Journal of Creation 31(2):94–102, August 2017
Browse our latest digital issue Subscribe
The irreducibly complex ribosome is a unique creation in the three domains of life
The evolution of the genetic code and the ribosome are intimately connected as the code is expressed through ribosomal activity. Models of genetic code evolution are analyzed. The error-minimization theory is faulty in that it supposes that highly error-prone genetic codes could produce more precise codes over time. The stereochemical theory posits complementarity between nucleotides and amino acids, but cannot demonstrate this for the whole code. The co-evolution theory states that the genetic code developed from an ancestral through an ancient to a modern state. There is no evidence for ancestral code protein generation. The big question remains why the code solidified in its present state. Finally, the accretion theory of ribosomal evolution is shown incapable of answering key questions.
Large and small subunit ribosome proteins are conserved within Archaea, Bacteria, and Eukarya but not between these domains. The size and weight of the subunit proteins are similar between Archaea and Bacteria only, whereas protein types are similar only between Archaea and Eukarya. This implies that the ribosome of all three domains has been created uniquely. The presence of many unique proteins and protein domains in the mitochondrial and chloroplast ribosomal proteins imply that they are not related to prokaryotic ones.
Protein synthesis is a fundamental function in the cell that involves ribosomes. The process involves first the transcription of the DNA sequence to messenger RNA (mRNA). Each sequence of three bases in mRNA is known as a codon. The information contained in the codon is used to produce functional proteins, as each codon specifies a particular amino acid. The vital step in protein formation occurs on the ribosomes with the cooperation of transfer RNA (tRNA) using ribosomal RNA (rRNA) as a binding site. It is evident that ribosomes are prerequisites to the life of the cell in that they convert genetic information into functional proteins. These structures consist of two different sized subunits, whose size is described in terms of Svedberg units (S), which is a measure of the sedimentation rate. Ribosomal proteins dominate these subunit structures, but there are up to 120 different molecules involved: rRNA, mRNAs, tRNAs, ribosomal proteins, aminoacyl-synthetases, and scanning factors. They are all needed to fulfil this basic, yet highly complex cellular housekeeping function. Besides histones (DNA packaging proteins in eukaryotes), ribosomal proteins are the most conserved proteins in the living world.1

The coupling of the protein-translation machinery to the DNA is fulfilled in the genetic code, which consists of four nucleotide bases arranged in groups of three (codon). The codons can be arranged in 64 combinations allowing selection of the 20 amino acids; special codons mark the start and stop point for a protein. Some evolutionists believe that the ribosomes came into being before cellular life and represent the first self-replicating entities. This means the ribosomal RNA they carry is a primitive genome.2
Here we will take a look at one of the greatest problems in biology. Classically, living systems produce copies of themselves. In ancestral systems, evolutionists consider that the descendants were different from their immediate ancestor as they needed to generate coding rules that then went on to evolve new systems.3,4
Evolutionary theories on the genetic code and the origin of the ribosome
There are up to 22 small ribosomal subunits in Escherichia coli, and up to 35 large subunit (LSU) proteins,5 whereas there are 33 small subunit (SSU) proteins in the human ribosome, compared to 47–48 proteins in the LSU. Across the eukaryotes, the number of ribosome proteins may not be constant.6 An added complication is that 70S ribosomes are found associated with the plastids (some eukaryotes). Whereas there are similarities in the 70S ribosomes between prokaryotes and eukaryotes, in the machinery associated with translation there are fundamental differences.7 Other ribosome types have been found in the mitochondria that show many differences in contrast to bacterial ribosomes.8
Our focus is on the evolution of the genetic code and the classical 70S prokaryote and 80S eukaryote ribosomes. It is a paradox of evolution that the composition of the prokaryotic ribosome is different to that of the eukaryotic one, yet the ribosome has supposedly evolved through a number of intermediary steps back into a ribosome. Evolution simply loses all meaning if a protein or set of proteins evolves into a structure, which fulfils the same purpose, which it started out from.
Today there are three major theories on the origin of the genetic code:4,9 the error-minimalization theory,10 the stereochemical theory,11 and the co-evolution theory.12 The recently enunciated accretion theory of ribosomal evolution assumes that the genetic code is also evolving since it supposedly accounts for rRNA, mRNA as well as tRNA changes.13 Other theories have been proposed,4,14,15 but these are not discussed except for the accretion model.
