Journal of Creation 33(2):107–112, August 2019
Browse our latest digital issue Subscribe
The green algae Chlamydomonas reinhardtii find safety in numbers by design
The development of multicellularity is one of the most significant steps during the purported evolutionary history of life. Multicellularity would theoretically allow for the further differentiation of cell types. This would allow for greater variability and facilitate evolutionary change. A study has been performed on the green algae Chlamydomonas reinhardtii that demonstrated a rapid transition from a unicellular to a para-multicellular phenotype due to predatory pressure. It can be demonstrated that evolution has nothing to do with this kind of transition. Rather, a pre-existing genetic mechanism had already been in place to adapt the algae to environmental cues, such as predation. The normally unicellular Chlamydomonas reinhardtii as well as the obligately multicellular Volvox carteri are both members of the same baramin. A preliminary analysis indicates that a genetic basis exists for the transition between unicellularity and multicellularity within this baramin.

One of the most critical steps during evolutionary history was the supposed transition from unicellular to multicellular life forms. Both unicellular and multicellular forms have been observed in algal species such as Chlamydomonas reinhardtii and Chlorella vulgaris. Because of this, these and related species have been the focus of intense evolutionary experimentation. Some evolutionists theorize that a transition to multicellularity was driven by predation. Most predators can consume prey only within a certain size range, and prey species larger than a given threshold are immune to such predation.
However, there is no fossil evidence that this transition happened. The earliest multicellular life forms are already fully formed. There is no nascent multicellular stage evident in the rocks. Evolutionists admit that surprisingly little is known about the origin of multicellularity,1 hence the interest in a unicellular algal species that demonstrates the ability to suddenly produce clumps of cells. This has given some evolutionists confidence, but a true transition to multicellularity would necessitate the evolution of a multitude of new genes, including ones that code for proteins involved in an extracellular matrix (ECM), cell-to-cell communication, cellular differentiation and specialization, and cellular cohesion, not to mention genes which alter the reproductive cycle.
But could it be that, instead of evolving multicellularity, algal species find safety in numbers by design?
Predation tests with C. reinhardtii strains
Herron et al.1 used ciliate predator species Paramecium tetraurelia to select for populations of C. reinhardtii which seemed to have evolved multicellularity. In their experiments, the C. reinhardtii populations were cultured for 50 weeks, representing about 750 generations. The strains were put into 24-well tissue culture plates and were observed with time-lapse videos. The resulting strains were classified based on the sizes of the parent and propagule clusters.
They began the experiment with a mother culture derived from crossing two different algal strains obtained from the Chlamydomonas Resource Center. They divided this initial culture into five experimental (B1–B5) and three control (K1–K3) ‘populations’. Within each population were multiple ‘strains’, each apparently grown in a separate well of a 24-well culture plate. A ‘strain’ is equal to all genetically identical descendants of an individual haploid cell isolated from a given population (i.e. B2-01 denotes strain #1 from population B2).
Table 1. Characterization of different categories of strains of C. reinhardtii in the Herron et al. experiment
| A | B | C | D | |
| Strains | K1-01, K1-06, B2-10, B5-06 | B2-01 | B5-05 | B2-03, B2-04, B2-11 |
| Predator present | X | √ | √ | √ |
| Multicellular | √ | √ | √ | √ |
| Extracellular matrix | X | √ | √ | √ |
| Multicellular propagules | X | X | √ | √ |
Yet, they do not describe their culture protocol in depth. For instance, how many replicates did they culture? Did they avoid pseudoreplication by culturing all eight populations, in triplicate, and in random positions, on each plate or did they fill each of the 24 wells with cells from the same population? They also say: “During transfers, populations were homogenized with a multichannel pipette …”. This means that all subcultures of each population were pooled, but this makes no sense. Why would they use a multichannel pipette, for instance, if they were pooling each subsample of each population? We can only assume that they mixed their samples randomly. The worst thing they could have done would be to create eight 24-well plates, with one population per plate, and then stack them in order in the incubator. This would give them an ‘n’ of 1 per population (because all 24 wells on each plate would be causally connected) and would introduce the possibility of uncontrolled variables like light or temperature differences across the stack. We can only assume they were aware of such things. However, it is impossible to reproduce their experiment given the description in their Methods section.
