Journal of Creation 15(2):55–58, August 2001
Browse our latest digital issue Subscribe
The weasel returns: Truman replies to Curtis
Let’s locate my missing selection procedure. Analogous to Professor Dawkins’ data, each generation is initiated by one new sequence by selecting between: one identical to the preceding; or one from a set of n-1 candidates, whichever is closer to the goal.1 I used a total of n=2 offspring2 and Dawkins presumably more.
Dawkins warned us: ‘You have to become an advocate and use the tricks of the advocate’s trade. This book is not a dispassionate scientific treatise’.3 My algorithm removes detracting details which for modelling purposes are superfluous to render transparent what is going on.
Mr Curtis writes, ‘There are 8 mutations in the 9 generations between generation 1 and generation 10 in Dawkin’s original example’. Dawkins’ original example was presented in the New Scientist. He reported far more than 8 mutations for the first generation (see Table 1).4

In the same page Curtis discusses, Dawkins published his second example (see Table 2).5 Almost all positions have ‘mutated’.

We can be accomodating by some creative ad hoc corrections. For instance, in the example in Table 2 one might arbitrarily perform a reading-frame shift on generation #0: move everything one position to the left starting at position 4; arbitrarily eliminate the last X; then slide the last four letters one position to the left. And so on.
Dawkins and Curtis wish to guarantee that mutations don’t destroy eons of evolutionary labour. ‘The existence of progeny exactly resembling their parents is necessary to conserve accrued matches.’ We agree this has been achieved in a programming manner open to discussion. From the computer-generated output reported5–7 we find no exceptions to the rule: once the desired letter is in place, keep it there.
We can speculate as to how Dawkins tuned his program to guarantee progress towards the goal and regeneration of the preceding genome to try again whenever necessary. Although he wrote both BASIC and Pascal programs,8 and demonstrated such programs repeatedly before television audiences,9 he is not able or willing to provide the original or a new source code.10
The algorithm favored by Mr Read11 is precisely how evolutionist David Wise programmed and mimicked Dawkins’ results,10 and has been circulated by the evolutionist National Center for Science Education:
‘Yet the simulation uses mutation in a peculiar way. Each progeny is identical with the parent except that a single letter position is selected at random and a randomly selected character is inserted in that position.’9
Professor Dembski agrees Dawkins prevented the letters correctly lined up from mutating.12 One can model the lining up of all rings by rotating a large number of n rings and keeping the best each ‘generation’ or by changing only 1 ring.1,2 The only interesting difference is that convergence is faster if multiple rings are used with a large n.
Nevertheless, Curtis’s favoured interpretation, which he is unable to justify (see his Addendum), and proposed rate of individual letter changes could be either close to what Dawkins may have programmed, or intended to program, or can be made (with suitable choice of progeny size) to mimic the reported data acceptably well. These are simply alternative manners of tuning the number of progeny and mutation characteristics to achieve the same ‘rigged fruit-machine’. At issue is not whether Dawkins’ or my algorithms have any biological realism. Both have none whatsoever.
Curtis’s proposed rate of ‘letter mutation’, 1/40–1/30, may be interesting in deciphering Dawkins’ program, but is biologically meaningless. In discussing the fidelity of DNA replication, Dawkins wrote, ‘The probability of any particular letter being miscopied on any one copying occasion turns out to be a little more than one in a billion’.13
Is it true that ‘Dawkins’ program is not at all like Truman’s program’?
What Dawkins wishes to show, and is implied by his results is: ‘Given, say, a hundred million Xs, we should be able to construct a plausible series of tiny gradations linking a human eye to just about anything!’14
So according to Dawkins:
1. One can start from any random sequence and very quickly converge to a pre-determined goal.
2. Selection is determined by correct letters, the effect of incorrect ones has no relevance.
Let us neglect the superfluous. Visualize boxes into which letters start to pop up as generations go by and don’t disappear once in place (Table 3).
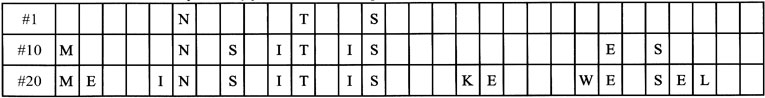
We need not overly trouble ourselves about the mutated letters which do not yet line up to their target (left blank above). They play absolutely no role in Dawkins’ program. They allegedly neither enhance nor hinder selection in any manner and for modelling purposes can be neglected: de facto, either a letter is properly lined up, or it can be ignored altogether. Since everyone agrees that the best single offspring is retained, no unwanted mutations are reported for the ‘losers’. Read proposed a mutational rate of 1/28 per letter, and Curtis slightly smaller, compensated for by a large number of progeny of which the best is selected.
I pointed out2 that the same convergence behaviour is demonstrated (ignoring letters not lined up properly) by generating as candidates one identical offspring, plus one new one for which each letter can mutate, every generation. Why convergence is unavoidable is now easier to understand.