The error-minimization or adaptation theory
According to Sonneborn’s argument reviewed by Carl Woese,10 selection pressure acted on a primitive genetic code that led to the generation of a mature genetic code where mutations in codons produced few adverse outcomes in terms of functional proteins. This represents an error-minimization strategy. Woese admitted that the error-minimization scheme involved innumerable “trials and errors” so that it, in his opinion, “could never have evolved in this way”.
Others have defended the theory. Some ingenious ideas were admitted subsequently as “utterly wrong”. Interestingly, one investigation of the theoretical susceptibility of a million randomly generated codes to errors, through mutations, showed that the standard genetic code was among the least prone to error.16 This indicated that, if the initial genetic code was primitive and error prone, then what is observed in nature is the best option. However, the question remains as to why only one single code survived. Why not several different ones? This rather stands as evidence that the Creator made a wise choice.
The ancestral translational machinery conceived in evolutionary schemes is, of necessity, very rudimentary, and thus highly prone to errors. This means that it would have been almost impossible to correctly translate any mRNA, and thus produced little more than statistical proteins (proteins with only random sequences). Yet through necessity, somehow, the codons of the ancestral code were gradually reconfigured in order to minimize translational error.4 The ‘somehow’ has been imagined as perhaps involving novel amino acids, existence of a positive feedback mechanism that would assign codons to amino acids with similar properties, direct templating between nucleic and amino acids, or other possibilities.16
Vetsigian and Woese 17 subsequently proposed that horizontal gene transfer (HGT) could possibly spread workable genetic workable codes across organisms, accounting for the near universality of the genetic code. However, HGT requires that the genetic codes of the host and the recipient species be similar enough for the new genetic code to work. There also needs to be evidence for a mechanism permitting transfer of genetic information in the ancient past.
The theory naturally cannot carry much weight, since if the translation machinery is so error-prone to begin with, no meaningful proteins can come from such a configuration. Errors only lead to more errors, not higher precision, which requires intelligent input. From a thermodynamic viewpoint, disorder only increases as mutations accrue.
The stereochemical theory
Over the past 60 years, several theories have been set forward which attempt to explain how information in the DNA translates to protein sequences. These are based on some sort of selective stereochemical complementarity or affinity between amino acids and nucleotides (base pair triplets). On a physico-chemical level, this is based on the negative charges of the nucleotide phosphates interacting with the positive charge of the basic amino acids. In Saxinger et al.’s study no conclusive selective binding occurred between certain amino acids and nucleotide triplets.18 More recently, Yarus et al.19 contended that coding triplets arose as essential parts of RNA-like amino acid-binding sites, but they could show this for only seven of the 20 (35%) canonical amino acids. However, they conceded that the code can change.
The take home implication is that different amino acids can be bound by different coding triplets, meaning that the code is not specific and thus meaningless historically. Overall, after decades of research, no evidence has been found which gives strong support to the stereochemical theory. Yarus’s group19 went on to argue that adaptation, stereochemical features and co-evolutionary changes were compatible and perhaps necessary in order to account for present codon characteristics. However, Barbieri3 has argued that there is “no real evidence in favour” of the stereochemical theory. This serves to illustrate the uncertainty prevailing.
The co-evolution theory
According to the co-evolution theory, the original genetic code was “excessively degenerate” meaning it could code for several amino acids. These originals were used in “inventive biosynthetic processes” to synthesize the other amino acids. The code then adapted to accommodate these new amino acids.20 Similarities in the codons of related amino acids were subject to computer analysis in order to determine if a better code could be found based on biosynthetically related amino acids. An extraordinary correlation was noted for the universal code, as against 32,000 randomly generated possibilities. Changing the pattern of relatedness among amino acids gave more codes equal to or of greater correlations than the universal code. However, the authors stated that these observations “cannot be used as proof for the biosynthetic theory of the genetic code”.21
Less than half of the 20 canonical amino acids found in proteins can be synthesized from inorganic molecules.3 Furthermore, the amino acids that are missing (the so-called secondary amino acids) are also missing from material recovered from meteorites.22 This is problematic for evolution, for it implies that early life-forms on this planet could only use ten amino acids for protein construction, something which we don’t observe today, thereby greatly reducing the possible number of functional proteins.