Table 2. Proteome characteristics of the nine species used in this analysis
| Species | Uniprot ID | no. proteins | no. mapped proteins |
| Chlorella sorokiniana | UP000239899 | 10201 | 4746 |
| Chlorella variabilis | UP000008141 | 9831 | 5050 |
| Chlamydomonas eustigmata | UP000232323 | 14139 | 6243 |
| Chlamydomonas reinhardtii | UP000006906 | 18828 | 7926 |
| Gonium pectorale | UP000075714 | 16224 | 6815 |
| Micromonas commoda | UP000002009 | 10115 | 6893 |
| Micromonas pusilla | UP000001876 | 10250 | 5786 |
| Saccharomyces cerevisiae (outlier) | UP000002311 | 6049 | 4537 |
| Volvox carteri f. nagariensis | UP000001058 | 14335 | 8181 |
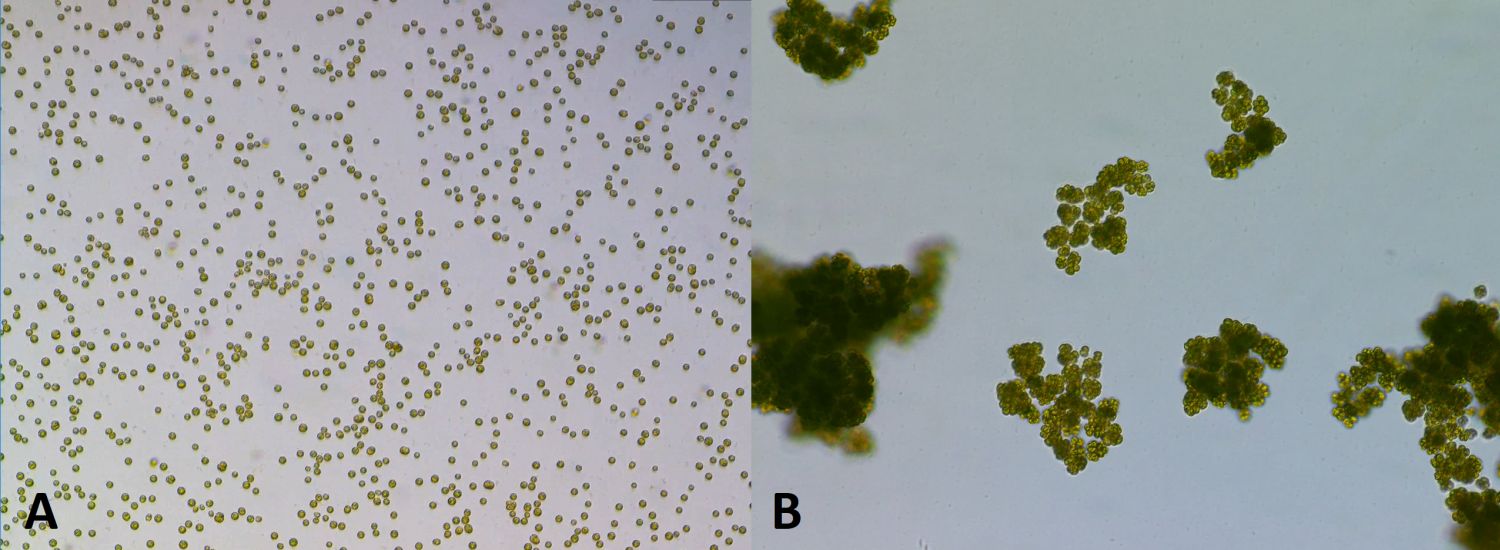
Para-multicellular structures appeared in two of the five predator-selected populations and one of the control populations. Seven strains were selected from the experimental populations (B2-01, B2-03, B2-04, B2-10, B2-11, B5-05, and B5-06) and two strains were selected from the control population (K1-01 and K1-06). They did not explain why they used five from the B2 population, but only two from the B5 and K1 populations. These nine strains were put into four different categories (A–D) based on a variety of lifestyle characteristics, such as the presence of unicellular to various multicellular forms, or the number and kind of propagules (daughter cells/cell clusters) that were released from the cell clusters, etc. These four categories are described in table 1. Figure 1 depicts samples from supplementary video 3 and 8 of Herron et al. showing cells in unicellular (figure 1A), and multicellular form (figure 1B).