Forty runs were made with my original program to provide us with some examples as to the number of correctly aligned letters after each generation. Figure 1 shows that these results straddle the examples Dawkins provides which are not badly garbled.15,7 Most of the curves lie to the right of Dawkin’s data, i.e. my algorithm always converges but on average a little slower than Dawkin’s.
The advantage of distilling the program to the relevant aspects becomes apparent. My algorithm is deterministic, as everyone realizes. Dawkins’ treatment merely confuses the issue by invoking ‘progeny’, ‘mutations’ and ‘generations’ to produce biological associations. Both algorithms work like funnels, with no possibility for failure nor variety since only one offspring is selected every generation.
Mathematician Dembski also recognized that Dawkins’ algorithm is deterministic.11
Here is the essence. Dawkins reports no mutational rates, almost no intervening sequences, and no number of progeny. I availed myself of a common method used in mathematics and modelling which could be summarized as: ‘if a cow can’t jump over a mountain then it can’t jump over the moon’. If my algorithm is deterministic and cannot fail to converge towards any preset sentence, then a second algorithm, beginning with the same random sequence, which converges even more rapidly to the same goal every time with no possibility of failure must also be deterministic!
In Figure 2, average convergence behaviour between both programs is illustrated using the same starting sequence as in Dawkins’ second example.16,7 My program1 had been run 10,000 times in 1998. Values between 35 and 331 for an average of 102 generations were found.7 947 runs took 64 or more generations. Although these results clearly straddle Dawkins’ data, on average my data converges more slowly (Figure 2), and for statistical reasons already explained1 cannot fail to converge. Dawkins’ reported data lies to the left of this curve and simply ‘rolls down the funnel’ a little faster.
Is Dawkins modelling anything real?
We have already pointed out that Dawkins is not simulating any real biological phenomena.1 Selection does not drive relentlessly towards a specific genetic goal; the order in which each base-pair mutation arises is deemed irrelevant; the proteins always fold properly (never gum up due to exposed hydrophobic regions); and there is, remarkably, no variety among reproducing offspring.
A biologically meaningful model would show that, on average, for every successful mutation acquired many destructive ones would be introduced.17 The neglected letters not lined up are biologically relevant. Natural selection cannot eliminate all flaws fast enough without repeated extinctions.
Protein folding studies18 suggest less than one DNA sequence of average gene length out of 1065 could have any biological activity. Yockey calculated19 1.5 x 1044 random trials would be needed to find the first minimally functional cytochrome c polypeptide. Even generating one billion progeny every second for 10 billion years provides only 3 x 1026 opportunities20 (<<1.5 x 1044) before Darwinian selection could start fine-tuning.