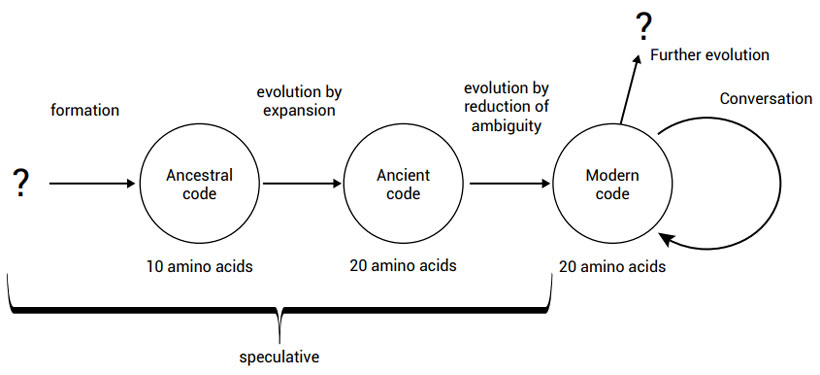
The primary amino acids were coded by an ancestral genetic code, which then expanded to include all 20 canonical amino acids. The present code is a non-random structure yet it is more robust as far as translational errors are concerned than the majority of alternative codes that can be generated conceptually according to accepted evolutionary trajectories. When the starting assumptions are altered so that the postulated codes start from an advantaged position, then higher levels of robustness are achieved. A better code could have been produced if evolution had continued, but it did not as the possibility of severe adverse effects was too great.23 This process can be seen in figure 1.
Several questions present themselves here, however. Why don’t we find any protein sequences in the fossils of ancient organisms, which only have primary amino acids? The fact that no such proteins exist is strong proof against the evolutionary origin of the genetic code. We only find proteins made up of all 20 amino acids. Why didn’t the genetic code keep on expanding to cover more than 20 amino acids? Why not 39, 48 or 62? Why did codon triplets evolve, and why not quadruplets? With 44 = 256 possible codon quadruplets, coding space could have increased, and thus a much larger universe of possible proteins could have been made possible.
An additional fundamental issue is that if life commenced in an RNA world, then amino acids could have been synthesized on the primitive codons associated with these molecules by primordial synthetases. How do similar coding rules now apply when codon recognition is performed by the anticodons of the tRNA with the assistance of the highly specific aminoacyl-tRNA-synthetases that attach to the amino acids? It has been suggested that perhaps there was a two-base code rather than a three-base one on account of the supposed limited number of amino acids available.4,24
The accretion model of ribosomal evolution
The accretion model of ribosomal evolution13,25 is one of the most recent models and describes how the ribosome evolves from simple RNA and protein elements into an organelle complex in six major phases through accretion, recursively adding, iterative processes, subsuming and freezing segments of the rRNA. It is argued that the record of changes is held in rRNA secondary and three-dimensional structures. Patterns observed in extant rRNA found among organisms were used to generate rules supposedly governing the changes.
First, it is assumed that evolution occurred with changes moving from prokaryotes leading finally to the eukaryotes and with the apex reached with humans. Using this framework, a chronological sequence was constructed of rRNA segment additions to the core structure found in Escherichia coli. The six-phase process envisaged provided no evidence for the emergence of ancestral RNA. The proto-mRNA is seen simply as arising from a random population of appropriate molecules. This proto-mRNA together with tRNA, formed through condensation of a cysteine: cysteine: alanine (CCA) sequence unit, gave rise to base-pair coding triplets (codons). The ribosomal units (small and large) are considered to have arisen from loops of the rRNA. The proposed RNA loops were ‘defect-laden’, which required a protection mechanism. During phase 2 the large ribosomal unit is thought of as a crude ribozyme almost as soon as it was a recognizable structure, catalyzing nonspecific, non-coded condensation of amino acids. Finally, the two developing ribosome units came together (phase 4) to form a complex structure recognizable as a ribosome. In the next phase (5), specific interactions began to occur between anticodons in tRNA and mRNA codons to produce functional proteins. In the final phase the genetic code was optimized.13
