Journal of Creation 33(3):55–65, December 2019
Browse our latest digital issue Subscribe
Baraminology data filtering method based on entropy measurement and its application in dinosaur and cephalopod data sets
Several recent dinosaur baraminology studies show areas of large degrees of continuity between baramins in the BDC matrixes where there should be none. These results suggest that the baramins predicted by the BDIST method in these studies tend to over-cluster, resulting in a smaller number of baramins, with falsified, inflated species memberships.
Also, evolutionists have used the BDIST method in an attempt to discredit baraminology, by trying to show that the number of dinosaur baramins gets smaller as more species are added to the analysis. This lumps dinosaur species together, showing the continuity of all life—which is the same as biological evolution. A potential problem was identified involving low-variability characters. Many such characters together tend to increase the correlation between any two given species and could possibly cause species lumping.
A new algorithm was developed to filter out such low entropy characters, and species with a high proportion of missing characteristics. The algorithm was applied on several dinosaur and two cephalopod data sets. It significantly cleans up the data sets and increases the number of baramins reported in previous studies, but eliminates many of the species and characters. There is a trade-off between the amount of available data and data quality. This affects the outcome of morphological baraminology studies.

Baraminology has a long publication record. In general, the field attempts to lump species into baramins and holobaramins. The holobaramin is the complete list of species, both living and extinct, belonging to a specific kind. The basic tenet of baraminology is that species within a baramin are related (continuous), whereas species from different baramins do not intermix (i.e. they are discontinuous). To reach the holobaramin, one can either keep on adding species together based on different lines of evidence until no more species can be added. Conversely, the holobaramin can be defined by dividing a larger group until species cannot be split any longer legitimately.1
The most common algorithm used is called BDIST.2 It is a phenetics-based algorithm, which calculates the pairwise correlation and baraminic distance between all possible species pairs in a study, based on a set of input characters with discrete values. The algorithm then creates a statistical graph (called a baraminic distance correlation matrix, or BDC), which shows how individual species relate to each other. Optimally, species from a single baramin cluster together on the graph. The designer has refused to share the program with this author. However, it was possible to reconstruct several features of the algorithm from published descriptions of BDIST.3
Several recent morphology-based baraminology studies seemingly tend to lump too many species into a single baramin. Baramins sometimes even overlap with one another. For example, a study by O’Micks lumped all decapods into a single holobaramin.4 This is a clade including organisms as diverse as squids and cuttlefish. Even evolutionists have found that the number of dinosaur baramins decreased from 50 to only eight in two studies that used the BDIST method that was developed by creationists.5 Wood replied in a subsequent analysis that selection of outliers, as well as a more holistic inclusion of all characters, may have given better results. Indeed, after applying these filters, his analysis gave more clear results.6
Following this, two recent studies lumped dinosaurs into a minimum of eight baramins,7,8 with the bold claim that not only are birds dinosaurs, but also that birds can be fitted into a morphological continuum with dinosaurs. Another study by Wood9 lumped Australopithecus sediba into the human holobaramin, although others have concluded that A. sediba is a mixed taxon and thus no analysis of this type can be performed before separating the ape and the human bones.10 When Wood took postcranial characters into consideration, he then changed his position, and stated that A. sediba was an australopith.11
The question arises, how far can we go with continually lumping species into a single holobaramin? How long can we continue decreasing the number of holobaramins, which individually have an ever-growing membership? Does it seem that baraminology methods would even seem to support evolutionary theory and the interrelatedness of all species? Whereas baraminologists are lumpers at a certain level, lumping can incorrectly be taken to an extreme. Using our scientific intuition, we should be able to break down large, over-lumped species clusters into smaller groups.
A recent review of baraminology methods by Cserhati and Tay12 has indicated several problems with morphology-based baraminology methods, such as the BDIST method, and has also suggested possible solutions to improve these methods, some of which are described below. Basically, the BDIST method is usable, although it should be refined and further developed.
On a practical level, it is important to note that many such data sets are messy, with many species having only partial data. For example, approximately two thirds of the Brusatte et al.13 data set of dinosaur remains used in a recent baraminological analysis by McLain et al.7 had undetermined character values. The other four data sets these authors used had between 54.2–69.8% undetermined character values. If a species has too many missing characters, its decreased information content may skew its relationship to other species. Missing data at a low level might be tolerable to some degree, but it’s an entirely different picture if more than half of the data is missing. This highlights the necessity of using more complete, quality data sets.
Furthermore, it is of utmost importance to select relevant characteristics (whenever possible), which are diagnostic of one of several baramins under study. Such diagnostic characters have the following characteristics: 1. They clearly differentiate between baramins, meaning that they are not uninformative or too general; 2. They have been measured for healthy adult individuals (and not juvenile or deformed individuals); 3. The measured character is not broken or fragmentary; and, 4. The character can be assigned an integer value (e.g. 0 = sagittal crest absent, 1 = present). In the case of continuous variables, character values can be put into range bins, or given binary values (e.g. 0 = <5 mm, 1 = >5 mm).
