Journal of Creation 30(3):78–87, December 2016
Browse our latest digital issue Subscribe
Origins of pathogenic microbes: part 2—viruses
From creation the ecosystem was stable and microbes participated in maintaining the homeostatic state. The basic types of microbes, including selected viruses, appear to have been created. Sin apparently brought a change to features of DNA repair and gene regulation in living systems and also stress was introduced into the equation. This resulted in mutations and other errors giving rise to cellular malfunctions. Hypotheses based on published data are presented to account for the emergence of the different categories of viruses. On the basis of genome similarity, horizontal transfer of genetic information apparently occurred among living organisms, which helps to explain viral diversity. Some potential exchange mechanisms have been identified, but new ones are likely to be discovered. Alternatively or in addition, a modification of original creation designs is being attempted by one who said “I will be like the Most High” (Isaiah 14:14). Some of the outcomes observed could represent a cunning reworking of the original genetic information to achieve results not previously imagined—as achieved by humans in their genetic engineering experiments. Via these processes, new pathogens may still be emerging. Versions of the theory of evolution commencing with viruses depend on a number of ‘miraculous’ events for their success. By contrast, the biblical story of creation appeals to Creation Week events and the disruption caused by the Fall.

In part 1 of this article, the origin of bacterial pathogens was considered. The involvement of viruses in bacteria acquiring pathogenic capabilities was mentioned. In this section, the origin of viruses will be discussed in detail. Viruses, particularly bacteriophages, represent the most numerous biological entities found in the natural world.1 The basic proposition used in this article is that microbes were not disease-causing initially in creatures with pain sensations. Bacteriophages (phages) are considered part of the original creation plan, for they carry out many significant functions in the bacterial world. Change after the Fall conceivably involved shifts in the ecosystem balance so that the nature and behaviour of organisms was altered. It is also postulated by some that an agent(s) altered or added novel genetic information to the genome of existing organisms after the Fall or an entirely new line of microbes was created (as part of God’s curse).2
The existence of beneficial phenomena in the natural world that are widely expressed will be taken to indicate their essential continuity from the beginning (parsimonious approach), except where biblical information dictates otherwise.
Origin of viruses
The origin of viruses is uncertain. In the evolutionary scheme they are sometimes seen as the commencing building blocks of life, which requires a series of miracles to accomplish.3 This view is rejected here. Virus genetic material is made of RNA or DNA. Creationists have adopted a number of approaches to their origin: viruses, or at least some, were created; they arose from existing elements and structures through naturalistic means; guided change was responsible; or a combination of these phenomena occurred.
Viruses, transposons (DNA sequences capable of changing position), and plasmids (small extrachromosomal DNA molecules) display some similarities, but no single gene is shared by all groups. However, there are different groups of shared genes that form links among these elements. This could indicate that exchange of genes may have occurred as well as host gene incorporation into some elements. Host gene incorporation is seen particularly in viruses with large genomes.1 The proposition forwarded in this paper is that the limited number of virus hallmark genes that have been identified may be taken to indicate that some basic virus forms were present from creation. The variants observed today arose from these basal types.
Created or basic types created
The concept that viruses were part of the original creation is sometimes promoted. The idea is that they, as with the more regular microbes, were made to contribute to the operation of the ecosystem and confer benefits on invertebrates and higher-order hosts.4 The proposal that viruses, in general, were in useful relationships with the entire creation before sin has limited observational support. Examples of beneficial relationships involving viruses (other than bacteriophages or phages—see below) include parvoviruses that enable wing development in aphids, parvoretroviruses of plants that protect against pathogenic viruses, and a mycovirus able to confer plant thermotolerance.5 Some parasitic wasps are dependent on viruses for their success. For example, an ascovirus in the wasp Diadromus pulchellus inhibits the deposition of melanin by the leek moth larvae (Acrolepiopsis assectella) that would normally encapsulate the wasp’s eggs and prevent their development. Other wasp viruses may act similarly.6 However, for plants and mammals, it has been difficult to identify benefits flowing from viral infection. Suggested examples are plant drought resistance and protection from other viruses and, in mammalian hosts, protection from damaging viruses.5 In some mammals another example is the expression of amylase activity in saliva, which confers adaptive advantages. This activity is facilitated by the presence of retrovirus elements.7
Others reject a creation origin for viruses due to the ill repute with which most viruses are held, their classification as non-living entities8 (some experts only), the concept that there was no death in Eden, that ecological balance is possible in the absence of such a culling process involving bacteria, and that all negative outcomes are attributable ultimately to Satan.
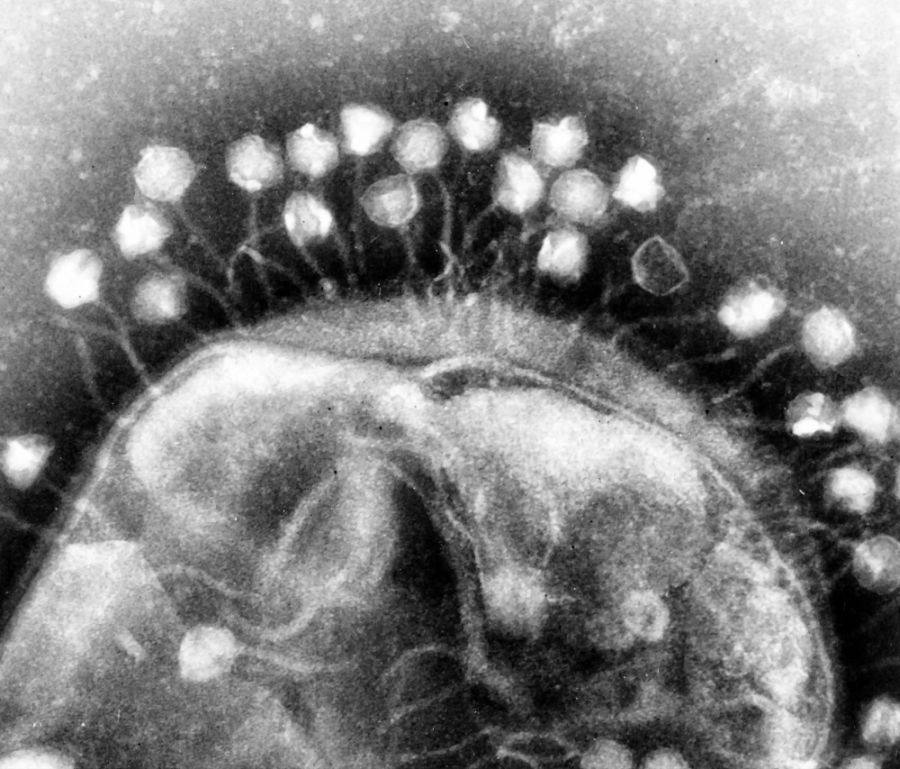
An intermediate view is that creation of special viruses occurred rather than the general creation of viruses. The mildest version of such an approach is to regard some asymptomatic animal and plant viruses potentially in this category. Indeed, it is noted that plants cannot be cured of their cryptic viruses (partitiviruses—function unknown).9 There are also asymptomatic viruses among the insects, which have no known function.10 Among the bacteria, phages (figure 1) ostensibly were made to assist in the control of explosive bacterial growth for the maintenance of ecosystem balance. Phages are widespread, function to improve bacterial growth, protect against chemical agents and stress, assist in biofilm formation (an essential feature of bacterial life), and occasionally prevent pathogenicity and dampen mutation.11 All these attributes represent benign, but very useful, features for bacterial existence. Some phages may function as plasmids,12 which seems to hint at their essential role. The argument for the creation of some categories of viruses may be strengthened on account of the cogent reasons given supporting the concept of death among the non-feeling organisms in Eden before the entrance of sin.13
The argument in favour of some basic virus types being created appears to stand on stronger ground than other proposals. It also has the benefit of denying Satan creation credentials. This means he would be restricted to manipulating that which was created.