This narrative suffers from major flaws, some of which also are inherent in previous models of the genetic code generation. No organisms have been found that contain ribosomes in any of these intermediary phases. If these intermediary phases are capable of ribosomal function, then why was it necessary to evolve further during additional steps? An insistent problem is how a genetic code could be generated that depends for its expression on proteins that can only be formed when it exists. Petrov et al.25 proposed a partial solution. The peptidyl transferase (enzyme) centre, an essential component of the ribosome, arose from an rRNA fragment. This means that its origin is conceived of as being in the RNA world.26 The peptidyl transferase centre is the place in the 50S LSU where peptide bond synthesis occurs. The machinery is very complex in extant organisms. In its original incarnation, the embryonic centre was less than 100 nucleotides long. The original RNA world quickly morphed into the familiar RNA/protein world. This argument is necessary as it “has proven experimentally difficult to achieve” a self-replicating RNA system. In a revealing aside, Fox even suggested that perhaps it is not necessary to validate the existence of the RNA world if it had a short life.26
Some of the additional problems with an RNA world origin were noted by Strobel.27 An RNA commencement to life on Earth rests on the ability of RNA to both share the task of encoding and also to replicate information. This proposition depends on the abilities of RNA copying enzymes (ribozymes). However, such enzymes are unable to copy long templates and at a sufficient rate to overtake decomposition processes. Even greater issues are that there is no sensible resolution to the question of the origin of the activated nucleotides through abiotic processes needed for RNA formation, or of the problem as to how randomly assembled nucleotides achieved the ability to replicate. This has led some to conclude that “the model does not appear to be very plausible”. Nevertheless, undaunted, other possibilities have been invented.28
The irreducibly complex character and conservation of the ribosome
The foregoing discussion leads us to conclude that the ribosome in itself is an irreducibly complex cellular organelle, requiring several dozen proteins to be present at the same time in order for it to work. Furthermore, the molecular machinery that regulates ribosomal gene co-expression involves just under 300 transcription regulators, which is also further modulated according to several cell types in humans and mice.29 The ribosome is also a prime example of the evolutionary paradox of sequence conservation of both functional genes and regulatory sequences.30 Among the approximately 60 proteins that are represented by an ortholog (gene in different species retaining the same function) in every single cellular lifeform with a sequenced genome, over 50 are components of the translation machinery.31 Some of these ribosomal protein genes (RPGs) are necessary for function or required for self-assembly, whereas others can be used for stimulation of the translation process.4 Ribosomal dysfunction is in the background of at least a dozen diseases, including some forms of cancer (table 1) (see appendix62).32 This is evidence that these proteins could not have evolved gradually over time, but are part of a complex functional unit.