Selection of diagnostic character traits stems from the creationist presupposition that different kinds of plants and animals can be visibly and intuitively distinguished from one another. For example, birds are clearly separate from reptiles, because they were both created on separate days (Genesis 1:20–25).
However, there is a robust and long-standing discussion about character trait selection in the field of taxonomy. Character inclusion vs exclusion is a well-known problem. Most taxonomists have given up on character selection because observer bias so often influences it. Thus, most in the field have adopted a ‘throw everything at the wall and see what sticks’ approach and most modern taxonomical methods can deal with uninformative traits easily.
But in order to achieve the perfect classification of a given set of species, we would need to measure every single conceivable trait of all species in our study. This would involve thousands, even millions, of characters. This is clearly infeasible. A character selection scheme will always be imperfect. We simply have to accept this fact because we do not know everything.
On the other hand, what would happen if we were to study the osprey, the hammerhead shark, the boll weevil, the fruit fly, mouse, human, and the alga Volvox? What if the characters that we selected for study were these: does it have DNA? Is it multicellular? Is it eukaryotic? Does it have a cell membrane? This way all seven species would be classified into the same group. Clearly, we have to get rid of general characters (i.e. warm-bloodedness in a study of mammals).
For example, many bird species have air sacs which intrude into their bones, such as their femur. The femur of birds is immobile, and located within the body, in contrast with reptiles. Furthermore, the hip structure of birds differs from that of dinosaurs, which can be classified into one of two main categories, either lizard-style hips (Saurischia) or bird-style hips (Ornithischia). It is a paradox that birds supposedly evolved from Saurischian dinosaurs, which have a different hip structure.14 Birds have a closed, cup-shaped acetabulum, which serves as a joint between three hip bones—the ilium, ischium, and pubis. In comparison, dinosaurs all have an open, or perforated, acetabulum. The centre of gravity in birds also lies closer to its forearms compared to dinosaurs.
Also, the brain structure of birds and reptiles is very different. Reptiles have relatively larger olfactory bulbs compared to birds. Birds and reptiles occupy different curves on a log-log graph plotting encephalic (brain) volume according to adult body mass.14 Birds are endothermic as opposed to reptiles, which are ectothermic. Birds have a four-chambered heart, whereas reptiles only have a three-chambered heart with poor separation between the two ventricles. For a detailed discussion on the anatomical differences between birds and reptiles/dinosaurs, see Thomas and Sarfati, 2018.15 If we used these as diagnostic features, birds and dinosaurs would clearly form distinct groups.
But there are other features birds and dinosaurs share in common. For example, birds and reptiles are both oviparous. They also have scales and claws on their feet. If we included these characters in a morphometric analysis it could cloud the results.
Due to these considerations, this paper presents a baraminology data filtering method based on the measurement of the entropy value of different characters. Entropy is a mathematical measure of the variety of a given data set. Other authors describe it as the ‘surprisability’ of a given character. For example, if all of the specimens had the same value for a certain character, this character would be too general, and would hardly be useful in distinguishing two baramins (e.g. both birds and dinosaurs lay eggs). The ‘entropy’ of such a character is low. On the other hand, if multiple states exist for a single character, with an equal or almost equal number of species taking up different values of that character, the entropy for such a character would be high. The present method achieves data filtering by filtering out species with a high percentage of undefined characters, and by filtering out characters with low entropy (low character variation, meaning non-diagnostic, ambiguous traits).
Results and discussion
Re-analysis of two cephalopod data sets
For both cephalopod data sets, the major problem was reducing the large tentative decapod holobaramin into smaller groups. The Decapodiformes superorder includes squid and cuttlefish and is made up of the orders Sepiida, Sepiolida, Spirulida, and Teuthida. Therefore, it would be intuitive to find at least two smaller groups within Decapodiformes. Compared to the dinosaur data sets, a much smaller proportion of characters were undefined (21.7% and 30.4% of the Lindgren16 and Sutton17 data sets). Their mean pre-filter entropy values were also relatively higher than that of the dinosaur data sets (0.528 and 0.462, respectively). Their post-filter data reduction was also relatively less than that of the dinosaurs.

For the Lindgren data set, a ‘maximum row and column undefined percentage’ of 15% and a ‘minimum entropy value’ of 0.35 was selected prior to running BDIST. These were relatively severe cutoff values, but which increased the mean entropy to 0.72, with a 10.3-fold reduction in data, with a loss of 33 species and 84 characters. With a BDIST character relevance cutoff of 0.75, it was possible to separate eight species within the orders Sepiida, Sepiolida, and Spirulida from the rest of the Decapodiformes holobaramin (figure 1). The stress graph in supplementary figure 1 shows a minimum unscaled stress value of 0.085 at six dimensions. These groups include species such as cuttlefish. Two Octopodiformes species remained after the filtering, namely Eledone cirrosa and Opisthotuethis sp. The remaining 35 species all belonged to the order Teuthida, or squids. It is notable that in this analysis, Vampyroteuthis infernalis is reclassified as a decapod, as opposed to octopod as in earlier studies.4

According to the entropy filter of the Sutton17 study, nine species and 78 characters were filtered out. A maximum row and column undefined percentage of 30% and a minimum entropy value of 0.25 was selected. The mean character entropy rose from 0.462 to 0.698. Only a relatively mild data reduction of 3.6-fold was achieved. A BDIST relevance cutoff of 0.75 was applied to the filtered data set. The results of the BDIST analysis can be seen in figure 2. The stress graph in supplementary figure 2 shows a minimum unscaled stress value of 0.034 at nine dimensions.