The following brief account of virus origins takes evidence from scientific work published in refereed journals. Some of these papers argue that eukaryotes arose from simpler forms of life whereas others champion the idea that viruses were derived from the genomes of their hosts. The concept adopted here is that in the beginning there was a near-simultaneous appearance of life-forms from the Creator’s hand. After sin entered, massive alterations appeared. This implies that microbes have adapted to changing conditions to give pathogens by exchange and recombination of existing information. The scientific evidence, when paired with this concept, leads to a reasonably satisfying fit.
Naturalistic derivation from existing elements
The processes occurring in nature are treated in this section, although a number of concepts are logical suppositions not yet supported by hard evidence. Hypothesis making precedes evidence gathering in the scientific endeavour, which means that propositions are refined over time.
Two types of nucleic acid are possessed by viruses requiring somewhat different emphases. First there is the ribonucleic acid (RNA) group and secondly the deoxynucleic acid (DNA) group. Some of the viruses in the latter group show similarities to genetic elements in the genome of living organisms, such as retrotransposons. These represent mobile elements that have other representatives too—transposons and other elements. Such pieces of genetic information may constitute up to 45% of the human genome and 37% of the mouse genome. Only a small group (80–100) of those present in a mammalian genome can move and influence the behaviour of other pieces of DNA (retrotransposons). They are responsible for genetic diversity, which includes diseases caused by insertional mutagenesis. Some of these retrotransposons are regarded as endogenous retroviruses14 due to selected retrotransposons possessing many features similar to those displayed by retroviruses (HIV is a member of this group).8
RNA viruses
In RNA viruses, the hallmark protein (enzyme) is RNA-dependent RNA polymerase. The structural similarity among the different categories of RNA viruses—positive, single-stranded (+ss)RNA, negative, single-stranded (–ss) RNA, and double-stranded (ds)RNA—is high. However, no similarities of the gene are present in eukaryotes. In order to solve the issue of origins, it has been postulated that the +ssRNA viruses of eukaryotes arose from +ssRNA bacteriophages or a more remote possibility is from the limited RNA virus representative(s) of Archaea. In turn these viruses are thought to have given rise to the –ssRNA and dsRNA viruses. Horizontal virus transfer to other hosts and intermixing of elements then may have occurred. Such a scenario appears to have taken place with fungal and plant viruses in particular. This makes sense as fungal-plant interactions are common. The close relationship observed between some plant and fungal viruses suggests that mixing of elements has given rise to new virus derivatives. Other groups of +ssRNA viruses also display many genome similarities, which suggests that the spread of genes among plant viruses occurs commonly, giving rise to new variants.1
Another possibility is that new plant viruses could be made by transfer of information from the plant to the virus. RNA plasmids are found in animals, fungi, and plants and replicate similarly to selected RNA viruses. Such plasmids would need to acquire genetic information allowing a coating protein to be fashioned.15 Evidence of such a scenario appears to be supported by studies with potato leafroll virus (+ssRNA). In one study, there was extensive similarity found in nucleotide sequences in potato leafroll virus and a tobacco chloroplast gene. This suggested to the investigators that recombination occurred between virus RNA and host plant messenger RNA.16 Many viruses that are non-pathogenic for a particular host can replicate in the initial cells inoculated but cannot spread. It is in these cells that recombination theoretically could occur, conferring on the virus particle altered abilities. Indeed, invasive chimeric viruses have been generated in the laboratory when plants containing a segment of a plant virus genome are inoculated together with a related virus, even a non-infecting one.17

Figure 2. Electron micrograph of Influenza A virus.
The –ssRNA viruses have narrow host ranges. The influenza A group of viruses (figure 2) have a segmented genome and illustrate rather well the capacity of segments from different sources to reassort and perhaps also mutate to create novel strains capable of causing deaths in the human and animal populations.18 Besides chance spread of respiratory viruses across the species barrier via droplet transmission, the arthropod parasites of animals and plants appear to facilitate horizontal transfer of some other viruses. In fact, the protein sequences and architecture of –ssRNA viruses is similar in plants and animals. It is thought that these viruses arose from the +ssRNA viruses; a suggestion based on crystal structure similarities involving selected members of each group. There is also a possibility that dsRNA phages were involved.1
With the double-stranded RNA viruses, the cystoviruses (phages) appear to have contributed most of the structural genes found in dsRNA viruses occurring in eukaryotes. Again, horizontal gene transfer among plants, vertebrates and arthropods may have contributed to the emergence of distinctive virus lineages.10 And there is some evidence that heritable information in a dsRNA virus-like particle (parasitic on a fungus) arose from RNA plasmids.15
Reverse transcribed elements in eukaryotes include the retroviruses. In these viruses, RNA is reverse transcribed into DNA. The only unifying feature of these entities is the reverse transcriptase feature. Some are able to integrate into the host genome, others behave as plasmids. Well-known retroviruses are infectious, but there are others that are inactivated by blockage of cell receptors because other retroviruses have integrated into the genome. Deletions and mutations also may render them inactive.19,7 Retroelements are common in eukaryotes and are represented among prokaryotes, but they do not possess envelope-forming capabilities. However, these retroelements may be of more than passing interest as follows.