The strains from categories B, C, and D, also formed an extracellular matrix (ECM). As cells grew and divided, some stayed embedded in the ECM, whereas sometimes the colony released propagules (either individual cells or groups of cells which broke away from the main colony). Category C was similar to category B, but with larger-sized colonies. As opposed to categories B and C, which released only single-celled propagules, category D released multicellular propagules.
Certain considerations strongly suggest that the unicellular algae such as C. reinhardtii and several multicellular algal species belong to the same created kind, or baramin. A baramin is a group of species which show continuity with one another and discontinuity with all other species. Member species generally form a reproductive continuum with one another. For example, Prochnik et al.2 have sequenced the genomes of both C. reinhardtii and Volvox carteri. They found that the overall genome size, the number of protein-coding genes, the number of different kinds of protein domains, and the distribution of gene family sizes are both very much the same.3 Hence, these two cellular forms are likely only phenotypically different expressions of the same baranome (the ‘baranome’ being the initial, created genome from which the genomes of all other species within this ‘kind’ derive).
Materials and methods
The Gene Content Method (GCM)4 analyzes the whole proteomes of the species under study and cross-compares them with one another. The species included in the current analysis include C. reinhardtii, Chlamydomonas eustigmata, Chlorella sorokiniana, Chlorella variabilis, V. carteri, Micromonas commoda, and Micromonas pusilla. Saccharomyces cerevisiae was used as an outgroup. The species, their Uniprot ID, the number of proteins in their proteome, and the number of mapped proteins can be seen in table 