In the real world, proportionally more than 27 of Dawkins’ 28-sentence positions would have to be already correctly lined up, by random attempts without any kind of selection, before Darwinian selection could act.21 Assuming a continuous genetic path from randomness to functional genes, with every fork selected flawlessly to guide towards a goal millions of years in the future, has no relevance to nature.
Dawkins discusses the histone H4 gene, for which the proportion of acceptable sequences to possibilities of comparable length is << 1 out of 1065:
‘there is a length of 306 characters in cows, which is virtually identical to a length of 306 characters in peas. Cows and peas differ from each other in only two characters out of these 306.’22
He placed their common ancestor about 1.5 billion years ago.22 Since mutations have generated so little variety in all this time, presumably the smallest of errors leads to nonviable or drastically inferior genes which thus cannot be produced by countless improvements starting from a random sequence.
Biological realities are incompatible with proposed changes of 1/40 to 1/30 per ‘letter’. From the context, Dawkins’ sentence appears to represent genes. Since mutations are random, this indicates that on average about 3% of the whole genome in the germ-line would change every generation on average. Since a large number of offspring is generated, this indicates a complex, multi-cellular organism such as a fruit fly.
Let us pretend worm the C. elegans (the ‘simplest’ multicellular animal showing complex development23,24) produces a large number of progeny. It has a 97-megabase genome coding for 19,099 proteins. Dawkins’ mutations have to be quarantined to the germ-line. Consider the effect of 3% of average random changes for every one of the thousands of genes, repeated dozens of times, as cells duplicate to produce an embryo or replace somatic cells during each organism’s lifetime! Self-destruction is guaranteed, but impossible in Dawkins’ program. Furthermore, the mutated gene would now appear in countless unrecognizable variants across all subsequent cells during the organism’s lifetime after a few dozen replications leaving selection with nothing consistent to identify.
Guaranteeing ‘The existence of progeny exactly resembling their parents’ may be useful in Dawkins’ make-believe computer world but is clearly absurd: logically, since generating an average of hundreds of thousands of mutations per genome every generation precludes providing a pristine original; and biologically since the program allows no possibility for extinction.
Conclusion
We don’t know exactly how Dawkins’ BASIC and Pascal programs were coded. My program is not exactly the same as Dawkins’, agreed. However, it gives similar results and uses a similar mechanism, where the ‘good’ mutations are effectively protected and a mechanism to guarantee progress to the goal is ensured. The essence of what was allegedly being modelled is now easier to grasp. Whether one does this by having a lower mutation rate and larger number of offspring to ensure that some are the same as the parent, or one copies the parent and then mutates another, it has fundamentally the same effect—protection of desired letters, which is not the real world. My program clearly demonstrates the ‘fail-safe’ nature of any such ‘simulation’ of evolution.
The statistical analysis1 demonstrates Dawkins’ program is a meaningless gimmick which cannot fail to converge to the same predetermined goal every time, which is evolutionary and biological nonsense contra Dawkins’ protests.25
Acknowledgement
I appreciate Bart Read’s thoughtful comments, both in his letter to the TJ and in discussing this reply to Curtis.
References
- Truman, R., Dawkins’ weasel revisited, Journal of Creation 12(3):358–361, 1998. Return to text.
- Truman, R., ‘Royal Truman replies’, Letters, Journal of Creation 13(2):74–75, 1999. Return to text.
- Dawkins, R., The Blind Watchmaker, Penguin Books, London, p. X, 1986. Return to text.
- Dawkins, R., Creation and natural selection, New Scientist, 34:34, 1986. Return to text.
- Dawkins, Ref. 3, p. 48. Return to text.
- Dawkins, Ref. 4, p. 34. Return to text.
- Details showing the workings of Dawkins’ and my algorithms can be found at Supplementary information for “The weasel returns: Truman replies to Curtis” Return to text.
- Dawkins, Ref. 3, p. 49. Return to text.
- Gitt, W., Weasel words, Creation 20(4):20–21, 1998. Return to text.
- ReMine, W., The Biotic Message, St. Paul Science, Minnesota, p. 235, 1993. Return to text.
- Read, B., ‘Dawkins’ weasel revisited’, Letters, Journal of Creation 13(2):73–74, 1999. Return to text.
- Behe, M.J., Dembski, W.A. and Meyer, S.C., Science and Evidence for Design in the Universe, Ignatius, San Francisco, p. 38, 2000. ‘(1) Start out with a randomly selected sequence of 28 capital Roman letter and spaces … ; (2) randomly alter all the letters and spaces in this initial randomly generated sequence; (3) whenever an alteration happens to match a corresponding letter in the target sequence, leave it be and randomly alter only those remaining letters that still differ from the target sequence. In very short order this algorithm converges to Dawkins’ target sequence.’ Return to text.
- Dawkins, Ref. 3, p. 124. Return to text.
- Dawkins, Ref. 3, p. 78. Return to text.
- Dawkins, Ref. 3, p. 47. Dawkins’ run which converged in 43 generation already had 3 letters lined up in the initial random sequence and the (corrected) example on p. 48 which converged in 64 generations had only 1 already lined up. Return to text.
- Dawkins, Ref. 3, p. 48. Return to text.
- A century of genetic research world-wide under optimal laboratory conditions using quick reproducing organisms has produced a very large number of mutations with probably none useful from the evolutionary viewpoint: fitness enhancing, information increasing to produce biological novelty. A very large number of destructive mutations have been reported, however. Return to text.
- Reidhaar-Olson, J. and Sauer, R., Proteins, Structure, Function and Genetics, 7:306, 1990; Bowie, J. and Sauer, R., Proceedings of the National Academy of Sciences, USA 86:2152, 1989; Bowie, J., Reidhaar-Olson, J., Lim, W. and Sauer, R., Science, 247:1306, 1990; Behe, M., Experimental support for regarding functional classes of proteins to be highly isolated from each other; in: Buell, J. and Hearns, G. (Eds), Darwinism: Science or Philosophy? Haughton Publishers, Dallas, pp. 60–71, 1994. Discussed in Ref. 11, p. 75. Return to text.
- Yockey, H., Information theory and molecular biology, Cambridge University Press, p. 254, 1992. Return to text.
- (1010 yrs) x (365.25 days/yr) x (24 hrs/day) x (3600 sec/hr) x 109 members = 3x1026 attempts using an average generation time of 1 second, 1 billion years and 109 offspring. Return to text.
- Each position has one chance out of 27 of being correct. Getting 27 positions correct by chance has a probability of (1/27)27 = 2 x 10-39 . Return to text.
- Dawkins, Ref. 3, p. 123. Return to text.
- Wilson, R.K., How the worm was won. The C. elegans genome sequencing project, Trends in Genetics 15(2):51–58, 1999. Return to text.
- A C. elegans Database (ACeDB). Return to text.
- Dawkins, Ref. 3, p. 72. After discussing another program (biomorphs) we read: ‘It sounds, once again, as though evolution deals in distant targets, homing in on things like scorpions. As we have seen, it never does’. Return to text.


Readers’ comments
Comments are automatically closed 14 days after publication.