a book containing nothing but ABCD repeated and a book of Shakespeare.
On a practical level, the information specifies the many parts needed to make machines work. Often, the removal of one part can disrupt the whole machine. Biochemist Michael Behe, in his book Darwin’s Black Box (right), calls this irreducible complexity.1 He gives the example of a very simple machine: a mousetrap. This would not work without a platform, holding bar, spring, hammer and catch, all in the right place. The thrust of Behe’s book is that many structures in living organisms show irreducible complexity, far in excess of a mousetrap or indeed any man-made machine.
Motors: a case study
Motors are irreducibly complex, because they need many parts working together to function. For example, an electric motor needs a power source, fixed stator, movable rotor, and a commutator or slip rings.
The more parts needed for a machine, the harder it is to make it smaller. Miniaturisation is such a vital part of the computer industry, and the best human minds are constantly working at it. And though miniaturised motors would be very useful, e.g. for unblocking clogged arteries and blood cleaning, the number of parts makes it difficult to make them below a certain size. But ingenious scientists are making them smaller all the time.2
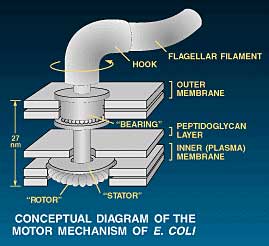
However the design in living organisms has far exceeded our most painstaking efforts. Bacteria propel themselves using flagella (singular flagellum, from the Latin for whip), filaments propelled by a true rotary motor. This motor is only the size of a virus, thus far smaller than anything man-made. Yet it can rotate at over 1000 times per second.3
But even this impressively tiny motor is not the tiniest in God’s creation. In a paper published in March 1997, Hiroyuki Noji et al. directly observed the rotation of the enzyme F1-ATPase, a subunit of a larger enzyme, ATP synthase.4,5 This had been suggested as the mechanism for the enzyme’s operation by Paul Boyer.6 Structural determination by X-ray diffraction by a team led by John Walker had supported this theory.7 A few months after Noji et al published their work, it was announced that Boyer and Walker had won a half share of the 1997 Nobel Prize for Chemistry for their discovery.8
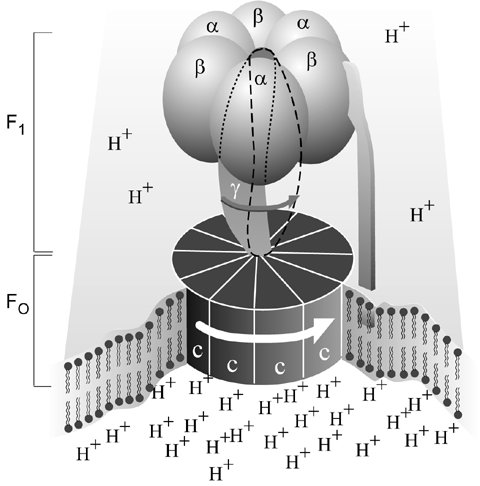
The F1-ATPase motor has nine components—five different proteins with the stoichiometry of 3a:3b:1g:1d:1e. In bovine mitochondria, they contain 510, 482, 272, 146 and 50 amino acids respectively, so Mr = 371,000. F1-ATPase is a flattened sphere about 10 nm across by 8 nm high—so tiny that 1017 would fill the volume of a pinhead. This has been shown to spin ‘like a motor’ to produce ATP, a chemical which is the ‘energy currency’ of life.9 This motor produces an immense torque (turning force) for its size—in the experiment, it rotated a strand of another protein, actin, 100 times its own length. Also, when driving a heavy load, it probably changes to a lower gear, as any well-designed motor should.
ATP synthase also contains the membrane-embedded FO subunit functioning as a proton (hydrogen ion) channel. Protons flowing through FO provide the driving force of the F1-ATPase motor. They turn a wheel-like structure as water turns a water wheel, but researchers are still trying to determine precisely how. This rotation changes the conformation of the three active sites on the enzyme. Then each in turn can attach ADP and inorganic phosphate to form ATP. Unlike most enzymes, where energy is needed to link the building blocks, ATP synthase uses energy to link them to the enzyme, and throw off the newly formed ATP molecules. Separating the ATP from the enzyme needs much energy.
