Journal of Creation 18(2):37–40, August 2004
Browse our latest digital issue Subscribe
Human/chimp DNA similarity continues to decrease: counting indels
Summary
It is conventionally held that humans and chimps differ only very slightly in their DNA. However, new evidence suggests that the difference might be much more drastic. Mutations resulting in DNA insertions and deletions cause much of the genetic difference between the two species, but are typically not included in estimates of diversity. Moreover, areas of significant similarity are often affected by selective constraints. An increasing number of functions are also being discovered for so-called ‘junk DNA’, suggesting similarity in such DNA is not necessarily due to common descent. Additional research should aid the understanding of such important data in the debate over origins.
Creationists have long maintained that the similarity between human and chimp DNA is not all that it is touted to be. A new study in the Proceedings of the National Academy of Sciences could help confirm this.
It is widely held that ‘The common chimpanzee (Pan troglodytes) is our closest relative. Its genome sequence is about 98.8% identical to our own, and we shared a common ancestor some six million years ago.’1 The assumption that humans diverged from chimps roughly this long ago also forms the basis of the mitochondrial clock,2 which ‘continues to be widely used to “time” human evolution and population movements, both ancient and modern.’3 In the popular-level book Genome, Matt Ridley states that:
‘Apart from the fusion of chromosome 2, visible differences between chimp and human chromosomes are few and tiny. In thirteen chromosomes no visible differences of any kind exist. If you select at random any “paragraph” in the chimp genome and compare it with the comparable “paragraph” in the human genome, you will find very few “letters” are different: on average, less than two in every hundred. We are, to a ninety-eight per cent approximation, chimpanzees, and they are, with ninety-eight per cent confidence limits, human beings. If that does not dent your self-esteem, consider that chimpanzees are only ninety-seven per cent gorillas; and humans are also ninety-seven per cent gorillas. In other words we are more chimpanzee-like than gorillas are.’4
One creationist response to such arguments regarding human/chimp DNA similarity has been that ‘Chimp DNA has not been anywhere near fully sequenced so that a proper comparison can be made’,5 and that this evidence is just as easily explained (and predicted, for that matter) by the concept of a common designer:
‘Since DNA codes for structures and biochemical molecules, we should expect the most similar creatures to have the most similar DNA. Apes and humans are both mammals, with similar shapes, so both have similar DNA. We should expect humans to have more DNA similarities with another mammal like a pig than with a reptile like a rattlesnake. And this is so. Humans are very different from yeast but they have some biochemistry in common, so we should expect human DNA to differ more from yeast DNA than from ape DNA.’6
In a recent article,7 David A. DeWitt cited a study which found that the two species are only 95% identical when insertions and deletions are considered,8 showing that the estimate of divergence depends mainly on what type of DNA is being compared. A number of differences between humans and chimps were named that are difficult to quantify in an estimate of sequence divergence (that is, the differences in bases between the human and chimp genomes), including shorter telomeres in humans, a 10% larger chimp genome, and great differences in chromosomes 4, 9, 12 and the Y chromosome, for example. Indeed, DNA similarity estimates ‘do not adequately represent fine changes in genome organization.’9
Considering DNA gaps
Previous estimates of sequence divergence have focused exclusively on base substitutions in DNA—that is, one base (or one DNA ‘letter’—A, T, C or G) being replaced with another. The new calculation, resulting in much less sequence similarity, also includes insertions and deletions, or indels, (occurring when a base is added or removed, often resulting in what is known as a frameshift mutation), in addition to base substitutions. The author of the study, Roy J. Britten, stated:
‘It appears appropriate to me to consider the full length of the gaps in estimating the interspecies divergence. These stretches of DNA are actually absent from one and present in the other genome. In the past, indels have often simply been counted regardless of length and added to the base substitution count, because that is convenient for phylogenetics.’8
His findings lend support to the idea that much of the failure of DNA to hybridize between chimps and humans is the result of missing DNA due to indel events. Britten then became involved in a follow-up paper in which these initial results were confirmed; in fact, it was found that ‘the 5% human-chimp difference already published is likely to be an underestimate, possibly by more than a factor of 2.’10
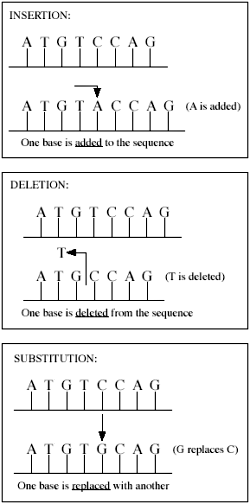
Various types of mutations. Much of the difference between human and chimp DNA can be attributed to insertions and deletions (indels).