Evolutionists argue that during ribosomal evolution, different proteins were co-opted from other processes. The ribosome is made up of approximately 80 proteins, around half of which play an exclusive role within the ribosome. The remainder of those studied have extra-ribosomal functions (although seven of them have not yet been studied, according to Wang et al.).33 Thus, since half of all ribosomal proteins (RPs) have no role outside of the ribosome, the co-option argument does not appear realistic.
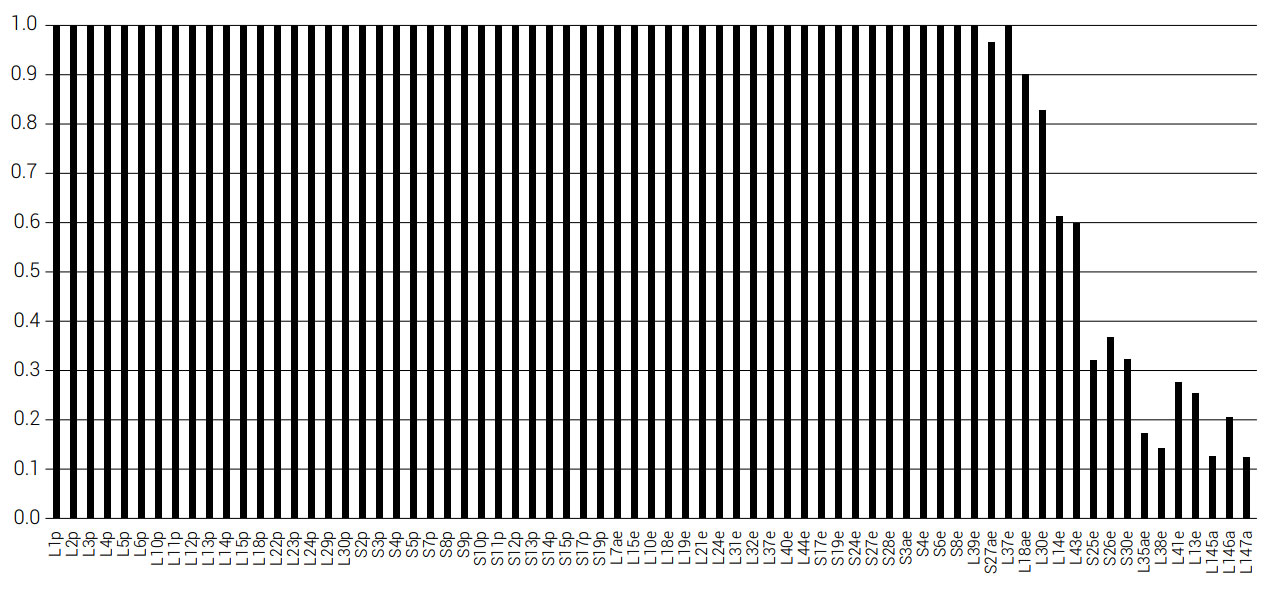
The conservation of the 80 ribosome RPs can be studied across a range of species using the Ribosomal Protein Gene Database (RPGD).34 For each protein, the number of database sequences, the number of species, and the percent conservation were noted (see materials and methods section). We found that 48.8% (39 out of 80) RPs from the database had at least 50% sequence conservation (table 2) (see appendix62). In a study of 41 of the 54 RPs from Escherichia coli, these genes were individually deleted to verify whether they are necessary for ribosomal function. Of these 41 proteins, nine (22%) were shown to be non-essential (RPs S9, S17, L15, L21, L24, L27, L29, L30, and L34).35 This is taken to mean that the ribosome performs basic functions essential for existence and that the operational design is robust hence accounting for the use of common elements. Further, the RPGD (136 RP alleles) was compared to the Saccharomyces Genome Database,36 to see what effect the null mutation (when the gene is functionally knocked out) had on each individual gene. It was found that 113 alleles showed a reduced phenotype, 22 were unviable, and one had no information. This was again taken to indicate the functional necessity of the majority of alleles. Related analyses have been extended to prokaryotes. Yutin et al.37 counted the number and distribution of RPs in a study of 995 bacterial species and 87 archaeal species. The level of conservation of the proteins is shown in figures 2 and 3. Fifty six out of 71 (78.9%) RPs were conserved in all 87 archaeal species, and 44 out of 56 (78.6%) RPs are conserved in all 995 bacterial species. Of the small and large subunit proteins, which are not universally conserved among the 995 bacterial species, S21 and L17 are found on the surface of the ribosome in E. coli, suggesting that they are free to mutate, and are thus unessential. Of these two, S21 was deleted from E. coli via λ-Red-mediated recombination.35 Only four RPs that are not present in the 995 species are not exposed on the surface of E. coli.38 Of these, S16 and L31 were present in 99.9%, L34 in 99.7%, and L30 was present in 85.4% of all 995 species. The proteins L9, L19, L25, L30, L34, and L36 within the LSU of E. coli can be seen in figure 4.

When comparing RPs across Eukarya, Archaea, and Bacteria, as well as mitochondria and chloroplasts, Mears et al.39 found that among both the SSU and the LSU, the degree of protein conservation was somewhat less, and even more so when comparing with the organelles (see table 3). Conservation of RPs in the LSU of the two organelles was as low as 23.7%, but this is on account of the comparison being made across differing domains of life. The conservation of RPs is as high as 85.7% in the SSU in Bacteria, within a single domain. The study was used to speculate about spatial changes occurring in ribosomal subunits during evolution. We do not resonate with this use of the data. It can be interpreted to mean that the ribosomes represented in the different domains of life were uniquely created and any similarity to the ribosomes of other domains was used to perform essentially similar functions. This section of our conclusion is not dissimilar to that of Mears et al.39 when they state that conserved residues “are generally found in areas that are known to be functionally significant”.