The bootstrapping values of both cephalopod data sets were combined to get three decapod holobaramins by selecting species pairs with a bootstrap value ≥ 95+ in at least one of the two studies. This way we break down the order Teuthida into two suborders, Oegopsina and Myopsina, besides a group of three orders, namely Sepiida+Sepiolida+Spirulida (figure 3). The BDIST algorithm itself also allows for the setting of a character relevance cutoff value, which filters out characters that are present at a proportionately smaller percentage than the cutoff.
Re-analysis of four dinosaur data sets
Figure 17 of McLain et al.7 shows the BDIST analysis of 78 species coming from a data set by Brusatte et al.,13 which seemingly partition into four groups. However, these groups show continuity not only between themselves but also between each other. The BDIST results of this data set was too messy, because it appears that even though there are four main clusters, species from these clusters are continuous with one another to a large extent. Therefore, the whole data set was subjected to entropy filtering.
Table 1. Parameters used for data filtering and BDIST re-analysis.

The parameters used during the data filtering and the BDIST re-analysis for each of the four dinosaur data sets studied by McLain are available in table 1. These parameters include the maximum unknown character per row, maximum unknown character per column, minimum character entropy, and the BDIST relevance cutoff. Table 2 contains the parameter values of certain characteristics in the four data sets, pre- and post-filter (number of species, number of characters, % undefined values, and mean entropy).
Table 2. Changes in different data sets after entropy filtering.

A maximum undefined character percentage of 35% per row and 35% per column as well as a minimum entropy percentage per column of 35% was set. This resulted in a reduction in the number of characters from 853 to 370, and the number of species was reduced from 152 to 19 (because many closely related species became indistinguishable without those characters). This meant a data reduction of 18.4-fold, but the mean character entropy rose greatly from 0.113 to 0.774.

The BDIST algorithm was re-run on these 19 species, with a relevance cutoff of 0.95. The stress graph shows a minimum unscaled stress value of 0.02 at six dimensions. The BDC results (figure 4) show four clusters, with one cluster of six species showing significant discontinuity with the other three groups, made up of 13 species. These six species are Tyrannosaurus rex, Tarbosaurus bataar, Alioramus, Daspletosaurus, Gorgosaurus libratus, and Albertosaurus sarcophagus. These six species are morphologically similar and all fall into the superfamily Tyrannosauroidea. The stress graph in supplementary figure 3 shows a minimum unscaled stress value of 0.02 at six dimensions.
One of these species, however, namely Alioramus, does not show discontinuity with four other species in this study, namely Guanlong, Dilong paradoxus, Sinraptor dongi, and Allosaurus fragilis. Analysis of the Lee et al.21 data showed that T. rex showed continuity between two of these species, namely S. dongi and A. fragilis. Therefore, these four other species may be part of the Tyrannosauroidea group as well.
The nine other species not mentioned yet are Sinorthomimus, Struthiomimus altus, and Gallimimus bullatus in one smaller group of three species and six species in another group: Citipati osmolskae, Velociraptor mongoliensis, Deinonychus antirrhopus, Bambiraptor feinbergi, Shuvuuia deserti, and Archaeopteryx lithographica. These two subgroups show neither continuity nor discontinuity between themselves. Therefore, with this study we cannot make a definitive statement as to whether they form one or two holobaramins.
Figure 32 of the McLain study7 shows the BDC analysis results of the Lee et al.21 study. The results seem to be too messy, although two large groups are apparent in the figure. Therefore, this data set was also subjected to entropy filtering. A maximum undefined character percentage of 50% per row and 50% per column as well as a minimum entropy percentage per column of 20% was set. This way the number of characters was reduced from 1,549 to 828, and the number of species was reduced from 120 to 15. This meant a data reduction of 15-fold, but the mean character entropy rose greatly from 0.104 to 0.686.