Virus-like particles may have been derived from retrotransposons. Viruses appear to have acquired the functional gene(s) for a virus envelope from some other source, possibly the host or other viruses.20,1 Indeed, some retrotransposons carry envelope-like genes.21 Hence, it is possible that the acquisition of a functional envelope gene may have resulted from protein domain fusion leading to the appropriate gene being formed.20 Another factor involved in the genesis of a virus is its release from a host cell. These cells possess similar release phenomena to enveloped viruses, as noted in the formation of microvesicles. Such vesicles are produced by cells into spaces outside them.22The vesicles may carry a limited amount of DNA and some RNA,23 which can include retrotransposon elements under stress conditions.24 Microvesicles also are present in human milk, together with an array of microbial entities. It is conceivable that information transfer may occur in this environment.25
In summary, RNA viruses appear to have arisen in various ways. These may involve phages, host RNA (plasmids), retroelements present in hosts, and other RNA containing viruses. The mixing of virions in host cells gives the opportunity for variants to arise too. It seems possible that all these features could have arisen through the operation of naturalistic phenomena.
DNA viruses
These viruses come as single-stranded (ss) or double-stranded (ds). I will commence the account of possible origins with the ssDNA viruses. The eukaryote ssDNA viruses replicate using a rolling circle mechanism initiated by the enzyme endonuclease. This method of replication is used by most of the prokaryote ssDNA viruses, many of the plasmids, and some transposons. On account of distinct structures in eukaryote ssDNA viruses, it appears improbable that they arose from similar viruses in prokaryotes. On the other hand, they share a number of significant similarities to bacterial plasmids (small DNA molecules replicating separately from the chromosome), suggesting possible origins from them.1 This suggestion is made on account of a number of lines of evidence. One example is that the bacterium Agrobacterium is able to support geminivirus (plant virus) replication when its DNA is experimentally inserted into the bacterium. This is on account of the viruses containing bacterial promoter sequences.26

Figure 3. Plasmids and chromosomal DNA shown schematically in a bacterial cell.
The conversion of a plasmid (figure 3) into a virus requires encapsulation of the nucleic acid by a protein coat and the acquisition of genetic information allowing intercellular transfer. The similarities between the structure of virus coat protein found in various viruses suggests that information transfer from bacterial plasmids and +ssRNA viruses has occurred, allowing the emergence of ssDNA viruses. Recombination events have been shown between RNA and DNA viruses so that the suggested marriage of plasmid DNA and RNA viruses is not outlandish. Indeed, since many bacteria are parasitic or mutualistic in eukaryotes and these may also be hosts to a variety of viruses, the opportunity for transfer and recombination exists. Genetic exchange could occur in cells infected simultaneously by both bacterium and virus. Such occurrences are known to occur.27,1
A greater amount of information is available on the possible origins of dsDNA viruses. Double-stranded DNA viruses are widely distributed in the eukaryotes and may have arisen in a number of ways. The first possibility involves transposons, which are transposable pieces of DNA. Large transposons have been termed polintons. They contain proteins needed for their own transposition. It is hypothesized that they arose originally from a linear plasmid.28 Most of these mobile elements contain two proteins similar to those found in the capsids of viruses residing in bacteria, archaea, and eukaryotes, and also contain additional proteins needed for virus development. This information can be interpreted to indicate that virions could arise from polintons under suitable conditions, 1,29although this has not yet been achieved in the laboratory.
The origin of another category of viruses, the virophages, appears to involve polintons too. Virophages are small dsDNA viruses that reduce the replication capacity of a category of large viruses (mimiviruses). Genome analysis indicates that virophage genes share genetic information with the polintons.29 They also show similarities to amoebal transposons and host genes as well as genes found in plasmids and bacteriophage.30 This can be taken to suggest horizontal gene transfer involving a number of organisms and entities and that virophages could have descended from polintons.
Plasmid involvement in the origin of viruses also is indicated with the replication of poxvirus (vaccinia—dsDNA virus). Virus duplication in animal cells may use proteins required for the replication of selected plasmids. Furthermore, a common cytoplasmic area of the cell is used for replication of both the virus and plasmid.31 This indicates the close similarity between what can be regarded as normal cell processes and those utilized by viruses. It also suggests that reassortment of genetic material present in cells could give rise to novel combinations characteristically found in selected viruses.
Turning to a different group of host organisms, a somewhat different set of circumstances may contribute to the emergence of viruses. Genomes from large viruses (dsDNA) contain homologues of genes found in their hosts, indicating that these genes have been transferred. For example, a large virus (mimivirus) found in amoebae (Entamoeba/Dictyostelium) contains proteins (serine/ threonine kinases) apparently derived from its host. One plausible route for such exchange has been suggested. Amoebae can ingest other microbes and break down (lyse) these cell occupants thereby releasing nucleic acid into the cell environment and creating a DNA soup allowing the possibility for gene acquisition. How this exchange and rearrangement of DNA might be accomplished is not known.32 However, the observation is that hallmark genes shared by many groups of viruses show similarities to cellular genes,33 which suggest that viruses may have arisen from several cellular sources through horizontal gene transfer.34
Some dsDNA viruses, such as herpes viruses, appear to have arisen from bacteriophages.1In turn, some bacteriophages (dsDNA phages) may have been derived from double-stranded DNA molecules (plasmids) that can replicate independently of the bacterial chromosome. A single mutation can enable some phages to exist as plasmids.35 Again, special antibiotic proteins termed bacteriocins may be phage-like in structure when they are released from the bacterial cell, which perhaps indicates recombination activity between plasmids and bacteriophage. Indeed, phage tail-like protein has been found within selected bacteriocin operons.36 These observations indicate that movement and recombination of genetic material within bacterial cells enjoy a wide range of possibilities in today’s environment. The dynamics of interactions among viruses is incompletely understood, but cells simultaneously infected with herpes (dsDNA) and retrovirus (RNA made into DNA by reverse transcriptase) allows integration of retrovirus genes into the herpes virus.37
In summary, DNA viruses appear to have arisen in several ways. These may involve transposons, plasmids, other eukaryote host genes, and phages. While no virus has been engineered in the laboratory from these starting materials, the suggestions made are based on similarities in genetic makeup and architecture among structures and a certain amount of experimental evidence indicating that a range of exchange, recombination, and reassortment of genetic information is possible. All these features appear capable of arising through the operation of naturalistic phenomena.