In studying ribosomal proteins, investigators have distinguished between essential and non-essential proteins. This refinement is performed to give greater insights into the supposed movement of genes during evolution. In one study the λ-Red-mediated recombination approach was used to alter the bacterial genome and study the function of 41 ribosomal subunit proteins. Nine proteins appeared to be non-essential (deletion was not lethal) according to the limited stress tests applied, but it is significant to note that complete removal of some of these ‘non-essential’ proteins led to alterations in growth and ribosomal function. This is not surprising as the ribosomal proteins are involved in conformational changes in ribosomes as well as in interactions with other components connected with translation. Further studies have added additional non-essential proteins to the list. 40 Akanuma et al.41 usefully have outlined some of the difficulties in determining whether a protein is essential or not. Fundamental is whether mutants contained the expected structure for a correct deletion. The percentage may be as low as 25%. Additional issues involve the presence of suppressor sequences that permit viability of mutants, and the presence of duplicate genes. Culture conditions may contribute to failures in correct identification, as well as technical problems involving primer design.40 Apart from these difficulties, a range of ribosomal proteins has been identified that have no apparent vital function. Whereas some of these conclusions may be correct, statements about non-essentiality will undoubtedly be revised as more sophisticated methods are used and a greater range of organisms is analyzed. One caution is merited. When investigators write about the non-essentiality of a gene, which gives rise to non-essential proteins, this does not necessarily mean that the absence of such a gene has no effect. This is illustrated nicely with Saccharomyces. The single copy of the RPL29 gene is considered nonessential. It codes for a 60S ribosomal subunit protein (Rpl29p). However, its absence retards the coupling of 60S and 40S subunits and also translation events. There are also interactions between RPL29 and essential genes.42
Taxonomic distribution of ribosomal proteins and patterns of gene loss for ribosomal proteins
Taxonomic distribution of ribosomal proteins between the three domains of life

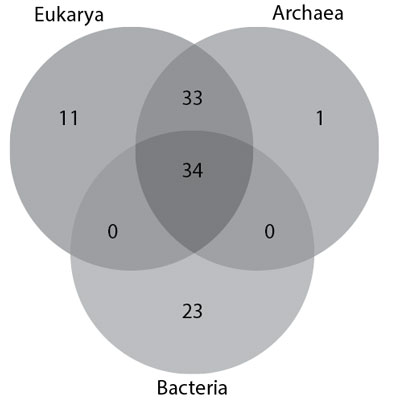
According to Fox26 and Barbieri4 the ribosome would have existed in essentially its modern form by the time of the Last Universal Common Ancestor (LUCA). However, there is variation across the three domains of life. Within each domain conservation is stronger than between domains.43 Out of in excess of a hundred ribosomal protein genes (RPGs), 32 were conserved in all three main domains of life (Eukarya, Archaea, and Bacteria), 33 are common to only Eukarya and Archaea, whereas there are no RPGs specific to Eukarya and Bacteria or Archaea and Bacteria. Twenty-three RPGs are unique to Bacteria, whereas 11 are unique to Eukarya, and only 1 to Archaea (figure 5).44 Márquez et al.45 also discovered nine RPs which hypothetically might be unique to Archaea. According to the Lecompte data, Archaea+Eukarya have a 0.7–0.71 Pearson correlation coefficient with each other, according to the absence/presence of RPs in the SSU, the LSU, or in both (table 4). The data as a whole poses a recognized problem in that no clear evolutionary scenario emerges and one is left to ponder whether prokaryotes emerged from an ancestral eukaryote genome, a bacterial-like genome or whether symbiosis was involved,42 as we will elaborate on later.
The information is noteworthy to us too, since eukaryotes have a 60S large subunit (LSU), and a 40S small subunit (SSU), whereas prokaryotes such as Archaea and Bacteria have a 50S LSU and a 30S SSU. This would imply that the ribosomes of Archaea and Bacteria are structurally different from than that of Eukaryotes. However, the absence/presence correlation of RPs discussed previously suggests that the ribosomes of Archaea and Eukarya also group away from Bacteria. This suggests to us that Archaea, Bacteria, and Eukarya are all distinctly created domains of life.