Note: the names of the two components are historical. The F1 unit comes from the term ‘Fraction 1’. The name FO (written as a subscript capital O, not zero) is due to its being the oligomycin-binding fraction. Oligomycin is an antibiotic that kills bacteria by blocking the proton channel of the FO subunit.
ATP synthase is the central enzyme in energy conversion in mitochondria (where they are embedded into the cristae, folds in the mitochondrion’s inner membrane), chloroplasts and bacteria. This probably makes ATP synthase the most ubiquitous protein on Earth. Since energy is required for life, and all life uses ATP as its energy currency (each of us synthesizes and consumes half our bodyweight of ATP per day!), life could not have evolved before this motor was fully functional. Natural selection by definition is differential reproduction, so requires self-reproducing entities to start with. So even if a series of gradual steps could be imagined up this peak of ‘Mount Improbable’, there would be no natural selection to enable that climb.
One of the Nature articles was appropriately entitled ‘Real Engines of Creation’. Unfortunately, despite the evidence for exquisite design, many scientists (including the editor of Nature) still have a blind faith that mutations and natural selection could build such machines.
Would any evidence convince evolutionists?
The famous British evolutionist (and communist) J.B.S. Haldane claimed in 1949 that evolution could never produce “various mechanisms, such as the wheel and magnet, which would be useless till fairly perfect.”10 Therefore such machines in organisms would, in his opinion, prove evolution false. These molecular motors have indeed fulfilled one of Haldane’s criteria. Also, turtles11 and monarch butterflies12 which use magnetic sensors for navigation fulfil Haldane’s other criterion. I wonder whether Haldane would have had a change of heart if he had been alive to see these discoveries. Many evolutionists rule out intelligent design a priori, so the evidence, overwhelming as it is, would probably have no effect.
Animations (off-site)
- CMI’s own animation is on the right
- ATP Synthase: an interpretation by Donald Nicholson
- Virtual Cell Animation Collection, Molecular and Cell Biology Learning Center
References
- Behe, M.J., Darwin’s Black Box: The Biochemical Challenge to Evolution, The Free Press, New York, 1996. Reviewed by Ury, T.H., Journal of Creation 11(3):283–291, 1997. Return to text.
- Hogan, H., Invasion of the micromachines, New Scientist 150(2036):28–33, 1996. Return to text.
- For a good description, see Behe, Ref. 1. Return to text.
- Hiroyuki Noji et al., Direct observation of the rotation of F1-ATPase, Nature 386(6622):299–302, 1997. Comment by Block, S. Real engines of creation. Same issue, pp. 217–219. Return to text.
- Wu, C., Molecular motor spins out energy for cells, Science News 151(12):173, 1997. Return to text.
- Boyer, P., Biochim. Biophys. Acta 1140:215–250, 1993. Return to text.
- Abrahams, J.P. et al., Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria, Nature 370(6491):621–628, 1994. Comment by Cross, R.L. Our primary source of ATP. Same issue, pp. 594–595. Return to text.
- Service, R.F., Awards for high-energy molecules and cool atoms, Science 278(5338):578–579, 1997. The third winner is Jens Skou of the University of Aarhus in Denmark. Forty years ago, he was the first to identify an enzyme that moves substances through cell membranes (in this case, sodium and potassium ions). This is a key function of all cells. Return to text.
- ATP stands for adenosine triphosphate. It is a high energy compound, and releases this energy by losing a phosphate group to give ADP, adenosine diphosphate. Return to text.
- Dewar, D., Davies, L.M. and Haldane, J.B.S., Is Evolution a Myth? A Debate between D. Dewar and L.M. Davies vs. J.B.S. Haldane, Watts & Co. Ltd / Paternoster Press, London, p. 90, 1949. Return to text.
- Sarfati, J.D., Turtles can read magnetic maps, 1997; creation.com/turtlemap. Return to text.
- Poirier, J.H., The magnificent migrating monarch, Creation 20(1):28–31, 1997; creation.com/monarch. But monarchs only use the earth’s magnetic field to give them the general direction, while they rely on the sun’s position for most of their navigation. Return to text.






Readers’ comments
Comments are automatically closed 14 days after publication.