Journal of Creation 35(2):87–90, August 2021
Browse our latest digital issue Subscribe
Doppelgänger protein ‘Signal Recognition Particle 14’ refutes evolution
Evolution has difficulty explaining the similarities in gene distribution and sequences in unrelated groups of organisms. Doppelganger genes are genes that are highly homologous to one another but are found on disjunct parts of the alleged evolutionary tree. One such doppelganger gene codes for a protein called the Signal Recognition Particle 14 (SRP14), which has a poly-alanine C-terminus. This protein is known to occur in most species of the primate apobaramin, but also in a species of bat, called the little brown bat. Another SRP14 isoform without a poly-alanine C-terminus also occurs not only in this bat species but also in primates, rodents, protists, and yeast. Evolution cannot explain why this isoform is conserved throughout the alleged evolutionary tree, yet two structurally significantly different isoforms occur in one species. Creation theory offers a better explanation, namely that these two SRP14 isoforms are distinctly created functional units.

The distribution of genes and their sequential similarities and differences challenge evolutionary theory. For example, the gradual transformation of one family of genes to another has never been observed, indeed it cannot even be observed. The de novo appearance of genes from random non-coding sequence has also never been observed. Evolutionary conservation of genes is also an oxymoron—how can proteins manage to evolve by staying the same for millions of years?1
Doppelganger genes perform the same function and have highly similar structures, appearing in two distinct clades far from one another on the alleged evolutionary tree.2 For example, the gene which codes the protein prestin, which plays a role in echolocation, is expressed in the inner ear of whales and bats.3 While it is broadly present in a number of vertebrate animals, it is also present in species of Ascomycetes as well as Blastocystis. This latter group of heterokont parasites belongs to a larger group called Stramenopiles, which includes organisms such as algae, diatoms, and water moulds.
Besides their structural similarity, doppelganger genes also show sequential similarity with one another. In contrast, analogous genes are found in unrelated organisms, and have the same function, albeit with totally different structures. They are thought to arise via convergent evolution, although it is difficult to understand how completely different sequences can give shape to the same structure.
Doppelganger genes pose a serious problem to evolutionary theory. The same, or very similar, sequences of mutations would have had to occur in different DNA sequences in two different branches of the evolutionary tree to produce two genes that are very homologous to one another. To explain this, evolutionists allege that the same kind of gene once existed in all of the organisms leading from the common ancestor of the two clades all the way up to the two clades themselves. Afterwards, all of these genes were lost in the intermediate species connecting the two clades during several rounds of massive gene deletion.
The probabilities of these events occurring are extremely low. A much simpler explanation is that these genes are created functional units which already existed in disjunct clades; that is, baramins. These proteins differ according to the cellular context they function in.
The signal recognition particle complex
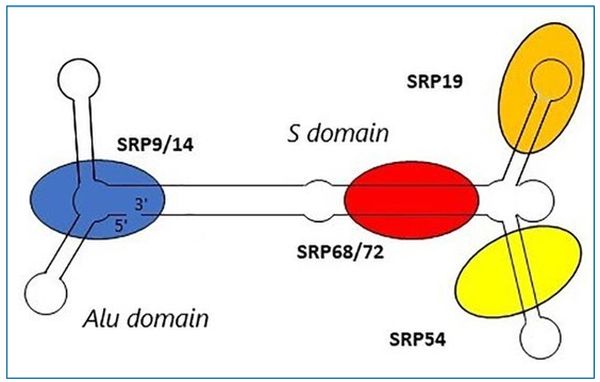
The cell produces several proteins, which fulfil their function in the endoplasmic reticulum (ER) or the thylakoid membrane in bacteria. These proteins contain a hydrophobic signal sequence at their N-terminus which tells the signal recognition particle (SRP) complex to translocate them into the ER. The SRP complex is a cytoplasmic ribonucleoprotein—a multi-unit macromolecule made up of proteins and RNA. The different components of the SRP complex are shown in figure 1.
The SRP consists of two domains, the S domain and the Alu domain. The S domain, consisting of SRP19, 54, 68, and 72, is responsible for binding the nascent signal sequence by SRP54 as it arises from the ribosome. This process is called ‘elongation arrest’.
The S domain is connected to the Alu domain by the SRP RNA. The Alu domain is responsible for the elongation of proteins once they exit the ribosome and consists of the proteins SRP9 and 14.4 SRP14 has a very interesting distribution in living organisms, which classifies its encoding gene as a doppelganger gene.