Genetic manipulation
The biblical account of life in Eden and on the earth excluded pain, shedding of blood, and death through old age (Genesis 3:3; Revelation 21:4). This raises the issue of the origin of the changes that are observed today and whether naturalistic phenomena alone have been responsible for them.
God-arranged change
Entrusting God with the privilege of making pathogens in a special creative act (curse) cannot be substantiated readily. God is loving, good, and incapable of evil (Matthew 19:17; James 1:13; 1 John 4:8). Thus, it seems implausible to some to argue that He created pathogens causing pain and suffering in feeling forms of life. God well understood evil’s nature and could have generated it, but Jesus’ statement “If a kingdom is divided against itself, that kingdom cannot stand” (Mark 3:24, ESV) precludes this possibility. The magnificent design of the immune system, His instruction to the Israelites on how to maintain health, and providing knowledge to moderns on how to fight disease are all clear evidences of His good will.
Changes in the mismatch DNA repair system allowing mistakes in repair of DNA damage and such like may constitute part of the Curse mentioned in Genesis 3:17. Changes may have occurred as a result of cellular stress such as imposed by dietary and environmental factors. These stress factors are known to activate transposable elements and give rise to mutations and to change the way in which microbes interact with the host. Restricting access to the tree of life also may have contributed through limited access to significant nutritional supplements, as indicated by the field of epigenetics (study of gene expression not involving DNA sequence changes). Epigenetics has to do with gene expression levels and the factors that influence them, which primarily are dietary and environmental stress. These stress factors act through changes to DNA, such as adding a methyl group to the cytosine base of DNA, and by modifications introduced into histone (proteins associated with eukaryote DNA) and ultimately to chromatin structure. Genes can be silenced or activated as a consequence of these changes.38 Finally, the general curse of death on all plants and creatures meant that much more organic matter was available for decay. In this cauldron, DNA of all types conceivable became available for re-assortment leading to the emergence of microbes with unusual capacities.
Human interference
Humans are able to effectively manipulate genes—i.e. generation of crops high in targeted nutrients; creation of insect, herbicide, and virus-resistant crops; transfer of spider and wasp toxins to plants; and introduction of computer-designed novel proteins.39 In addition, humans can alter the pathogenic capability of organisms.
Pathogenic ability can be acquired and lost. Recently, Dr Craig Venter’s group manipulated the smallest free-living pathogen Mycoplasma genitalium in unusual ways. First, they synthesized the organism’s genome in the laboratory. They then disrupted the gene that caused pathogenicity and finally inserted the artificial genome into yeast cells where it successfully replicated. This was followed by the replication of an artificial chromosome in a related bacterial cell robbed of its nucleus, so creating a semi-artificial microbe.40The gene involved with pathogenicity allowed the microbe to adhere to host cells, an ability that can be lost, or reduced, by mutation, or artificial manipulation.41
It is possible to block the expression of disease in plants through introducing silencing codes into the pathogen’s genome.42 Equally, disease-causing capabilities might be introduced through genetic engineering. Great advances are being made in taking functional genes and placing them in different organisms. Future advances propose the design of organisms to perform ordered tasks.43
Mechanisms present in created organisms can be copied and reworked into something positively sinister, such as is done in germ warfare. This has been accomplished with an animal influenza virus. The original virus infected humans but did not pass easily between individuals. However, targeted mutations of the virus particle made it an effective airborne entity.44 This paralleled work with a pox virus some years previously. The virus was made highly damaging by incorporating mouse-derived molecules. It was then able to undergo uncontrolled replication causing death in the experimental mice, which normally were resistant.45
An unexpected recent discovery is that in the nonretroviral RNA virus (lymphocytic choriomeningitis) genetic material may hybridize with retrotransposon DNA. Such hybridization has been observed in the mouse. This observation raises the possibility that virus genes may be integrated into a mammalian genome and that humans could facilitate such an outcome in their own kind by using RNA virus vectors in gene therapy experiments.46
Other interferences
After Satan failed to find majority support in heaven, he was expelled and denied significant interaction with heavenly beings (Revelation 12:7–9; Job 1:6–7, 2:1–2). This indicates that limits were placed on his activities just as limits were placed on his annoyance of Adam and Eve (Genesis 3:1–3). The forces of evil are permitted to work within the bounds set by God and His plan to bring the reign of wickedness to a just end (Psalm 34:7; Revelation 13:5, 7, 15).
Satan’s power over nature is beyond human capabilities (Job 1:12–19; Psalm 8:5; Hebrews 2:7). He has the ability to manipulate diverse organisms as suggested in the account recorded in Job. At that time, Satan was able to induce experimental infection at will (Job 2:7). In understanding the latter phenomenon refer to my first article on the origin of pathogenic bacteria (e.g. staphylococci). It is also fruitful to refer to relatively recent community outbreaks where special environmental and contact conditions were shown to permit mass infection by Staphylococcus to occur.47 These observations may be taken to indicate that Satan possessed advanced knowledge on microbial behaviour and ecology at the time of Job. The Bible also speaks of thorns and thistles arising as a consequence of sin (Genesis 3:18). One possibility is that thorn-like structures arose through directed crossing and mutations, as has been demonstrated experimentally in the laboratory.48 Alternatively, they could have arisen through genetic manipulation. Such unusual outcomes have been achieved through using both classical breeding methods and genetic engineering.49
Genetic engineering involves horizontal transfer of genes. It seems untenable to acknowledge Satan’s great power over nature and his ability to perform the spectacular (Job 1:18–19; Matthew 4:8), which is well in excess of human capabilities, and then argue he cannot match human endeavours in the field of genetic engineering. Satan can alter existing information within limits, and he is at times able to use his human agents to do the unthinkable, as in biological warfare.50
After Satan’s expulsion from heaven, it might be expected that he would disrupt God’s creation and blame Him for the suffering caused, as creative ability is God’s hallmark (Isaiah 42:5, 65:17; Colossians 1:16). One of my basic premises is that pre-existing structures and mechanisms have functioned as prototypes to devise malignant forms. Ingenious combinations and innovations among genetic resources, particularly involving opportunistic organisms, could have facilitated pathogenic organism generation. The emergence of some viruses might represent an engineered result. However, a strategy to detect such occurrences would be difficult if not impossible to devise.