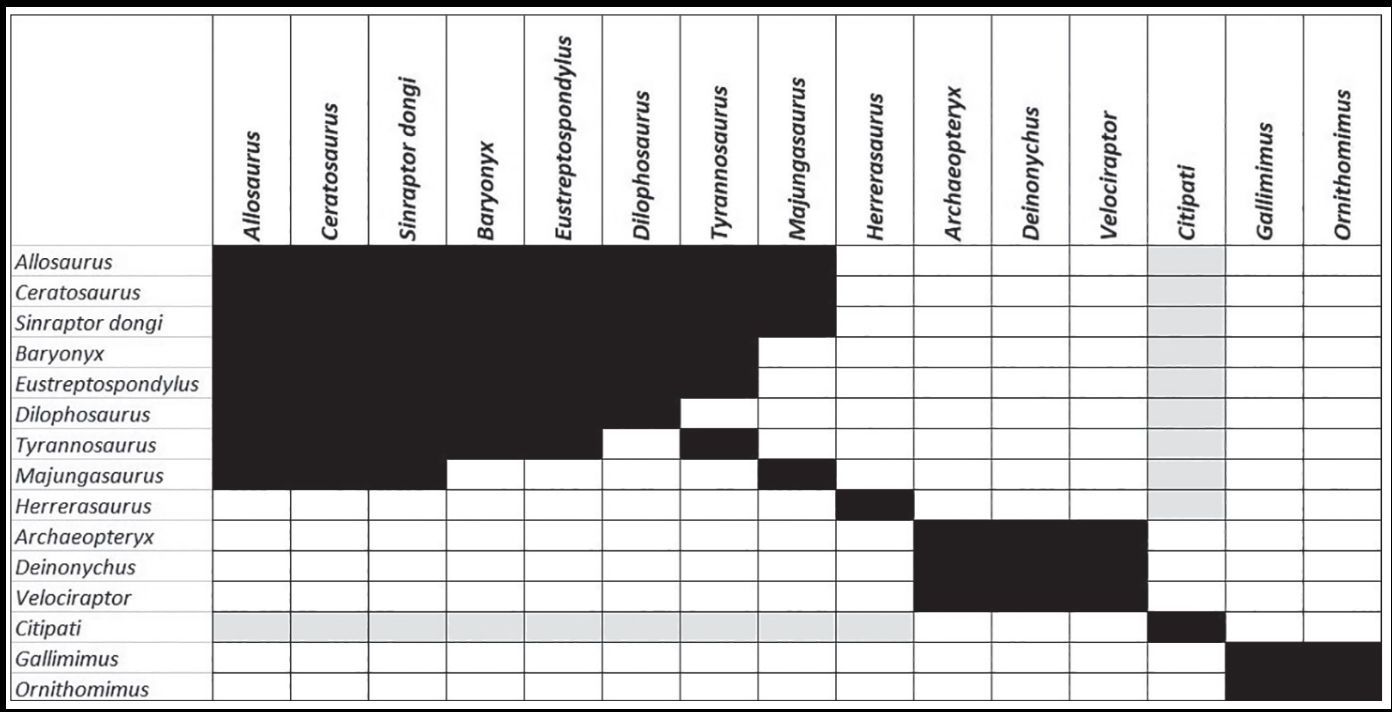
The BDIST algorithm was re-run on these 15 species, with a relevance cutoff of 0.95. The stress graph shows a minimum unscaled stress value of 0.0225 at 10 dimensions. The BDIST results show three main clusters with at least two species, besides several singleton species (figure 5). The stress graph in supplementary figure 4 shows a minimum unscaled stress value of 0.022 at ten dimensions. The first cluster is made up of eight species, Majungasaurus, Tyrannosaurus, Dilophosaurus, Eustreptospondylus, Baryonyx, Sinraptor dongi, Ceratosaurus, and Allosaurus. Three species, Velociraptor, Deinonychus, and Archaeopteryx, form a smaller group. Two Ornithomimosaurians form another smaller group of two species in the BDIST results in the upper-right corner.
Figure 56 of the McLain study depicts what could be either three or four clusters from the van der Reest data set.22 Therefore, entropy filtering was applied to the species in this data set. A maximum undefined character percentage of 35% per row and 35% per column as well as a minimum entropy percentage per column of 20% was set. This way the number of characters was reduced from 366 to 249, and the number of species was reduced from 93 to 22. This meant a data reduction of 6.8-fold, but the mean character entropy rose greatly from 0.286 to 0.730. The BDIST algorithm was re-run on these 22 species, with a relevance cutoff of 0.75. The stress graph shows a minimum unscaled stress value of 0.0298 at eight dimensions.

In the BDIST results we can see four very clearly defined clusters of five, three, eight, and five species, respectively, along with a singleton species, Shuvuuia deserti (figure 6). The stress graph in supplementary figure 5 shows a minimum unscaled stress value of 0.03 at eight dimensions. However, significant discontinuity exists between the third cluster of eight species and the first two clusters of five and three species. The first cluster of five species is made up of Sinosauropteryx prima, Tyrannosaurus rex, Gorgosaurus libratus, Sinraptor sp., and Allosaurus fragilis. The second cluster of three species is Ornothomimus edmontonicus, Struthiomimus altus, and Gallimimus bullatus. There is neither discontinuity nor continuity between these two clusters, so therefore they could possibly be part of the same holobaramin, although they may also form different holobaramins.