2. For each species, each protein is assigned to an orthology group (if possible) by the OrthoMCL algorithm.5 A pairwise overlapping percentage is then calculated for each pair of species. Clusters of species are calculated using the k-means algorithm and are visualized on a heat map.
The algal and yeast proteomes were downloaded from the Uniport Database at uniprot.org. The OrthoMCL pipeline for each proteome was run at the EuPathDB Galaxy website at eupathdb.globusgenomics.org to map the proteins to OrthoMCL groups. Jaccard Coefficient Values were calculated according to the GCM method. R (version 3.5.2) was used to create the heat map between the five algae species using the heatmap.2 function.
Table 3. JCV statistics for the three putative baramins analyzed in this study
| Cluster | No. species | Min JCV | Max JCV | St.dev. | p-value |
| Volvox | 4 | 0.572 | 0.722 | 0.058 | 1.24 x 10-4 |
| Chlorella | 2 | 0.697 | 0.697 | NA | 2.10 x 10-8 |
| Micromonas | 2 | 0.750 | 0.750 | NA | 2.30 x 10-9 |
Results from the baraminological status of C. reinhardtii
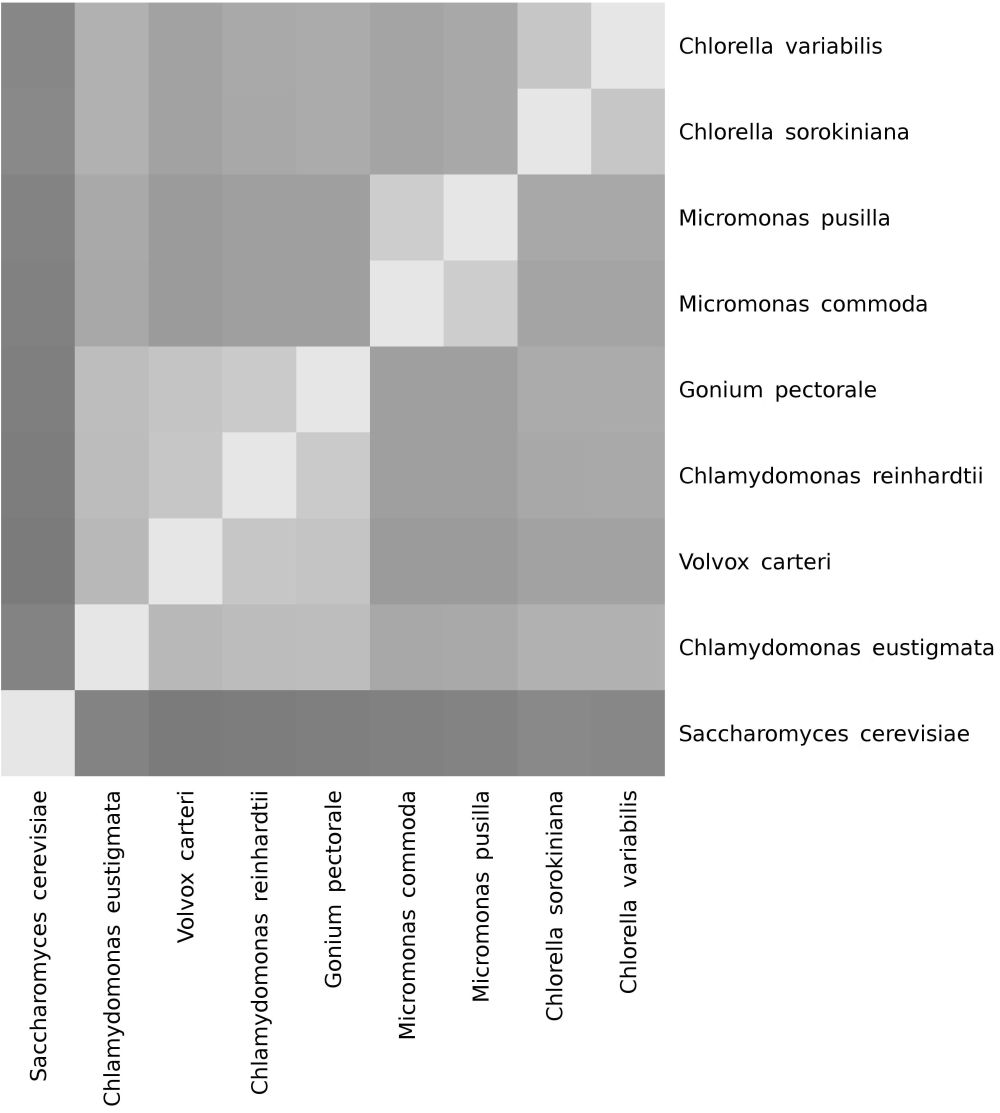
The GCM software was run by the author to fit three baramins (figure 2). Indeed, the two Chlamydomonas species cluster together with V. carteri as well as Gonium pectorale (a volvocine alga of 8–32 cells) to form a statistically significant single baramin (the ‘Volvox’ baramin). This baramin has a mean Jaccard Coefficient Value (JCV) of 0.648 and a range of 0.572 to 0.722. JCV is defined as the overlap of common genes divided by the union of such genes,6 with higher numbers representing more significant similarities between species. For perspective, the JCVs between this group and all other species is 0.188 to 0.527 (p = 1.2 x 10-4, Student’s t-test between JCV values between species within the Volvox kind versus JCV values between Volvox and all other species).