Conservation and distribution of SRP14 contradicts evolution
The proteins in the SRP occur in primates, bats, rodents, yeast, and even protists such as Plasmodium falciparum.5 Even though there are structural differences within different groups, they perform the same function.6 The Alu RNA structure of the Archaean Pyrococcus horikoshii is so similar to that of human that even the human SRP9/14 heterodimer can bind to it.7 Interestingly, the yeast SRP has no SRP9 homologue, yet still shows elongation arrest which is dependent upon two Srp14p proteins which substitute SRP9. Furthermore, yeast also contains the protein Srp21p which has no known homologues in other organisms.8,9
SRP9/14 homologues are absent in Archaea and Eubacteria. However, Nakamura et al. think that the histone-like protein HBsu from Bacillus subtilis is a functional analogue in Eubacteria. In B. subtilis, this protein binds the Alu domain of the scRNA, which plays a role in membrane docking, the lack of which leads to depletion of extracellular enzymes.10
Since the structure and function of SRP14 is so conserved in very different organisms, their very existence is evidence of non-evolutionary stasis. Furthermore, the existence of functional SRP14 analogues in prokaryotes contradicts sequential transition between prokaryotes and eukaryotes.11
Comparison between primates and bats
Interestingly, in primates, the SRP14 protein contains a 16-amino-acid-long segment containing 15 alanines: A9PA4TA2 in its C-terminus. This poly-alanine tract could be due to the expansion of the trinucleotide GCA, with the G mutating to A or C to get threonine (T) or proline (P). However, the presence of the poly-alanine C-terminus does not confer any differential RNA-binding onto SRP14, although this may not be important.12 The poly-alanine C-terminus is absent in some primates, such as Carlito syrichta (the Philippine tarsier, XP_008068029.1) and Microcebus murinus (the Gray mouse lemur, XP_012621845.1).
Among bats, the only known SRP14 protein that contains a poly-alanine C-terminus is found in Myotis lucifugus, the little brown bat (G1PG47_MYOLU). In M. lucifugus, the poly-alanine C-terminus is somewhat different than in primates, with a sequence of A2GA9. Nevertheless, both primates and M. lucifugus share a stretch of nine alanines in their C-terminus. An alignment of 11 SRP14 proteins from four primate and six bat species is shown in figure 2. The identity matrix comparing the sequence similarity between these 11 SRP14 proteins as well as the proteins’ accession numbers are found in supplementary file 1 (see ‘Materials and methods’ section below).
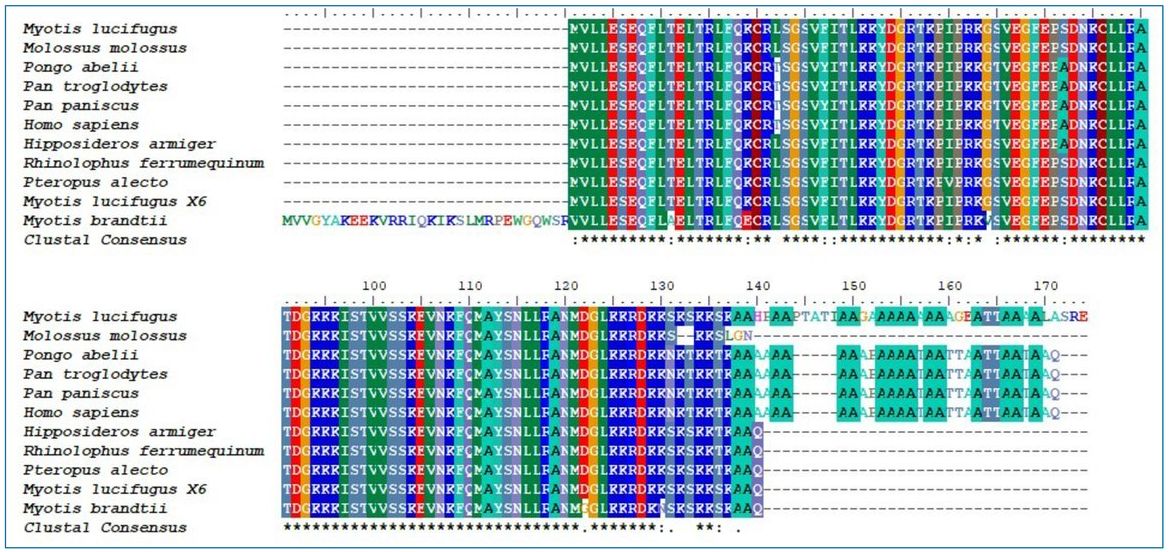
Even more interestingly, M. lucifugus has an X6 isoform of the SRP14 protein (XP_006093069.1), which is at least 98% similar in its sequence to the SRP14 protein found in three other bat species: Hipposideros armiger, Pteropus alecto, and Rhinolophus ferrumequinuum. In comparison, its other SRP14 isoform is only 74.3–75% similar to the SRP14 proteins in these three species.