There is a large degree of discontinuity between these species and the following two clusters. The third cluster of eight species is made up of Confuciusornis sanctus, Velociraptor mongoliensis, Deinonychis antirrhopus, Bambiraptor feinbergi, Mei long, Gobivenator mongoliensis, Archaeopteryx lithographica, and Anchiornis huxleyi. The last cluster of five species includes Khaan mckennai, Ingenia yanshini, an unnamed oviraptorid IGM100-42, Citipati osmolskae, Caudipteryx sp., and Confuciusornis sanctus. There is neither discontinuity nor continuity between these two clusters, so therefore they may be part of the same holobaramin, but they may also form different holobaramins.
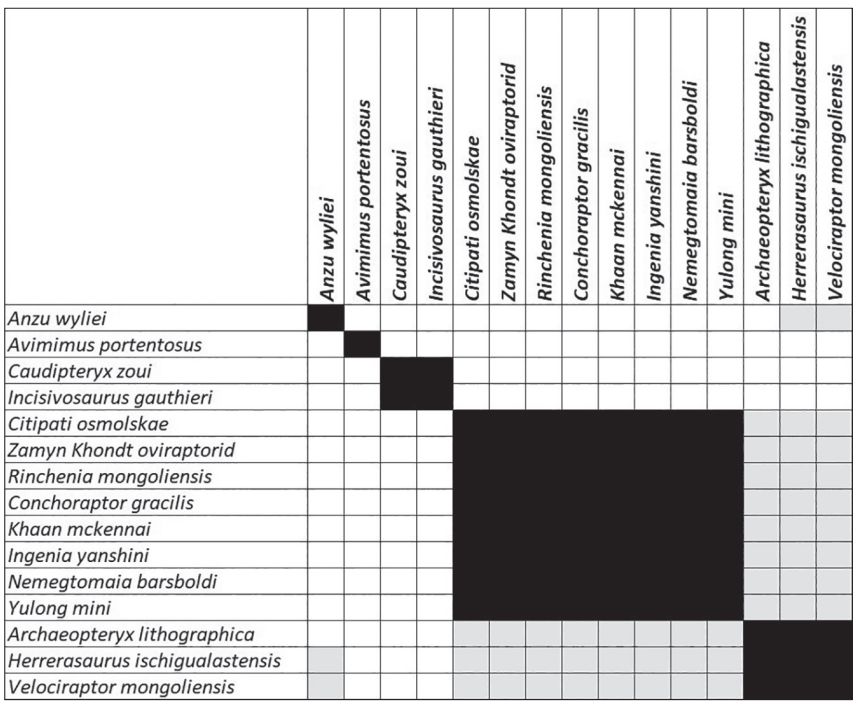
Figure 64 of the McLain et al.7 study depicts BDIST results from the Lamanna et al. study.23 In this study, 41 species were studied with 230 morphological characters. A maximum undefined character percentage of 50% per row and 50% per column as well as a minimum entropy percentage per column of 25% was set. Entropy filtering reduced the number of characters from 230 to 173. This meant a data reduction of 6.1-fold, but the mean character entropy rose greatly from 0.113 to 0.793. The BDIST algorithm was re-run on these 22 species, with a relevance cutoff of 0.75. The stress graph shows a minimum unscaled stress value of 0.084 at five dimensions.
Three clusters of two, eight, and three species are visible in the BDIST results (figure 7). The stress graph in supplementary figure 6 shows a minimum unscaled stress value of 0.086 at five dimensions. There is visible discontinuity between the third cluster and the previous two clusters. The first cluster includes two species: Incisivosaurus gauthieri and Caudipteryx zoui. The second, larger cluster consists of eight species: Yulong mini, Nemegtomaia barsboldi, Ingenia yanshini, Khaan mckennai, Conchoraptor gracilis, Rinchenia mongoliensis, Zamyn Khondt, and Citipati osmolskae. There being neither continuity nor discontinuity between these two groups we cannot yet say whether these two groups form one or two holobaramins. The third group, however, clearly separates from the first two clusters. This third group is made up of three species: Velociraptor mongoliensis, Herrerasaurus ischigualastensis, and Archaeopteryx lithographica.

Table 3, column 3, lists 50 dinosaur species found in the BDIST analyses of the reduced morphology matrixes after entropy filtering. For each species the corresponding set of studies is listed in the first column. This same information is depicted in the Venn diagram in figure 8. In this figure each possible combination of 15 data sets shows the number of species that were discovered in that particular combination of data sets. For example, according to table 3 and figure 8, three species—Archaeopteryx lithographica, Velociraptor mongoliensis, and Citipati osmolskae— were found by the Brusatte, Lee, Lamanna, and van der Reest data sets.
All of these results can be summed up into a single baraminic classification. For each of the four analyses, we can take those species pairs where the bootstrapping results have a value higher than 95%. These species pairs would then form an edge in a graph. The thickness of such an edge would be proportionate to the number of studies that this species pair shows up in (1–4). This species graph can be seen in figure 9, which shows three holobaramins with 22, 12, and four species, respectively. These 38 species are all therapod saurischian dinosaurs, and are listed in supplementary table 1, along with their cluster number, as well as their taxonomic classification into order, clade, and superfamily/family.