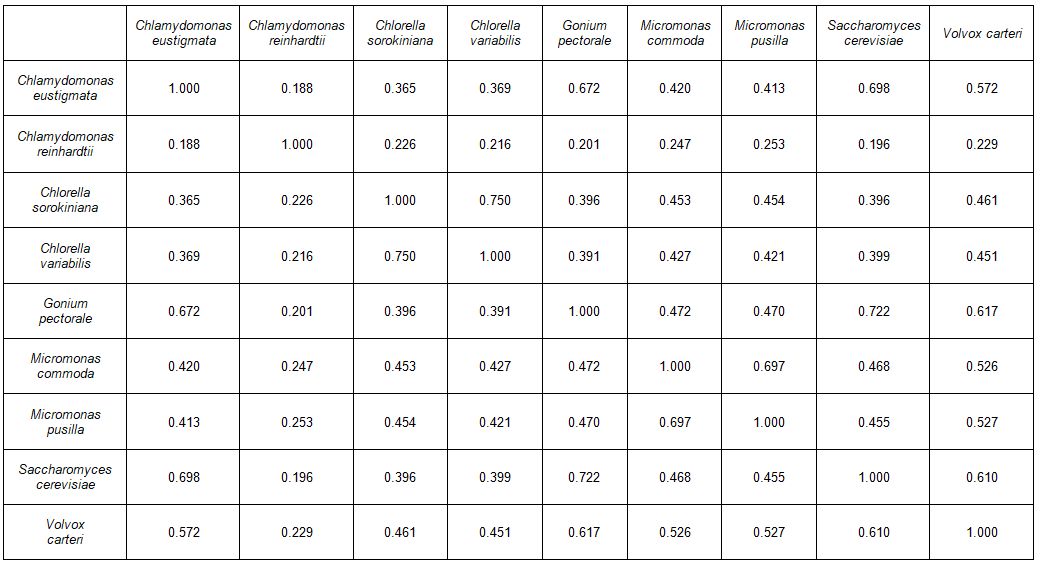
The two Micromonas species also cluster together to form a statistically significant putative baramin (p = 2.3 x 10-9), with a JCV of 0.750, which is the highest JCV between any two pairs of species in the study. The two Chlorella species have a JCV of 0.697 (p = 2.1 x 10-8) (table 3). The JCV matrix can be seen in table 4. Figure 3 depicts JCV values within a given baramin, between a given baramin and all other species, and between members of a given baramin and the outlier species. It is easy to see that the within-baramin JCV values separate well from the among-baramin JCV values and the outlier JCV values.
Discussion
Herron et al. speculate that multicellular phenotypes have a genetic basis involving the co-option of a previously existing plastic response.1 Therefore, the shift from a unicellular to a multicellular lifestyle may involve a shift to the expression of genes involved in multicellular life forms. This means that the genetic machinery responsible for multicellular life is already in existence. No explanation of any kind was offered to how these genes themselves supposedly came into existence via evolution. Thus, the only thing that happened was a phenotypic change due to differential expression of genes specific to a sort of pseudo-multicellularity. We do not yet know what genes were allegedly involved.
In fact, the time frame that was made available for the Chlamydomonas strains to evolve new multicellularity-specific structures was only 750 generations. This happened despite claims by evolutionists that the gene content of C. reinhardtii and V. carteri have stayed the same for ~310 million years.2 Since the transition happened in parallel in multiple cultures, it happened much too rapidly for any kind of genetic mutation to cause the differences in phenotype. Rather, this supports the idea that multicellularity-specific genes serve as design elements which are differentially expressed according to different external conditions (e.g. the absence or presence of predators), helping the algae to adapt to such conditions. Yet, the fact that the transition occurred in one of the control populations indicates that we have a lot more to learn about the environmental triggers that control the life history strategies within C. reinhardtii.