Now, Anzai et al. have published a new report in the Proceedings of the National Academy of Sciences that confirms this statement. In the study, nearly one-half of the MHC (major histocompatibility complex) region was sequenced, ‘which to date represents the longest continuous sequence within this species [chimps], our closest evolutionary relative’, and has been described as a ‘rapidly evolving’ part of the genome.9 Although it has been held that human/chimp similarity in the MHC is ‘so great that the alleles must have originated before the supposed chimp/human evolutionary divergence’,11 the sequence results actually dropped the DNA similarity estimate down to 86.7%!12 Indeed, the actual difference between the two species (when counting indels) is greater than 5% by well more than a factor of two. Not only this, but ‘evolutionists now recognize that complex MHC genetic motifs can arise independently’ in primates—that is, at least some similarities that do exist are not attributable to common descent.13
The human genome contains two MHC Class I genes, the MICA and MICB, yet chimpanzees contain only one gene at this location, the Patr-MIC. According to evolutionary speculation, a 95-kb deletion occurred between the two human genes, forming the hybrid chimpanzee gene ~33–44 million years ago, by far predating the commonly held divergence date between the two species of 6 million years. Because the two ends of the chimpanzee gene seem to match up with the beginning of the human MICA and end of the human MICB genes, it may seem reasonable that common ancestry is feasible. However, even some humans contain a single gene at this location (called the HLA-B*4801 allele) very similar to the one found in chimps. The study notes that it ‘is quite intriguing that an equal-sized deletion involving this very same region and genes (MICA/B) has happened at distinct points in time in several different primate species’.12 Yet it is also claimed that other such similar changes in DNA structure cannot be attributed to convergence, but must be due to common ancestry! Clearly, similar ‘mistakes’ can arise independently in separate species (as expanded upon by Woodmorappe13). The hypothesis that a Designer would create the same structures for the same functions seems to explain the data much more easily. As noted by Woodmorappe,11 strong selective pressures must have existed in order to prevent the MHC similarities between primates from being scrambled over supposed millions of years, further weakening the evolutionary scenario.
The Anzai et al. study also mentions a number of differences between humans and chimps that may be a result of genetic changes in the MHC genes, including the difference in handling infectious agents such as HIV, hepatitis B and C, and susceptibility to Plasmodium falciparum. Therefore, the differences observed in these genes may portray the believed ‘true’ divergence between the two species much better than previous estimates.
Although these results are interesting, there has been debate over whether or not indels should be included in sequence divergence estimates. For example, a mutation called a translocation can occur, in which a segment of DNA breaks off from one chromosome and is inserted in another. The original Britten study discussed such rearrangement events briefly and found them to be frequent. Due to the fact that indel differences were defined as ‘the full length of the gaps’ in the genomes, the estimates would not be able to consider this kind of mutational change easily.14 New research will hopefully aid in the understanding of changes in genome organization, and give clues as to how these changes can be included in estimates of human/chimp similarity.
Difference between coding and noncoding DNA
Other studies have resulted in estimates of similarity higher than 98.6%, also. For instance, Wildman et al.15 compared ~90 kilobases of human DNA to chimps and found a similarity of 98.86%, even when counting indels. This is important evidence, considering that it is in direct opposition to the data presented by Britten and Anzai et al. However, it must be understood that the various estimates use different types of DNA. Wildman’s team examined only coding DNA from a number of genes. Here, non-synonymous changes (those affecting protein structure by changing the specific amino acid encoded) are subject to purifying selection. This means that they can be selected against if they have any effect on the function of the protein.
Similarly, a study of human chromosome 21 (the smallest chromosome in the human genome) found only 3,003 nucleotide differences in over 400 kilobases. It was shown that: ‘The differences in coding, promoter, and exon-intron junction regions were 0.51 ± 0.02%, 0.88 ± 0.03%, and 0.85 ± 0.02%, respectively, much lower than the previously reported 1.23% in genomic regions’,16 with an overall similarity of 99.3%. Within an evolutionary framework, these results would confirm chimps as our closest relatives. However, this finding seems to contradict the knowledge of a high substitution rate on chromosome 21, also leading to the conclusion
‘ … that the higher level of similarity observed in the transcript units in this study is attributable to the presence of purifying natural selection exerted on the most important functional portions of the genes, including promoters, coding regions, and intronic regions near the exon-intron boundary.’16
Therefore, high similarity estimates specifically involve regions of coding DNA that are functionally constrained. The studies by Britten et al. and Anzai et al. both consider non-coding DNA, which might be less constrained, and therefore more free to accumulate random mutations. This non-coding DNA thus serves as a more accurate portrayal of true divergence. Of course, it is very reasonable within the context of biblical creation that the most similarity should exist where protein function is vital, since the same proteins would be used for the same structures by a common Designer.6 It naturally follows that non-coding DNA, being less constrained, possibly contains more divergence.