The first, largest holobaramin is called Maniraptora, with 22 species. This holobaramin could possibly be split up into two smaller monobaramins of 10 and 12 species, respectively, called Avialae and Oviraptosauria. These species come from a number of different families, including Alvarezsauridae, Anchiornithidae, Archaeopterygidae, Confuciusornithidae, Dromaeosauridae, Herrerasauridae, and Troodontidae.
The second group is called Tyrannosauroidea and contains species coming from six different reptile families/superfamilies, namely Tyrannosauridae, Allosauridae, Ceratosauridae, Proceratosauridae, Megalosauridae, and Metriacanthosauridea. Members of Tyrannosauroidea differ little in their morphology from species to species and are characterized by short forelimbs on robust pectoral girdles, with only two fingers. They have relatively large skulls in proportion to their bodies.
Table 3. List of different dinosaur species found by different combination of studies.

In his study of tyrannosauroid taxa, M. Aaron concluded that the family Tyrannosauridae+Bistahieversor+Appalachiosaurus+Dryptosaurus+Raptorex+Xiongguanlong+Eotyrannus all constitute a holobaramin. The species Eotyrranus seemingly represents the extreme form of tyrannosauroids, with longer, grasping hands and three digits. The author also stated that with the discovery of further tyrannosauroid fossils, the definition of this holobaramin might change, including other tyrannosauroid species, such as Dilong paradoxus.18 According to the results of the present study, Dilong is continuous with Allosaurus fragilis, which itself is continuous with Tyrannosaurus rex.
Therefore, the species Guanlong, Sinraptor dongi, Baryonyx, Eustreptospondylus, Ceratosaurus, and Sinosauropteryx prima might be added to the existing tyrannosauroid holobaramin. This could mean that species belonging to the superfamily Tyrannosauroidea all form a single holobaramin. This would include species from the family Proceratosauridae (such as Guanlong), which have a positive correlation with Eotyrannus, according to two studies.19,20
The third group consists of four species, all from the family Ornithomimidae, namely Gallimimus bullatus, Ornithomimus edmontonicus, Sinornithomimus sp., and Struthiomimus altus. These species are characterized by their slender, light frame, long, slim hindlimbs, and elongated forelimbs ending in uniquely structured hands. Since there are only four species in this small group, it may be that these four species only form a monobaramin.
Entropy filtering applied to Senter, 2010 data set
Senter,5 an evolutionist, claims that after using the BDIST method on a data set of 40 characters for 33 fossil dinosaur species, compiled in 2009, the dinosaur groups Oviraptorosauria, Avialae, Deinonychosauria, and Coelurosaurian outlier species all fall into a morphological continuum, implying that the application of this baraminological method decreases the number of holobaramins, and shows continuity between dinosaurs and birds. Wood6 responded by pointing out that Senter’s selection of characters was too stringent, appealing to a holistic inclusion of most if not all characters using less stringent selection criteria. In Wood’s new character matrix, taxa had at least 50% of their characters in a known state, raising the number of included taxa to 42, with 177 characters. Figure 2 of Wood6 shows that the distance correlation results for the larger character matrix, with less stringent criteria, shows discontinuity between Oviraptorosauria+Avialae+Deinonychosauria and Coelurosaurian outlier species. This study indicates that character selection influences the results of the BDIST analysis, and therefore impacts the number of holobaramins as a result.
In the present paper, an expanded data set from Senter5 covering 89 species and 364 characters was analyzed using the BDIST software with and without entropy filtering. Supplementary figure 7 shows the distance correlation results (relevance cutoff of 0.75), showing general high continuity between all species in the study. The species do not cluster at all into well-defined holobaramins. The stress graph in supplementary figure 8 shows a minimum unscaled stress value of 0.24 at four dimensions. This value is very high, and even shows an increase starting from 5 dimensions. This is a clear indication that this data set needs to be filtered.
Thus, entropy filtering was applied to Senter’s data set, with a maximum undefined character percentage of 50% per row and 50% per column, and a minimum entropy percentage per column set at 25%. Entropy filtering only reduced the number of species to 42 and the number of characters to 215. The mean character entropy rose greatly from 0.379 to 0.726, meaning a data reduction of 3.6-fold. The BDIST algorithm was re-run on these 42 species, with a relevance cutoff of 0.75.
In supplementary figure 9 we can see that the groups Oviraptorosauria and Avialae+Deinonychosauria show significant negative baraminic correlation with Ornithomimosauria+Tyrannosauroidea. Both Oviraptorosauria and Avialae+Deinonychosauria, as well as Ornithomimosauria and Tyrannosauroidea do not show significant negative baraminic correlation with each other, but neither do they show significant positive baraminic correlation. These results indicate a separation of birds from dinosaurs and are similar to the results in figure 2 in Wood.6 The stress graph (supplementary figure 10) shows a minimum unscaled stress value of 0.067 at seven dimensions.