Genesis and expression of pathogenic capabilities
Virulence, the capacity to cause disease, is not strictly an intrinsic microbial characteristic but includes both microbial and host factors.51 With viruses, which are technically non-living entities, features other than intrinsic characteristics are emphasized. In all considerations relating to pathogenic ability, the environment also can be highly significant.
Microbe changes
The generation of pathogenic viruses from asymptomatic or defective ones is a possibility. Defective viruses integrated into the chromosome may give rise to pathogenic variants when recombinants form as a result of genetic exchange. This has been shown with a number of retroviruses and indicates that in the right circumstances, exchange of genetic information may take place without great difficulty.52
Virus may exchange information and even take genes from the host organism. This has been documented in the laboratory following the chance acquisition of a retrotransposon from insect cells into a baculovirus (dsDNA).53 The resulting changes in host range, virulence, and other features of such an acquisition are unknown.

The emergence of human immunodeficiency virus (HIV—figure 4) from simian immunodeficiency virus (SIV) appears to be the result of two phenomena. One is transmission of SIV from monkeys to humans, thought to have occurred by contamination of fresh wounds by monkey blood during their slaughter for human consumption. The contaminating virus particles are thought to have possessed a limited capacity to replicate in human tissues. However, among the contaminants thus introduced a mutant apparently was selected conferring the ability to replicate well within human cells (second phenomenon). On an experimental basis, when HIV was used to infect simian hosts, a reverse mutational change was noted; that is, the virus now became adapted to growth in apes rather than in humans. This observation gives credibility to the suggested mechanism behind the host range extension seen.54
Other retroviruses may arise as a result of recombination events between those integrated into the chromosome or those external to it. This outcome is indicated by the identification of a unique avian leucosis virus.55
Host changes
The best known example of host changes leading to disease manifestation is with immunocompromised and immunosuppressed organisms. These hosts may be predisposed to attack by microbes not normally considered pathogens or by rare pathogens that flourish in unusual locations. Virus infection of the central nervous system fits into the latter category. An example is John Cunningham virus, which is capable of invading and causing disease of the central nervous system only when the individual is immunocompromised.56 Influenza A viruses also disrupt the immune response of the host, leading to a more severe form of the disease, and predispose the host to secondary infections from other microbes.57
Environmental influences facilitate emergence and expression
Changes in the environment may be responsible for the emergence and expression of virulence traits. For example, nutritional status may be a significant environmental variable in viral virulence. Mice infected with a strain of coxsackievirus B3, which under normal circumstances is not capable of causing disease, caused moderate to severe disease (myocarditis) when the animals were selenium deficient. Re-isolation of the virus from these animals and subsequent injection into animals with an adequate level of selenium demonstrated that the virus had been changed as a result of the exposure to altered selenium levels. In this case the virus became virulent as a result of mutations, a conclusion confirmed by genome sequencing.58
The methylation status of host nucleic acid influences the susceptibility and resistance of an organism to virus infection. For example, mice are predisposed to show a high frequency of thymic lymphomas when their nucleic acid is poorly methylated. This observation was made following the knockout of the gene regulating methylation. This change apparently led to genetic instability and activation of the retroviral elements.59 In another example, involving chickens that contained a leucosis virus integrated into chromosomal DNA, abundant methylation was associated with resistance against tumour formation.60 These and other studies indicate that methylation status has some significance in disease susceptibility and resistance. The level of methylation can be influenced by diet and environmental factors such as the presence of toxicants. For example, cancer patients typically show unusual changes in DNA methylation (lower levels) that predispose heritable material to instability and mutations. Air pollution is one factor contributing to these changes.61
Expression may also be influenced by ambient environmental conditions. Studies on the incidence of viral disease (gastrointestinal, respiratory, and vector borne) have shown that many of them display a marked seasonality with a peak in winter.62 Both the pathogen and the host immune system are influenced by cold weather conditions. Exposure to cold or induced hypothermia increases the risk of upper and lower respiratory tract infections. Suppression of the body’s immune responses and the reactions of the respiratory tract membranes have been associated with the increased susceptibility. This outcome is due to the restriction of blood supply to the respiratory system surface tissues caused by body surface cooling. This resulted in fewer white cells being available in this area to fight infection. The impact of low temperatures is most acutely felt by the young and the elderly and, not surprisingly, the risk of infection increases with exposure duration.63
Coinfection of a host by one pathogen may predispose it to severe infection by another. For example, the presence of active bacterial pathogens may predispose the host to a more severe form of influenza A virus infection.18
Conclusions
The emergence of pathogenic viruses in feeling organisms can be accounted for using the creation of viruses in non-sentient forms of life as the starting point. In these simple forms of life, viruses have been shown to have a number of benign to useful and seemingly indispensable functions along with population controlling roles.
Cells in prokaryote (bacteria) and eukaryote organisms possess genes in structures other than the chromosome. Particularly in bacteria, movement of genetic information between chromosome, plasmids, and phages has been documented. This introduces the possibility for novel combinations of genetic material. The phages in turn may have been responsible for the generation of some dsDNA and dsRNA viruses. Recombination events are possible between RNA and DNA viruses, which raises the possibility that recombination could occur between plasmid DNA and RNA viruses, hence leading to additional viral lines.
Transposable pieces of DNA are commonly found in various cells. They contain proteins similar to those found in some viruses that suggest they could have given rise to the necessary virus capsid proteins. Transposable elements also show a number of sequence similarities to information held by plasmids and phages.
The involvement of retroelements in virus emergence cannot be overestimated, as such elements are abundant in various genomes of higher organisms. These elements are incorporated into the cell DNA after transcription from a RNA molecule. They may also behave as plasmids. Retrotransposons may be released from cells under stress conditions in microvesicles that are classically generated by host cells. Some of the retrotransposons possess the capacity to form envelope-type proteins, which are a suitable starting point for virus protein envelope construction. These observations and others indicate that retroelements possibly have played a significant role in the generation of retroviruses.