Returning to the Anzai et al. study, which found chimps and humans to be 86.7% similar, a general trend may be noticed with higher similarity in coding regions. Whereas most ‘non-MHC genes are involved in basic (homeostatic) cellular functions that require interindividual as well as interspecies homogeneity’, the MHC genes ‘have to constantly adapt themselves to the microbiological habitat of every species.’ Therefore, purifying selection tends to maintain the structural conservation of non-MHC genes because of their specific functions. We can conclude that the 86.7% estimate ‘may be a better representation of whole-genome sequence similarity between the human and the chimpanzee’ than previous estimates of 98.6%. Since ‘the major difference between the human and chimpanzee sequences is overwhelmingly attributable to indels’,12 estimates not including these mutations ignore a huge source of potential differences. Recent studies have consistently found indels to be the main source of variation between humans and chimps.8,10,12 It should also be noted, in contrast to examples of high-sequence similarity, that sequence divergence in certain regions can exceed 20%.8 As noted by DeWitt, estimates can be ‘misleading because it depends on what is being compared.’7
Junk DNA
Introns are regions of DNA in the genome that do not code for a protein product, and are therefore assumed to have no function. Because of this, ‘introns in a particular gene are often compared between organisms, with the base pair differences seen between their sequences supposedly indicating the degree and time of divergence since they last shared a common ancestor.’17 Indeed, functionless introns should be very different in humans and chimps, or even nonexistent, within the context of biblical creation. However, evidence is mounting that introns are not, after all, void of function, and the assumption that they were may ‘come to be a classic story of orthodoxy derailing objective analysis of the facts.’18 Other forms of ‘junk’ DNA, obviously said to lack function and thus able to mutate at random, actually contradict evolutionary phylogenies, such as pseudogenes shared by humans and gorillas but not chimps, the CYP pseudogene being present only in chimps, and a substitution in the Alpha-1,3GT pseudogene shared by cows, squirrel monkeys and gorillas. Many substitutions that are shared take place in a non-random manner, also weakening the explanatory power of common descent.13 Numerous articles have been published discussing the functions of various alleged forms of ‘junk’ DNA,13,17,19–22 and it is encouraging to actually see evolutionary journals awakening to this important fact. The preservation of introns
‘… suggests they do something indispensable. And indeed a large number are transcribed into varieties of RNA that perform a much wider range of functions than biologists had imagined possible. Some scientists now suspect that much of what makes one person, and one species, different from the next are variations in the gems hidden within our “junk” DNA.’23
Similarities in introns do, therefore, fit the creationist paradigm quite nicely.
DNA is not everything
I suggest that further research is required in order to sort through this evidence, research that will also find differences inherent within the chimp kind. Indels can easily be viewed as intrinsic differences between kinds. The DNA sequence is not all that distinguishes different kinds of organisms—as geneticist Steve Jones was quoted in Creation as saying, ‘We also share about 50% of our DNA with bananas and that doesn’t make us half bananas, either from the waist up or the waist down.’24 Evidence has certainly emerged that ‘DNA is not everything’; for example, mitochondria, ribosomes, the endoplasmic reticulum and the cytosol are passed unchanged from parent to offspring (save for possible mutations in mtDNA). In fact, gene expression is itself under the control of the cell.25 Some animals have undergone extremely dramatic genetic changes, and yet their phenotype has remained virtually identical.26 Such epigenetic marks ‘can dramatically affect the health and characteristics of an organism—some are even passed from parent to child—yet they do not alter the underlying DNA sequence.’27 This evidence lends great support to reproduction after kinds (Genesis 1:24–25; 1 Corinthians 15:39), as structures present within parents are preserved in their offspring.
Conclusion
This is an exciting time for creationists as estimates of human/chimp similarity continue to decrease when indels are considered. Although it is obvious that the two species are very much alike in the mere DNA sequences (many of the same structures are present in both, so this would be expected in a creation model), the previous estimate of ~98.6% sequence identity may have been dealt a significant blow. Upcoming research will likely shed new light on the many differences between humans and other animals, and continue to affirm the truth of Genesis.
Acknowledgements
I wish to thank Kim Risely for reviewing an earlier draft of this manuscript. Also, I recognize Reed Cartwright, David DeWitt and Carl Wieland for providing helpful information and criticism. I am also indebted to Randy Kim, for having supported, encouraged and taught me at the times when it was most needed.
About the author
J. Warren Nelson is a high school sophomore with a particular interest in biology as it relates to evolutionary theory. He has attended a number of Creation conferences, including Creation Conference 2003. He often gives talks at his school’s Fellowship of Christian Athletes, on topics ranging from Genesis to Christian love. His current research interests involve population genetics, chemical evolution and Christian friendship. Return to top.