Conclusions
Character selection influences the result of BDIST studies, whether all possible characters are chosen or whether they are selected based on special selection criteria. This is true, for example, when multiple protein sequence alignments are trimmed to obtain the most informative set of amino acids which produce an optimal evolutionary tree. The BDIST method was re-applied to several dinosaur morphology character sets, and, as we have seen, a reduction was made possible in the number of dinosaur baramins, averting the scenario of all species coalescing into one common baramin (which is equal to the evolutionary tree of life, suggesting general evolutionary relatedness between all species).
It must be noted here that the most optimal scenario would be to make measurements of all characters for all species, which however is hardly possible for fossil remains. Fossil character data sets tend to be messy and easily skew results, thereby making entropy filtering of the data sets necessary. Filtering the data sets involves removing rows (species) and columns (characters) which contain too much unknown information and too little variation. This way character entropy/diversity increases, with a parallel decrease in the number of species. However, the result is that the boundary of baramins can be made much clearer. Setting cutoff values for these parameters can fine-tune BDIST correlation results.
A radical reduction in characters may also skew BDIST results. But this was necessary due to the fact that many of the data sets contained a very large percentage of missing values. Whereas the combined results in this paper show that Oviraptorosauria, Avialae, and Deinonychosauria form a single holobaramin, they are connected by only one species, Citipati osmolskae. The results from Wood6 suggest that they separate from one another, even though significant baraminic discontinuity is lacking. However, when the 2010 Senter data set was subjected to entropy filtering, Oviraptorosauria separate from Avialae+Deinonychosauria, but also Ornithomimosauria and Tyrannosauroidea, do not show significant baraminic continuity with one another, suggesting four baramins.
Materials and methods
The entropy filter algorithm was applied to 90 cephalopod and 227 dinosaur species from four data sets. The goal was to refine at times overlapping baraminic predictions which have possibly been over-lumped into a smaller number of possible holobaramins. This was done on the Lindgren16 and Sutton17 cephalopod data sets, and the Brusatte13, Lee21, van der Reest22, and Lamanna23 dinosaur data sets. The BDIST results of the Zanno24 dinosaur data set showed that the discovered putative baramins were well-segregated enough so as to make further analysis using the entropy filter unnecessary.
For each data set, a small subset of species was identified which made up what was deemed to be an over-lumped cluster. Character entropy filtering was performed on these species within the data set. The BDIST method was re-run on the remaining filtered data set to see if the entropy filtering was able to split up the selected species into a larger number of baramins, each with a smaller species membership.
The results of data filtering and reclustering are reported here. For the analysis of each data set the original data set is included as well as a list of species for which entropy filtering was done. Furthermore, the filtered character set and the BDIST results are also provided for each analysis in a separate Excel file, which are available on github.com/csmatyi/EntropyFilter.
Data sets for the cephalopod and dinosaur baraminology studies listed in table 1 were downloaded. The script EntropyFilter. R was written in R studio, version 1.1.442. The script itself, as well as supplementary figures and data files, are available on the github web page.
The script applies several filters to the data. First, it filters out those species which have a percentage of undefined (“?”) characters above a certain cutoff. Next, it selects those species which are over-lumped. The names of these species should be listed in a separate txt file. The third filtering step is the most important and is essential to the whole method. This filtering step involved calculating entropy for each character. A column of character values is extracted from the double-filtered data set. Those characters are filtered out, and contain a certain percentage of undefined characters, just as with the row filtering criterion. Shannon entropy is calculated for each of the characters, minus the undefined states of a given character. Mixed characters, such as {0,1} are treated as separate characters (thus, 0, 1, and {0,1} count as three states of a given character). Shannon entropy is calculated in the following manner for a given character j:

Where n is equal to the number of states for character j, pi is equal to the probability of observing state i of the given character, and is equal to the number of occurrences of state i/the total number of occurrences for a character j. A minimum undefined character ratio for rows and columns and a minimal entropy value was selected for all data sets.
Figures 3 and 9 were made using Cytoscape version 3.7.1. The bootstrapping values of the BDIST results of the entropy-filtered Brusatte, Lee, Lamanna, and van der Reest analyses were combined. An edge was placed between two species (vertices) if their bootstrapping value was ≥ 95+. Edge thickness was adjusted to reflect the number of BDIST studies which showed continuity between a given pair of species.
The baraminic distance correlation matrix as well as the stress graphs for all studies were generated using the BDIST software at coresci.org/bdist.html. The Venn diagram (figure 9) was created using the software available at bioinformatics. psb.ugent.be/webtools/Venn/.
Acknowledgements
The author would like to thank Matthew McLain of The Master’s University for providing the data set from the van der Reest and Currie publication.
References and notes
- Wood, T.C., A baraminology tutorial with examples from the grasses (Poaceae), J. Creation 16(1):15–25, 2002. Return to text.