The obvious question is, how did such sequentially different variants of the same gene arise in M. lucifugus? Does evolution really say that this one sequence underwent significant change in the life of this one bat species? It makes much more sense to view both the regular and the X6 isoforms of the SRP14 genes as separately created functional units. It is as if God directly created a poly-alanine SRP14 protein isoform in M. lucifugus to defy evolution.
Comparison of secondary structure
Figure 3 shows a secondary structure plot for three SRP14 proteins from human, M. lucifugus, and R. ferrumequinuum. H stands for an amino acid which is a part of an alpha-helix, E for a beta-sheet amino acid, C for a coiled coil amino acid, and T for a turn amino acid. Interestingly, all three proteins contain an 11-amino-acid-long structure TTTEEEEEECC, which corresponds to the protein motif GKKKISTVVSS, from positions 63 to 74. Both bat proteins contain the secondary structure TTTTTCEEEEETTTTTTCEEECTTCCCEETCTTTT, from positions 19 to 54. However, M. lucifugus SRP14 is more similar to human SRP14 in its alpha-helix C-terminus, as well as its overall length, not to mention the poly-alanine segment discussed earlier.

Baraminic relationships based on SRP14 sequence identities

A sequence identity matrix was derived from the alignment of the 11 SRP14 proteins in figure 2. This matrix was then displayed in the heatmap shown in figure 4. The Hopkins statistic is 0.641, which indicates fair clusterability. This way, baraminic relationships can be visualized based on sequential similarities. The caveat is that since this baraminogram is based on the sequence identity matrix of a single protein, we cannot draw strong conclusions from these results.
There are four clusters, two large ones and two singleton species. Both larger groups are statistically significant, with a p-value of 2.4 × 10–4 for the poly-alanine C-terminus isoform in the primate apobaramin, and 9.3 × 10–18 for the other isoform in the bat apobaramin (see ‘stats’ tab in supplementary file 1). A silhouette plot also shows that there are four optimal clusters (supplementary figure 1). The maximum silhouette value is 0.756, which indicates statistically significant clusters.
These four SRP14-based clusters centre around two basic SRP14 isoforms, the poly-alanine isoform present in the primate apobaramin, as well as the non-poly-alanine isoform present mainly in the bat apobaramin. The Myotis brandtii SRP14 (EPQ10995.1) differs from the other bat non-polyalanine protein isoforms sequences due to a 30-amino-acid leader sequence in the N-terminus which is absent in the other protein sequences.
Discussion and conclusion
The distribution and sequence similarities of the SRP14 protein in the SRP complex are a thorny riddle for evolution. Its conserved structure and function in organisms from yeast to human defy evolution because evolution is change, not stasis. Furthermore, analogous proteins in archaea and bacteria contradict gradual sequential variation in this protein over time.
Besides this, the presence of the poly-alanine C-terminus isoform of SRP14 in primates and M. lucifugus is difficult for evolution to resolve. Since bats and primates are disjunct clades (apobaramins), it is hard to explain how the same protein could even arise through random mutations in lineages leading up to these two groups. This classifies SRP14 as a doppelganger gene. Furthermore, M. lucifugus has two different SRP14 isoforms, each with a significantly different structure. How evolution posits significant sequential changes within a single species, but also that the SRP14 protein is conserved all throughout the evolutionary tree, is difficult to understand.
Creation theory offers a much simpler explanation without forcing the data into an evolutionary tree. Rather, different protein isoforms, such as those seen in SRP14, can be seen as discrete functional units created within individual species or baramins.
Materials and methods
All protein sequences were downloaded from the National Center for Biotechnology Information (NCBI) database. Secondary protein structures were generated for the three SRP14 proteins in figure 4 using the ̔garnier̕ program from the EMBOSS bioinformatics software suite at bioinformatics.nl/cgi-bin/emboss/garnier.13 The heatmap was constructed using the ‘heatmap’ command in R version 4.0.3. The clustering method was set to ‘ward.D2’.