In a related experiment, Herron et al. studied transcription in a unicellular strain of C. reinhardtii and a multicellular strain over a characteristic life cycle spanning 72 hours. Approximately 20% (3,453) of all C. reinhardtii genes were found to be differentially expressed by fourfold or greater in at least one time point. The ontology of the over-represented genes is related to cell cycle and reproductive processes.7
Brueggeman et al. showed that roughly 32% of all C. reinhardtii genes were significantly differentially expressed (> 1 absolute log2-fold change) when transitioning from high to very low (≤ 100 ppm) levels of CO2.8 These genes were part of a CO2-concentrating mechanism which accumulates the CO2 necessary for photosynthesis. Thus, we know that the species already contains an elaborate system that controls a massive metabolic change. It only makes sense that the phenotypic changes they report also involve predetermined pathways. This is evidenced by the fact that it arose independently in multiple cultures and in a very short space of time. In fact, since the predator and the prey would be expected to interact naturally and frequently in the wild, one might expect that what was discovered is only a pre-engineered system to regulate the populations of both. Phenotypic switches are caused by the interplay between hundreds of genes. For example, in yeast, the cytotoxin CuO induces the differential expression of some 654 genes, shifting metabolism from alcoholic fermentation to aerobic respiration.9
A facultative multicellular phenotype is not rare and occurs not just in algae but also in bacteria and yeasts, such as Saccharomyces cerevisiae.10 Herron and co-workers have documented it in at least 25 separate lineages across all life forms.11 The supposed evolution of multicellularity is thus all the more unlikely, since it must have evolved more than once, according to the principle that evolutionary trajectories do not repeat themselves.12 The freshwater bacterial strain Flectobacillus displays both a filamentous phenotype when in the presence of protist predators, as well as a rod-like shape during carbon deprivation (low CO2 concentrations). Similar rapid phenotypic changes by some bacterial species is based on the adaptation of underlying genetic mechanisms.13
Table 4. JCV matrix for nine species under study
It may be possible to expand the Volvox baramin to include species from the genus Vitreochlamys, as well as the colonial volvocine groups Tetrabaena, Goniaceae, and other species from Volvocaceae, the latter three of which have been shown to be monophyletic by a number of studies.14,15 It is also possible that the unicellular algae are derived from multicellular forms. This implies devolution and not evolution, which is a much easier process to explain,16 like how parasitic bacteria devolved from wild species through genomic reduction.17
Even though the genomes of C. reinhardtii and Volvox carteri are almost identical, there are some genes which are active in the Volvox developmental processes which are not active in the developmental pathway of Chlamydomonas, although the Chlamydomonas gene versions can rescue Volvox mutants. An example of such a gene is glsA, which is responsible for asymmetric division leading to colony inversion.18 Furthermore, two gene families which are responsible for the production of the ECM have more members in Volvox than in Chlamydomonas.5

According to evolutionary theory, when species reach a new level of complexity, the new species is greater than the sum of the previous levels. Each new level of complexity is built upon, but cannot be explained merely in terms of the previous levels alone. These experiments do not prove evolution, which requires the augmentation of a prior genetic system to fit a new need. This would involve the great quantity of new information necessary to coordinate newly founded relationships between individual cells. New genes with entirely new functions would be necessary to coordinate individual cells with each other in a multicellular system.19 This did not happen here with Chlamydomonas and Volvox. Chlamydomonas cells indeed form an ECM, but afterwards individual cells, or clumps of cells, break off after reaching a pseudo-multicellular state. But could this be considered an incipient, transitional stage on the way to multicellularity? The genes for ECM formation are present in both Chlamydomonas and Volvox, showing that no genetic changes accompanied this phenotypic change between the two species. Herron et al. claim that C. reinhardtii has no multicellular ancestors, and that they discovered a completely new origin of obligate multicellularity. Yet it might be the other way around, namely that unicellular algae are descended from multicellular forms, with a diminished capacity of forming an ECM.
Summary and conclusion
With this experiment, evolutionists have shown absolutely no evidence for the evolution of multicellularity. Neither have they shown the transition of unicellular life to multicellular life. Rather they have uncovered what appears to be a pre-existing, complex, designed genetic system that is, perhaps, responsible for phenotypic adaptation against predators. In fact, there is some evidence that the unicellular volvocine algae are actually derived from multicellular forms via devolution, and not evolution. Furthermore, three preliminary algal holobaramins were predicted, that of the genera Chlorella, Micromonas, and Volvox/Chlamydomonas. The Volvox baramin displays a dual unicellular/multicellular phenotype.
Acknowledgements
I would like to thank Dr Robert Carter for his critical analysis of the paper and also his helpful comments and suggestions.