References
- Cyranoski, D., Almost human …, Nature 418(6901):910–912, 2002.
- Gibbons, A., Calibrating the mitochondrial clock, Science 279(5347):28–29, 1998.
- Howell, N., Smejkal, C.B., Mackey, D.A., Chinnery, P.F., Turnbull, D.M. and Herrnstadt, C., The pedigree rate of sequence divergence in the human mitochondrial genome: there is a difference between phylogenetic and pedigree rates, American J. Human Genetics 72(3):659–670, 2003.
- Ridley, M., Genome, HarperCollins Publishers, New York, p. 28, 1999.
- Batten, D., Human/chimp DNA similarity: Evidence for evolutionary relationship? Creation 19(1):21–22, 1997.
- Sarfati, J., Refuting Evolution 2, Creation Ministries International, Brisbane, Australia, pp. 112–113, 2002.
- DeWitt, D.A., >98% Chimp/Human DNA similarity? Not any more, TJ 17(1):8–10, 2003.
- Britten, R.J., Divergence between samples of chimpanzee and human DNA sequences is 5%, counting indels, Proc. Nat. Acad. Sci. USA 99(21):13633–13635, 2002.
- Hacia, J.G., Genome of the apes, Trends in Genetics 17(11):637–645, 2001.
- Britten, R.J., Rowen, L., Williams, J. and Cameron, R.A., Majority of divergence between closely related DNA samples is due to indels, Proc. Nat. Acad. Sci. USA 100(8):4661–4665, 2003.
- Woodmorappe, J., Noah’s Ark: A Feasibility Study, Institute for Creation Research, El Cajon, p. 203, 1996.
- Anzai, T., Shiina, T., Kimura, N., Yanagiya, K., Kohara, S., Shigenari, A., Yamagata, T., Kulski, J.K., Naruse, T.K., Fujimori, Y., Fukuzumi, Y., Yamazaki, M., Tashiro, H., Iawmoto, C., Umehara, Y., Imanishi, T., Meyer, A., Ikeo, K., Gojobori, T., Bahram, S. and Inoko, H., Comparative sequencing of human and chimpanzee MHC class I regions unveils insertions/deletions as the major path to genomic divergence, Proc. Nat. Acad. Sci. USA 100(13):7708–7713, 2003.
- Woodmorappe, J., Are pseudogenes ‘shared mistakes’ between primate genomes? TJ 14(3):55–71, 2000.
- Cartwright, R.A., personal communication, 8 July 2003.
- Wildman, D.E., Uddin, M., Liu, G., Grossman, L.I. and Goodman, M., Implications of natural selection in shaping 99.4% nonsynonymous DNA identity between humans and chimpanzees: enlarging genus Homo, Proc. Nat. Acad. Sci. USA 100(12):7181–7188, 2003.
- Shi, J., Xi, H., Wang, Y., Zhang, C., Jiang, Z., Zhang, K., Shen, Y., Jin, L., Zhang, K., Yuan, W., Wang, Y., Lin, J., Hua, Q., Wang, F., Xu, S., Ren, S., Xu, S., Zhao, G., Chen, Z., Jin, L. and Huang, W., Divergence of the genes on human chromosome 21 between human and other hominoids and variation of substitution rates among transcription units, Proc. Nat. Acad. Sci. USA 100(14):8331–8336, 2003.
- Walkup, L.K., Junk DNA: evolutionary discards or God’s tools? TJ 14(2):18–30, 2000.
- Mattick J.S., quoted in: Gibbs, W.W., The unseen genome: gems among the junk, Scientific American 289(5):46–53, 2003.
- Batten, D., More junk reclaimed, TJ 16(2):8, 2002.
- Woodmorappe, J., Pseudogene function: regulation of gene expression, TJ 17(1):47–52, 2003.
- Woodmorappe, J., Pseudogene function: more evidence, TJ 17(2):15–18, 2003.
- Woodmorappe, J., Junk DNA indicted, TJ 18(1);27–33, 2004.
- Gibbs, W.W., The unseen genome: gems among the junk, Scientific American 289(5):46–53, November 2003.
- Jones J., quoted in: Wieland, C., Furry little humans? Creation 24(3):10–12, 2002.
- Williams, A.R., Jumping paradigms, TJ 17(1):19–21, 2003.
- Fox, D., Wallaby nations, New Scientist 175(2354):32–35, 2002.
- Gibbs, W.W., The unseen genome: beyond DNA, Scientific American 289(6):106–113, 2003.


Readers’ comments
Comments are automatically closed 14 days after publication.