- Wood, T.C., BDIST software, v. 1.0, Center for Origins Research and Education, Bryan College, 2001. Distributed by the author. Return to text.
- github.com/jeanomicks/bdist_weighted Return to text.
- O’Micks, J., A preliminary cephalopod baraminology study based on the analysis of mitochondrial genomes and morphological characteristics, ARJ 11:193–204, 2018. Return to text.
- Senter P., Using creation science to demonstrate evolution: application of a creationist method for visualizing gaps in the fossil record to a phylogenetic study of coelurosaurian dinosaurs, J. Evol. Biol. 23(8):1732–1743, 2010 | doi:10.1111/j.1420-9101.2010.02039.x. Return to text.
- Wood, T.C., Using creation science to demonstrate evolution? Senter’s strategy revisited, J. Evol. Biol. 24(4):914–918, 2011 | doi:10.1111/j.1420-9101.2010.02208.x. Return to text.
- McLain, M.A., Petrone, M., and Speights, M., Feathered dinosaurs reconsidered: new insights from baraminology and ethnotaxonomy; in: Whitmore, J.H. (Ed.), Proceedings of the Eighth International Conference on Creationism, Creation Science Fellowship, Pittsburgh, PA, p. 508, 2018. Return to text.
- Doran, N., McLain, M.A., Young, N., and Sanderson, A., The Dinosauria: baraminological and multivariate patterns; in: Whitmore, J.H. (Ed.), Proceedings of the Eighth International Conference on Creationism, Creation Science Fellowship, Pittsburgh, PA, pp. 404–457, 2018. Return to text.
- Wood, T.C., Baraminological analysis places Homo habilis, Homo rudolfensis, and Australopithecus sediba in the human holobaramin, ARJ 3:71–90, 2010. Return to text.
- Rupe, C. and Sanford, J., Contested Bones, FMS Publications, Waterloo, NY, 2017. Return to text.
- Wood, T.C., Australopithecus sediba, statistical baraminology, and challenges to identifying the human holobaramin; in: Horstemeyer, M. (Ed.), Proceedings of the Seventh International Conference on Creationism, Creation Science Fellowship, Pittsburgh, PA, 2013. Return to text.
- Cserhati, M. and Tay, J., Comparison of morphology-based and genomics-based baraminology methods, J. Creation 33(3):49–54, 2019. Return to text.
- Brusatte, S.L., Lloyd, G.T., Wang, S.C., and Norell., M.A., Gradual assembly of avian body plan culminated in rapid rates of evolution across the dinosaur-bird transition, Current Biology 24:2386–2392, 2014. Return to text.
- Clarey, T., The Science of the Biblical Account of Dinosaurs, Master Books, Green Forest, AR, 2015. Return to text.
- Thomas, B. and Sarfati, S., Researchers remain divided over ‘feathered dinosaurs’, J. Creation 32(1):121–127, 2018. Return to text.
- Lindgren, A.R., Giribet, G., and Nishiguchi, M.K., A combined approach to the phylogeny of Cephalopoda (Mollusca), Cladistics 20:454–486, 2004. Return to text.
- Sutton, M., Perales-Raya, C., and Gilbert, I., A phylogeny of fossil and living neocoleoid cephalopods, Cladistics (32):297–307, 2016 | doi:10.1111/cla.12131. Return to text.
- Aaron, M., Discerning tyrants from usurpers: a statistical baraminological analysis of tyrannosauroidea yielding the first dinosaur holobaramin, ARJ 7:463–481, 2014. Return to text.
- Xu, X., Wang, K., Zhang, Q., et al., A gigantic feathered dinosaur from the Lower Cretaceous of China, Nature 484(7392):92–95, 2012. Return to text.
- Lü, J., Yi, L., Brusatte, L.S., et al., A new clade of Asian Late Cretaceous long-snouted tyrannosaurids, Nature communications 5(3788), 2014. Return to text.
- Lee, M.S.Y., Cau, A., Naish, D., and Dyke, G.J., Sustained miniaturization and anatomical innovation in the dinosaurian ancestors of birds, Science 345(6196):562–566, 2014. Return to text.
- van der Reest, A.J. and Currie, P.J., Troodontids (Theropoda) from the Dinosaur Park Formation, Alberta, with a description of a unique new taxon: implications for deinonychosaur diversity in North America, Canadian J. Earth Sciences 54(9):919–935, 2017. Return to text.
- Lamanna, M.C., Sues, H.-D., Schachner, E.R., and Lyson, T.R., A new large-bodied oviraptorosaurian theropod dinosaur from the latest Cretaceous of western North America PLoS ONE 9(3):e92022, 2014. Return to text.
- Zanno, L.E., A taxonomic and phylogenetic re-evaluation of Therizinosauria (Dinosauria: Maniraptora), J. Systematic Palaeontology 8(4):503–543, 2010. Return to text.





Readers’ comments
Comments are automatically closed 14 days after publication.