References
- Herron, M.D., Borin, J.M., Boswell, J.C. et al., De novo origins of multicellularity in response to predation, Sci. Rep. 9(1):2328, 2019 | doi:10.1038/s41598-019-39558-8. Return to text
- Prochnik, S.E., Umen, J., Nedelcu, A.M. et al., Genomic analysis of organismal complexity in the multicellular green alga Volvox carteri, Science 329(5988):223–226, 2010 | doi:10.1126/science.1188800. Return to text
- Miller, S.M., Volvox, Chlamydomonas, and the evolution of multicellularity, Nature Education 3(9):65, 2010. Return to text
- O’Micks, J., Baraminology classification based on gene content similarity measurement, CRSQ 54:27–37, 2017. Return to text
- Fischer, S., Brunk, B.P., Chen, F. et al., Using OrthoMCL to assign proteins to OrthoMCL-DB groups or to cluster proteomes into new ortholog groups, Curr. Protoc. Bioinformatics, chap. 6, unit 6, 12:1–19, 2011 | doi: 10.1002/0471250953.bi0612s35. Return to text
- Jaccard, P., The distribution of the flora of the alpine zone, New Phytologist 11:37–50, 1912. Return to text
- Herron, M.D., Ratcliff, W.C., Boswell, J., and Rosenzweig F., Genetics of a de novo origin of undifferentiated multicellularity, R. Soc. Open Sci. 5(8):180912, 2018 | doi: 10.1098/rsos.180912. Return to text
- Brueggeman, A., Gangadharaiah, D.S., Cserhati, M.F. et al., Activation of the carbon concentrating mechanism by CO2 deprivation coincides with massive transcriptional restructuring in Chlamydomonas reinhardtii, Plant Cell 24(5):1860–1875, 2012 | doi: 10.1105/tpc.111.093435. Return to text
- Tan, J., He, Q., Pentz, J.T., Peng, C. et al., Copper oxide nanoparticles promote the evolution of multicellularity in yeast, Nanotoxicology 7:1–9, 2019 | doi: 10.1080/17435390.2018.1553253. Return to text
- Grosberg, R.K. and Strathmann, R.R., The evolution of multicellularity: a minor major transition? Annu. Rev. Ecol. Evol. Syst. 38:621–654, 2007. Return to text
- Herron, M.D., Rashidi, A., Shelton, D.E., and Driscoll, W.W., Cellular differentiation and individuality in the ‘minor’ multicellular taxa, Biol. Rev. Camb. Philos. Soc. 88(4):844–861, 2013 | doi: 10.1111/brv.12031. Return to text
- Gould, S.J. Wonderful Life: The Burgess Shale and the nature of history, W.W. Norton & Company, New York, 1990. Return to text
- Corno, G. and Jürgens, K., Direct and indirect effects of protist predation on population size structure of a bacterial strain with high phenotypic plasticity, Appl. Environ. Microbiol. 72(1):78–86, 2006. Return to text
- Arakaki, Y., Kawai-Toyooka, H., Hamamura, Y. et al., The simplest integrated multicellular organism unveiled, PLoS One 8(12):e81641, 2013 | doi:10.1371/journal.pone.0081641. Return to text
- Herron, M.D., Hackett, J.D., Aylward, F.O., and Michod, R.E., Triassic origin and early radiation of multicellular volvocine algae, PNAS 106(9):3254–3258, 2009 | doi: 10.1073/pnas.0811205106. Return to text
- Herron, M.D., Origins of multicellular complexity: Volvox and the volvocine algae, Mol. Ecol. 25(6):1213–1223, 2016 | doi: 10.1111/mec.13551. Return to text
- O’Micks, J., Bacterial genome decay from a baraminological viewpoint, J. Creation 29(2):122–130, 2015. Return to text
- Cheng, Q., Fowler, R., Tam, L.W. et al., The role of glsA in the evolution of asymmetric cell division in the green alga Volvox carteri, Dev. Genes Evol. 213(7):328–335, 2003. Return to text
- Doyle, S., Evolution of multicellularity: what is required? J. Creation 23(1):5–7, 2009. Return to text




Readers’ comments
Comments are automatically closed 14 days after publication.