Journal of Creation 14(2):11–13, August 2000
Browse our latest digital issue Subscribe
Is the evolutionary tree turning into a creationist orchard?
Introduction
Evolutionists often infer that all organisms are related based on similarities at the physiological and anatomical level and, more recently, on ribosomal RNA (rRNA) homology at the gene level. Eldredge has highlighted this in his statement: ‘ … the major prediction of evolutionary theory is that there is one single nested pattern of resemblance linking all organisms’.1 As a result, there has been an attempt to group all life into one phylogenetic or family tree. But the large gaps between many supposed relatives have been a constant headache.
The recent elucidation of the complete genome sequences of 20 microorganisms has given fresh hope that this new data will help to reduce the gaps and strengthen the rRNA tree of life. But the data is instead proving to be a Trojan horse, as sequence comparison between homologous genes is yielding unexpected relatives and evolutionary lines, and different trees from the one originally predicted.2
Revised evolutionary model
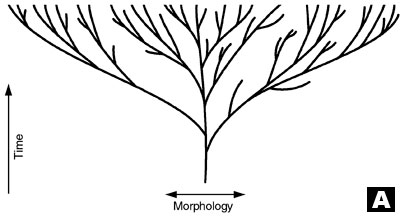

A. Original evolutionary tree, which postulates that all today’s species are descended from one common ancestor (after Wieland, concept by Wise).13,14
B. Revised evolutionary tree, depicting horizontal transfer among branches and a community of ancestral cells (after Doolittle).5
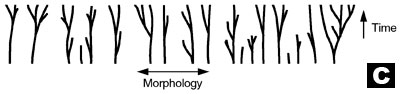
C. The Creationist orchard. Diversity has occurred with time within the original Genesis ‘kinds’ (baramins) (after Wieland, concept by Wise).13,14
In order to accommodate the new data, a new model for the evolutionary tree of life has been proposed which embraces the endosymbiont hypothesis (see below) and horizontal/lateral gene transfer (HT; gene exchange between unrelated microorganisms).2 Also, instead of one single organism at the root of the tree, a community of primitive cells is now believed to be the common ancestor:
‘It was communal, a loosely knit, diverse conglomeration of primitive cells that evolved as a unit, … and became the three primary lines of descent [archaea, bacteria and eukarya].’3
As a result, the once simple tree with a single trunk, rooted to a hypothetical 3.5 billion year old, ancient prokaryote, has become a tangled brier (see trees A and B in diagram), causing much frustration and discouragement.
‘There’s so much lateral transfer that even the concept of the tree is debatable.’ 4
‘It is as if we have failed at the task that Darwin set for us: delineating the unique structure of the tree of life.’5
But what is the scientific evidence for the involvement of the endosymbiont hypothesis and HT, two main tenets of this new model, in evolution?
Problems with the endosymbiont hypothesis
In the original endosymbiont model, an ‘ancient’ eukaryote with a nucleus supposedly evolved from an archaea-like prokaryote. It later engulfed an ancient prokaryote and a cyanobacterium (at separate times) that became mutually advantageous with the eukaryote host and developed into a mitochondrion and a chloroplast, respectively.
This simplistic original model has now been challenged, since ‘Many eukaryotic genes turn out to be unlike those of any known archaea or bacteria; they seem to have come from nowhere.’ 6
To try to solve this dilemma, two prominent phylogenetic theorists have proposed that an unknown and now extinct fourth domain of organisms transferred those genes horizontally into the eukaryotic nuclear genome.6 But there is no evidence for such a fourth domain of organisms, and this new idea unfortunately appears to be another ‘just so’ addition to help patch up an ailing hypothesis.
As shown in the following examples, current evidence also conflicts with the stepwise evolution from prokaryote to primitive eukaryote and then to eukaryote, proposed by the endosymbiont model:
‘Mitochondrion-free’ eukaryotes were believed to be descendants of ancient proto-eukaryotes, but mitochondrial genes have now been discovered in the eukaryotes’ nuclear genomes.7
A preserved protistan (single celled eukaryote) apparently dated to 1.4 billion years old appears to have had well-developed organelles, confirming the lack of evolutionary development of these structures and that fully functional organelles were always present in eukaryotes.8
Horizontal transfer (HT)
Bacteria have three main modes of exchanging DNA:9
Conjugation is used for exchanging plasmids. Plasmids are extrachromosomal, circular DNA molecules that carry not only genes needed for their own replication and transfer, but genes coding for proteins that protect bacteria, e.g. metabolic enzymes that break down toxic chemicals such as polychlorinated biphenyls, antibiotics, etc.
The plasmid-containing donor cell uses a pilus, an appendage that attaches to a recipient cell with a pilus receptor, to pull the cells together—some bacteria use chemicals called clumping factors instead of pili. A bridge, or pore, forms between the two cells and a copy of the plasmid is transferred. After transfer, the cells separate. It is possible, although quite rare, to transfer an entire chromosome by this process.
Bacteria can also pick up free DNA from their surroundings by transformation. For this to occur, cells must have specialised surface proteins to bind and to internalise the DNA. Transformation, however, is very rare in bacteria.