Construction of Hidden Markov Model
A Hidden Markov Model (HMM) was constructed using the ‘hmmbuild’ command on an alignment of ten primate SRP14 proteins. The accession numbers of these proteins are listed in supplementary file 2. The software used to construct the HMM was downloaded from hmmer.org. Using an HMM is a more sensitive way of finding homologous proteins than BLAST14 This HMM and supplementary files 1 and 2 are available at github.com/csmatyi/SRP14. The SRP14 HMM was then used to locate bat protein sequences at the HMM search website at ebi.ac.uk/Tools/hmmer/search/hmmsearch, using default parameters, with the Organism option set to ‘Chiroptera’.15
References and notes
- Cserháti, M., Creation aspects of conserved non-coding sequences, J. Creation 21(2):101–108, 2007. Return to text.
- Cserhati, M., Lazarus and doppelganger genes, Creation Research Society Conference, Ann Arbor, MI, 2016. Return to text.
- Morell, M., Vogl, A.W., IJsseldijk, L.L., Piscitelli-Doshkov, M., Tong, L., Ostertag, S., Ferreira, M., Fraija-Fernandez, N., Colegrove, K.M., Puel, J-L., Raverty, S.A., and Shadwick, R.E., Echolocating whales and bats express the motor protein prestin in the inner ear: a potential marker for hearing loss, Frontiers in Veterinary Science 7(429):1–14, 2020. Return to text.
- Weichenrieder, O., Wild, K., Strub, K., and Cusack, S., Structure and assembly of the Alu domain of the mammalian signal recognition particle, Nature 408(6809):167–173, 2000. Return to text.
- Panchal, M., Rawat, K., Kumar, G., Kibria, K. M., Singh, S., Kalamuddin, M., et al., Plasmodium falciparum signal recognition particle components and anti-parasitic effect of ivermectin in blocking nucleo-cytoplasmic shuttling of SRP, Cell death & disease 5(1):e994, 2014. Return to text.
- Pool, M.R., Signal recognition particles in chloroplasts, bacteria, yeast and mammals (review), Molecular membrane biology 22(1–2):3–15, 2005. Return to text.
- Bousset, L., Mary, C., Brooks, M. A., Scherrer, A., Strub, K., and Cusack, S., Crystal structure of a signal recognition particle Alu domain in the elongation arrest conformation, RNA 20(12):1955–1962, 2014. Return to text.
- Mason, N., Ciufo, L.F., and Brown, J.D., Elongation arrest is a physiologically important function of signal recognition particle, The EMBO J. 19(15):4164–4174, 2000. Return to text.
- Strub, K., Fornallaz, M., and Bui, N., The Alu domain homolog of the yeast signal recognition particle consists of an Srp14p homodimer and a yeast-specific RNA structure, RNA 5(10):1333–1347, 1999. Return to text.
- Nakamura, K., Yahagi, S., Yamazaki, T., and Yamane, K., Bacillus subtilis histone-like protein, HBsu, is an integral component of a SRP-like particle that can bind the Alu domain of small cytoplasmic RNA, J. Biological Chemistry 274(19):13569–13576, 1999. Return to text.
- O’Micks, J., Molecular structures shared by prokaryotes and eukaryotes show signs of only analogy and not homology, ARJ 9:284–292, 2016. Return to text.
- Bovia, F., Wolff, N., Ryser, S., and Strub, K., The SRP9/14 subunit of the human signal recognition particle binds to a variety of Alu-like RNAs and with higher affinity than its mouse homolog, Nucleic Acids Research 25(2):318–326, 1997. Return to text.
- Garnier, J., Osguthorpe, D.J., and Robson, B., Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins, J. Mol. Biol. 120(1):97–120, 1978. Return to text.
- Eddy, S.R., A new generation of homology search tools based on probabilistic inference, Genome Informatics 23(1):205–211, 2019. Return to text.
- Prakash, A., Jeffryes, M., Bateman, A., and Finn, R.D., The HMMER Web Server for Protein Sequence Similarity Search, Current Protocols in Bioinformatics 60:3.15.1–3.15.23, 2017. Return to text.










Readers’ comments
Comments are automatically closed 14 days after publication.