Transfer of information across the species barrier may be facilitated by invertebrates and pathogens, such as fungi. Mixing of genetic information may occur in these environments when viruses are capable of multiplying. Exchange of genetic information may also occur in amoebal-type organisms and human milk, which can contain an admixture of genetic information and microbes.
Genes conceivably can be acquired from the host organism and incorporated into the viral genome permitting new combinations to emerge. Then again it is known that recombination among related viruses can lead to the emergence of new variants. Changes in the status of a virus particle from harmless to disease-causing may be associated also with both host changes and environmental influences.
While there are obvious gaps in our knowledge, the scenarios painted are reasonable from a scientific viewpoint. Those who believe that God created the fundamental lifeforms and biological entities have an additional piece of information to add to the puzzle on origins. Fundamental changes in the fidelity of cellular events following the Fall are indicated in Genesis and the account given by patriarch Job indicates the involvement of malevolent intelligent agencies in the generation of pathogens. Direct evidence for this latter proposition cannot be guaranteed, but human endeavours have shown that novel structures, as spoken about in Genesis, can be generated by both conventional and modern genetic manipulation techniques. The account given by Job on the generation of a pathogenic bacterium is not an outlandish proposition for modern genetic engineers. This means that in the end, faith must be placed in the biblical account or the miracles needed64to bridge the gap between non-life and the generation of living cells propounded by those who choose not to believe.
References and notes
- Koonin, E.V., Dolja, V.V. and Krupovic, M., Origins and evolution of viruses of eukaryotes: the ultimate modularity, Virology 479:2–25, 2015. Return to text.
- Gillen, A., Microbes and the days of creation, Answers Research J. 1:7–10, 2008; Shipton, W.A., Thorns also and thistles, J. Adventist Theological Society 23(1):18–45, 2012. Return to text.
- Koonin, E.V., Senkevich, T.G. and Dolja, V.V., The ancient virus world and evolution of cells, Biology Direct 1:29, 2006 | doi:10.1186/1745-6150-1-29; Krisch, H. M., The view from Les Treilles on the origins, evolution and diversity of viruses, Research in Microbiology 154(4):227–229, 2003. Return to text.
- Bergman, J., Did God make pathogenic viruses? Creation Ex Nihilo Technical J. 13(1):115–125, 1999; Francis, J. W., The organosubstrate of life: a creationist perspective of microbes and viruses, Proceedings of the Fifth International Conference on Creationism, pp. 433–444, 2003, answersingenesis.org/articles/aid/v4/n1/organosubstrate-of-life. Return to text.
- Márques, L.M., Redman, R.S., Rodriguez, R.J. et al., A virus in a fungus in a plant: three-way symbiosis required for thermal tolerance, Science 315(5811):513–515, 2007; Roossinck, M.J., The good viruses: viral mutualistic symbioses, Nature Reviews Microbiology 9:99–108, 2011. Return to text.
- Renault, S., Stasiak, K., Federici, B. et al., Commensal and mutualistic relationships of reoviruses with their parasitoid wasp hosts, J. Insect Physiology 51(2):137–148, 2005; Roossinck, ref. 5; Schaack, S., Gilbert, C. and Feschotte, C., Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution, Trends in Ecology and Evolution 25(9):537–546, 2010; Stasiak, K., Renault, S., Federici, B.A. et al., Characteristics of pathogenic and mutualistic relationships of ascoviruses in field populations of parasitoid wasps, J. Insect Physiology 51(2):103–115, 2005. Return to text.
- Weiss, R.A., On the concept and elucidation of endogenous retroviruses, Philosophical Transactions of the Royal Society B, London 368:20120494, 2012 | doi:10.1098/rstb.2012.0494. Return to text.
- Wessner, D.R., The origin of viruses, Nature Education 3(9):37, 2010. Return to text.
- Roossinck, M.J., Lifestyles of plant viruses, Philosophical Transactions of the Royal Society B 365(1548):1899–1905, 2010. Return to text.
- Noda, H. and Nakashima, N., Non-pathogenic reoviruses of leafhoppers and plant hoppers, Seminars in Virology 6:109–116, 1995. Return to text.
- Ackermann, H.-W., Bacteriophage observations and evolution, Research in Microbiology 154:245–251, 2003; Diallo, M.D., Monteil, C L., Vinatzer, B.A. et al., Pseudomonas syringae naturally lacking the canonical type III secretion system are ubiquitous in non-agricultural habitats, are phylogenetically diverse and can be pathogenic, ISME J. 6:1325–1335, 2012; Matos, R.C., Lapaque, N., Rigottier-Gios, L. et al., Enterococcus faecalis prophage dynamics and contributions to pathogenic traits, Plos Genetics June 6, 2013 | doi:10.1371/journal.pgen.1003539. Return to text.
- Briani, F., Dehò, G., Forti, F. et al., The plasmid status of satellite bacteriophage P4, Plasmid 45(1):1–17, 2001; Goldstein, R., Sedivy, J. and Ljungquist, E., Propagation of satellite phage P4 as a plasmid, Proceedings of the National Academy of Sciences USA 79:515–519, 1982; Roossinck, ref. 5. Return to text.
- Brand, L., What are the limits of death in paradise? J. Adventist Theological Society 14(1):74–85, 2003. Return to text.
- Deininger, P.L., Moran, J.V., Batzer, M.A. et al., Mobile elements and mammalian genome evolution, Current Opinion in Genetics and Development 13:651–658, 2003; Solyom, S. and Kazazian, H.H., Mobile elements in the human genome: implications for disease, Genome Medicine 4:12, 2012 | doi:10.1186/gm311; Temin, H.M., Origin of retroviruses from cellular moveable genetic elements, Cell 21(3):599–600, 1980. Return to text.
- Gorbalenya, A.E., Origin of RNA viral genomes, in: Gibbs, A., Calisher, C.H. and García-Arenal, F. (Eds.), Molecular Basis of Virus Evolution, Cambridge University Press, Cambridge, 1995, pp. 49–66. Return to text.
- Mayo, M.A. and Jolly, C.A., The 5-terminal sequence of potato leafroll virus RNA: evidence of recombination between virus and host RNA, J. General Virology 72(10):2591–2595, 1991. Return to text.