The third mode of exchange—transduction—involves bacterial viruses (bacteriophages). After a virus has injected itself into the host cell, it uses the cell’s DNA replication machinery to make many copies of itself. The cell eventually bursts, and the released virus particles are then able to infect other cells. Normally, viral DNA is packaged into the new virus particles, but sometimes some virus particles instead acquire bacterial host DNA. It is these particles that are used to deliver whole plasmids, as well as pieces of bacterial chromosome, to other bacterial cells.
Although these modes of genetic exchange are known to occur within bacterial species, it is believed that they are also involved in the HT of genetic material between unrelated microorganisms. The most quoted example is that of drug resistant pathogenic bacteria, such as Staphylococcus aureus (golden staph), which appear to have arisen from the exchange of plasmids, carrying multiple antibiotic resistance genes, between different bacterial species in the human body. It is important to note, however, that this exchange simply involves the sharing of genes already present in bacteria, and does not result in the appearance of complex, new genetic information. When the selective conditions are removed, the acquired genetic information becomes redundant and is eventually discarded by the cells to enable them to survive among the faster-growing wild-type bacteria. This is hardly an ‘uphill’ evolutionary process leading to the formation of complex organs and structures.10
What is overlooked then, is that bacteria live in a variety of environments (such as soil, water and inside plants and animals) and have the growth dynamics and the capability, as single cells, for genetic exchange, which they require to survive and adapt to changes in their environments. This merely shows the in-built, creative design in bacteria that allows them to occupy the bottom of the food cycling chains in a range of ecosystems. But it does not support microbe-to-man evolution.
The creationist orchard
As the evolutionary’ picture of origins is still unclear, scientists are hoping that analysis of additional genomic sequences and molecular phylogeny will bring some new light. However, they admit that, ‘Now new hypotheses, having final forms we cannot yet guess, are called for.’11
Comparisons of the DNA sequence data from the recently sequenced genomes, which have been determined by testable and repeatable scientific means, conflict with Darwin’s single evolutionary tree of life. As a result, polyphyly, in one sense the opposite of evolution or common ancestry, has been embraced in the form of HT and a community of ancestral cells. This is not surprising, as ReMine predicted that evolutionary ideology is bound to naturalism and simply ‘accommodates’ all evidence to fit reworked evolutionary models, thus showing that it is not falsifiable and therefore not scientific according to science philosopher Carl Popper’s primary criterion.12 He even predicted that evolutionists would increasingly resort to such lateral transfer.
If we tease apart the evolutionary brier and remove the hypothesised evolutionary HT ‘links’ between the branches, we obtain separate trees with individual trunks and roots. This is highly reminiscent of the creationist ‘orchard’, a biblical model for the origin and diversity of all life (see tree C in diagram) which was predicted by the scientific creation movement at its inception.13 In the creationist orchard, the trunk of each tree represents an original created kind or baramin, and the branches correspond to the diversity within a kind due to (limited) speciation, as for example, seen in the dog/wolf/jackal/coyote kind.
Conclusion
The creationist orchard agrees with the current genome sequence data, since anatomical, physiological and/or genetic similarities between kinds/baramins do not represent phylogeny but a designer. Evolutionary reasoning, on the other hand, cannot unravel the tangled phylogenetic brier and is unable to understand the true origin and diversity of created life, because its premises of naturalism and materialism axiomatically exclude the original work of a divine Creator.
The introduction of HT, the opposite of evolution or common ancestry, into the revised evolutionary model, clearly shows that nested hierarchy/phylogeny was never a prediction of evolutionary theory. Evolutionary theory simply keeps changing its ‘goalposts’ to produce reworked models that suit the current scientific beliefs.
References
- Eldredge, N., Do gaps in the fossil record disprove descent with modifications? Creation/Evolution IV:17–19, 1981. Return to text.
- Jerlström, P., Shaky tree of life, Journal of Creation 13(1):10–11, 1999. Return to text.
- Woese, C., The universal ancestor, Proc. Nat. Acad. Sci. USA 95:6854–6859, 1998. Return to text.
- Pennisi, E., Is it time to uproot the tree of life? Science 284:1305–1307, 1999. Return to text.
- Doolittle, W.F., Uprooting the tree of life, Scientific American 282(2):72–77, 2000;p. 77. Return to text.
- Doolittle, Ref. 5, p. 76. Return to text.
- Doolittle, W.F., A paradigm gets shifty, Nature 392:15–16, 1998; p. 15. Return to text.
- Bergman, J., The putative evolution of the animal eukaryote cell ultrastructure, Creation Research Society Quarterly 35(4):221–231, 1999; p. 228. Return to text.
- Miller, R.V., Bacterial gene swapping in nature, Scientific American 278(1):46–51, 1998. Return to text.
- Wieland, C., Antibiotic resistance in bacteria, Journal of Creation 8(1):5–6, 1994. Return to text.
- Doolittle, Ref. 5, p. 77. Return to text.
- ReMine, W.J., The Biotic Message: Evolution versus Message Theory, St Paul Science, St Paul, Minnesota, 1993. Return to text.
- Wieland, C., The beak of the finch: evolution in real time, Journal of Creation 9(1):21–24, 1995. Return to text.
- Wise, K.P., Baraminology: a young-earth creation biosystematic method; in: Walsh, R.E. and Brooks, C.L. (eds), Proceedings of the Second International Conference on Creationism, Creation Science Fellowship, Pittsburgh, Pennsylvania, vol. 2, pp. 345–360, 1990. Return to text.




Readers’ comments
Comments are automatically closed 14 days after publication.