- Allison, R.F., Schneider, W.L. and Greene, A.E., Recombination in transgenic plants, Seminars in Virology 7:417–422, 1996. Return to text.
- Taubenberger, J.K. and Morens, D.M., Influenza viruses: breaking all the rules, mBio 4(4):e00365-13 | doi:10.1128/mBio.00365-13. Return to text.
- Tarlinton, R.E., Meers, J. and Young, P.R., Retroviral invasion of the koala genome. Nature 442:79–81, 2006. Return to text.
- Malik, H.S., Henikoff, S. and Eickbush, T.H., Poised for contagion: evolutionary origins of the infectious abilities of invertebrate retroviruses, Genome Research 10(9):1307–1318, 2000. Return to text.
- Lerat, E. and Capy, P., Retrotransposons and retroviruses: analysis of the envelope gene, Molecular Biology and Evolution 16(9):1198–1207, 1999. Return to text.
- Meckes, D.G. and Raab-Traub, N., Microvesicles and viral infection, J. Virology 85(24):12844–12854, 2011. Return to text.
- Schorey, J.S., Cheng, Y., Singh, P.P. et al., Exosomes and other intracellular vesicles in host-pathogen interactions, EMBO reports 16( 1):24–43, 2015. Return to text.
- Balaj, L., Lessard, R., Dai, L. et al., Tumor microvesicles contain retrotransposon elements and amplified oncogene sequences, Nature Communications 2:180, 2011 | doi:10.1038/ncomms1180. Return to text.
- Yin, S-d., Bacteria, viruses, membrane-enclosed microentities, and fungi as the environmental evolutionary entities coexisting in human milk, J. Theoretical Fimpology 2(2):e-20120718-2-2-11, 2014. Return to text.
- Rigden, J.E., Dry, I.B., Krake, L.R. et al., Plant virus DNA replication processes in Agrobacterium: insight into the origins of geminiviruses, Proceedings of the National Academy of Sciences USA 93(19):10280–10284, 1996; Wang, W.C., Hsu, Y.H., Lin, N.S. et al., A novel prokaryotic promoter identified in the genome of some monopartite begomoviruses, Plos One 8(7):e70037, 2013. Return to text.
- Krupovic, M., Ravantti, J.J. and Bamford, D.H., Geminiviruses: a tale of a plasmid becoming a virus, BMC Evolutionary Biology 8:112, 2009; Stedman, K., Mechanisms for RNA capture by ssDNA viruses: grand theft RNA, J. Molecular Evolution 76(6):359–364, 2013. Return to text.
- Kapitonov, V.V. and Jurka, J., Self-synthesizing DNA transposons, Proceedings of the National Academy of Science USA 103(12):4540–4545, 2006. Return to text.
- Krupovic, M. and Koonin, E.V., Polintons: a hotbed of eukaryotic virus, transposon and plasmid evolution, Nature Reviews Microbiology 13(1):105–115, 2015. Return to text.
- Katzourakis, A. and Aswald, A., The origins of giant viruses, virophages and their relatives in host genomes, BMC Biology 12:51, 2014 | doi:10.1186/s12915-014-0051-y. Return to text.
- De Silva, F.S. and Moss, B., Origin-independent plasmid replication occurs in vaccinia virus cytoplasmic factories and requires all five known poxvirus replication factors, Virology J. 2:23, 2005 | doi:10.1186/1743-422X-2-23. Return to text.
- Moreira, D. and Brochier-Armanet, C., Giant viruses, giant chimeras: the multiple evolutionary histories of mimivirus genes, BMC Evolutionary Biology 8:12, 2008 | doi:10.1186/1471-2148-8-12. Return to text.
- Koonin, E.V., On the origin of cells and viruses, Annals of the New York Academy of Sciences 1178:47–64, 2009. Return to text.
- Bamford, D.H., Do viruses form lineages across different domains of life? Research in Microbiology 154:231–236, 2003. Return to text.
- Briani, F., Dehò, G., Forti, F. et al., The plasmid status of satellite bacteriophage P4, Plasmid 45(1):1–17, 2001; Goldstein, R., Sedivy, J. and Ljungquist, E., Propagation of satellite phage P4 as a plasmid, Proceedings of the National Academy of Sciences USA 79(2):515–519, 1982. Return to text.
- Chavan, M., Rafi, H., Wertz, J. et al., Phage associated bacteriocins reveal a novel mechanism for bacteriocin diversification in Klebsiella, J. Molecular Evolution 60(4):546–556, 2005; Šmarda, J. and Benada, O., Phage tail-like (high molecular weight) bacteriocins of Budvicia aquatica and Pragia fontium (Enterobacteriaceae), Applied and Environmental Microbiology 71(12):8970–8973, 2005. Return to text.
- Brunovskis, P. and Kung, H-J., Retrotransposition and herpesvirus evolution, Virus Genes 11(2–3):259–270, 1996. Return to text.
- Esteller, M. and Herman, J.G., Cancer as an epigenetic disease: DNA methylation and chromatin alterations in human tumours, J. Pathology 196(1):1–7, 2002; Herceg, Z., Epigenetics and cancer: towards an evaluation of the impact of environmental and dietary factors, Mutagenesis 22(2):91–103, 2007. Return to text.
- Gupta, P.K., Biotechnology and Genomics, Rastogi Publications, New Delhi, pp. 459–480, 2009. Return to text.
- Gibson, D.G., Benders, G.A., Andrews-Pfannkoch, C. et al., Complete chemical synthesis, assembly, and cloning of Mycoplasma genitalium genome, Science 319(5867):1215–1220, 2008; Pennisi, E., Synthetic genome brings new life to bacterium, Science 328(5981):958–959, 2010. Return to text.
- Dhandayuthapani, S., Blaylock, M.W., Bebear, C.M. et al., Peptide methionine sulfoxide reductase (MsrA) is a virulence determinant in Mycoplasma genitalium, J. Bacteriology 183(19):5645–5650, 2001. Return to text.
- Galambos, A., Zok, A., Kuczmog, A. et al., Silencing Agrobacterium oncogenes in transgenic grapevine results in strain-specific crown gall resistance, Plant Cell Reports 32(11):1751–1757, 2013. Return to text.
- Gi Na, L. and Na, J., The impact of synthetic biology, ACS Synthetic Biology 2:210–212, 2013. Return to text.
- Herfst, S., Schrauwen, E.J.A., Linster, M. et al., Airborne transmission of influenza A/H5N1 virus between ferrets, Science 336(6088):1534–1541, 2012; Saey, T.H., Designer flu, Science News 181(11):20–25, 2011. Return to text.
- Jackson, R.J., Ramsay, A.J., Christensen, C.D. et al., Expression of mouse interleukin-4 by a recombinant ectromelia virus suppresses cytolytic lymhphocyte responses and overcomes genetic resistance to mousepox, J. Virology 75(3):1205–1210, 2001; Müllbacher, A. and Lobigs, M., Creation of killer poxvirus could have been predicted, J. Virology 75(18):8353–8355, 2001. Return to text.
- Geuking, M.B., Weber, J., Dewannieux, M. et al., Recombination of retrotransposon and exogenous RNA virus results in nonretroviral cDNA integration, Science 323(5912):393–396, 2009. Return to text.
- Landen, M.G., McCumber, B.J., Asay, E.D. et al., Outbreak of boils in an Alaskan village, Western J. Medicine 172:235–239, 2000; Wang, J., Barth, S., Richardson, M. et al., An outbreak of methicillin-resistant Staphylococcus aureus cutaneous infection in a saturation diving facility, Undersea Hyperbaric Medicine 30:277–284, 2003. Return to text.
- Bogdanov, Y.F., Fedotova, Y.S., Sosnikhina, S.P. et al., Bar- and thorn-like abnormalities in synaptonemal complexes of a mutant rye, Secale cereale. Genome 41(2):284–288, 1998—thorn-like abnormalities arise through mutation; Lester R.N. and Daunay, M.-C., Diversity of African vegetable Solanum species and its implications for a better understanding of plant domestication, Proceedings of a Symposium Dedicated to the 100th Birthday of Rudolf Mansfield, Gatersleben, Germany, 8-9 October, 2001, Schriften zu Genetischen Ressourcen, 22:136–152, 2001—prickles appear when certain crosses are made; Lester, R.N. and Thitaiz, G.N.W., Inheritance in Solanum aethiopicum, the scarlet eggplant, Euphytica 40:67–74, 1989—reverse mutation noted causing prickles to be lost; Marcotrigiano, M., Herbivory could unlock mutations sequestered in stratified shoot apices of genetic mosaics, American J. Botany 87:355–361, 2000—prickle expression fails to occur because of a mutation in a developmental pathway. Return to text.
- Bletson, F., Roupakias, D., Tsaksira, M. et al., Production and characterization of interspecific hybrids between three eggplant (Solanum melongena L.) cultivars and Solanum macrocarpon L., Scientia Horticulturae 101(1-2):11–21, 2004; Okamura, M., Pomato: Potato protoplast system and somatic hybridization between potato and wild tomato; in: Bajaj, Y.P.S. (Ed.), Somatic Hybridization in Crop Improvement, Springer-Verlag, Heidelberg, 1:209–223, 1994. Return to text.
- Riedel, S., Biological warfare and bioterrorism: a historical review, Proceedings (Baylor University Medical Center) 17(4):400–406, 2004. Return to text.
- Casadevall, A. and Pirofski, L., Host–pathogen interactions: redefining the basic concepts of virulence and pathogenicity, Infection and Immunity 67(8):3703–3713, 1999; Casadevall, A. and Pirofski, L., Host–pathogen interactions: the attributes of virulence, J. Infectious Diseases 184(3):337–344, 2001. Return to text.
- Jern, P. and Coffin, J.M., Effects of retroviruses on host genome function, Annual Review of Genetics 42:709–732, 2008. Return to text.
- Schaack et al., ref. 6; Shadan, F.F. and Villarreal, L.P., The evolution of small DNA viruses of eukaryotes: past and present considerations, Virus Genes 11(2–3):239–257, 1996. Return to text.
- Bibollet-Ruche, F., Heigele, A., Keele, B.F. et al., Efficient SIVcpz replication in human lymphoid tissue requires viral matrix protein adaptation, J. Clinical Investigation 122(5):1644–1652, 2012; Mourez, T., Simon, F. and Plantier, J-C., Non-M variants of human immunodeficiency virus type 1, Clinical Microbiology Reviews 26(3):448–461, 2013. Return to text.
- Benson, S.J., Ruis, B.L., Fadly, A.M. et al., The unique envelope gene of subgroup J avian leukosis virus derives from ev/J proviruses, a novel family of avian endogenous viruses, J. Virology 72(12):10157–10164, 1998. Return to text.
- Sundaram, C., Shankar, S.K., Thong, W.K. et al., Pathology and diagnosis of central nervous system infections, Pathology Research International, 2011, article ID 878263, 4 pages. Return to text.
- Chertow, D.S., Contribution of bacterial coinfection to severe influenza infection, Critical Care Medicine 40(5):1664–1665, 2012. Return to text.
- Beck, M.A., Handy, J. and Levander, O.A., Host nutritional status: the neglected virulence factor, Trends in Microbiology 12(9):417–423, 2004. Return to text.
- Howard, G., Eiges, R., Gaudet, F. et al., Activation and transposition of endogenous retroviral elements in hypomethylation induced tumors in mice, Oncogene 27(3):404–408, 2008. Return to text.
- Yu, Y., Zhang, H., Tian, F. et al., Quantitative evaluation of DNA methylation patterns for ALVE and TVB genes in a neoplastic disease susceptible and resistant chicken model, Plos One March 5, 2008 | doi.org/10.1371/journal.pone.0001731. Return to text.
- Barchitta, M., Quattrocchi, A., Adornetto, V. et al., Association between LINE-1 hypomethylation and cancer risk: a systematic review and meta-analysis, European J. Public Health 24(Suppl. 2):62, 2014; Bollati, V. and Baccarelli, A., Environmental epigenetics, Heredity 105:105–112, 2010. Return to text.
- Fisman, D., Seasonality of viral infections: mechanisms and unknowns, Clinical Microbiology and Infection 18(10):946–954, 2012. Return to text.
- Mourtzoukou, E.G. and Falagas, M.E., Exposure to cold and respiratory tract infections, International J. Tuberculosis and Lung Diseases 11(9): 938–943, 2007. Return to text.
- Krisch, ref. 3, nominated six ‘miracles’ or ‘miraculous’ events. Return to text.





Readers’ comments
Comments are automatically closed 14 days after publication.