Ribosomal proteins of the mitochondrion and chloroplast
Eukaryotic mitochondrial RPs have originated from prokaryotes in the view of some evolutionists. They point as evidence to size similarity between the two types of ribosomes, as well as sequence homology. However, mitochondrial ribosomes are missing some RPs present in prokaryotes, and have unique proteins of their own, which are not homologous to bacterial RPs.46 These unique proteins replace the function of rRNAs in the prokaryotic organisms, such as the 50% of rRNAs that correspond to a RP in the protist Leishmania tarentolae.47 When ribosomal proteins of E. coli and the human mitochondrial ribosome are compared (table 5) (see appendix62), only 42 of the 88 proteins (47.7%) are common between the two types of ribosomes, with 12 being unique to E. coli, and 34 unique to the human mitoribosome. In a further example, yeast MRP51 gene encodes a novel protein responsible for global mitochondrial translation.48 Again, the function of MPS33 in Drosophila melanogaster is unknown, but its absence can cause cardiomyopathy49 and MRPL55 has been shown to be necessary for mitochondrial biogenesis.50
Similar to the mitochondrion, the chloroplast ribosome is also missing RPs compared to the prokaryotic ribosome, and also contains RPs not found in the ribosomes of prokaryotes. The RP content of E. coli and Chlamydomonas reinhardtii is compared in table 5. There is an 80.3% commonality.51 Despite the high number of common RPs, the chloroplast RPs that are in common with Bacteria also have novel protein domains. For example, a plastid-specific ribosomal protein (PSRP-3) takes part in the regulation of translation, whereas PSRP-7 assists the positioning of mRNA during translation initiation. RAP38 and RAP41 are ribosome-associated proteins, which take part in translation.52
Once again, just because prokaryotic ribosomes are superficially similar to eukaryotic mitochondrial ones, does not necessarily mean that they are descended from them.53
Loss of ribosomal protein genes (RPGs)
Evolutionists surmise that, with the exception of the LXa gene, the full complement of RPs was present in the ancestor of Archaea and Eukarya, with reductive evolution happening on the scale of a whole domain. Furthermore, this led Lecompte et al.44 to suggest that the full complement of archaeal RPs was present in the cenancestor of both Archaea and Eukarya, leading to the conclusion that the prokaryotes evolved by simplification of this ancestral eukaryotic-like genome, an idea suggested by several authors.54,55 This creates an uncomfortable dilemma for evolutionists in that they have to choose between such a theory of origin as opposed to one based on a symbiotic hypothesis involving eubacteria and methanogenic Archaea or even other possibilities.56 An added dilemma is introduced in that it is admitted that closeness of sequence similarity need not mean a close phylogenetic relationship unless there were similar rates of evolution involved with the different lineages considered.57
RPG losses are restricted to small numbers of divergent species or genera, meaning that these gene deletion events occur independently in these lineages. RPG losses tend to happen more frequently in intracellular pathogens, such as Mycoplasma genitalium, M. pneumoniae and Encephalitozoon cuniculi.
The LSU RNA of the microsporidian Encephalitozoon cuniculi is greatly reduced in length compared to other eukaryotes, and so is its SSU RNA, which is only ~1300 bp, as compared to ~1600 bp in prokaryotes and ~1800 bp in other eukaryotes. Furthermore, the internal transcribed spacer DNA 2 region (ITS2) located between the 5.8S and 28S region in eukaryotes is lacking in E. cuniculi, suggesting, to those seeking evolutionary clues to origins, that this species is one of the earliest eukaryotic species to diverge from prokaryotes.58 However, the parasitic lifestyle of this species suggests otherwise, namely that the loss of the ITS2 spacer and the smaller size of the two ribosomal RNA subunits indicates that it has been derived from other eukaryotes and not prokaryotes. Further analysis of E. cuniculi has indicated that it contains mitochondrial genes (e.g. the Fe-S cluster assembly), suggesting that this organism group arose as a result of degenerative processes.59 The clear implication is that though some prokaryotes may show similarities to eukaryotes this doesn’t mean that such features could not have come from prokaryotes.53 Conversely, certain prokaryotic features present in eukaryotes also do not necessarily mean that eukaryotes evolved from prokaryotes.
Conclusion
The ribosome is an example of sudden, early complexity, if evolution is true. Its appearance raises many speculative events, sequences, and unresolved issues,60 making such a scenario highly dubious. A number of theories surrounding the evolution of the genetic code and of ribosomal evolution have been examined, but have been found lacking in any form of convincing evidence. Rather, due to its content and intricacy of operation, the ribosome is an example of irreducible complexity, with several dozen proteins making up its functional-structural core. The ribosome differs in make-up between Archaea, Bacteria, and Eukarya, with a number of RPs which are unique to each, suggesting that these structures did not evolve from each other but rather came into being separately by special creation. Also, some of the ribosomes in prokaryotes are missing from the mitochondria and chloroplasts of eukaryotes. Novel proteins have been found. This is taken to highlight their independent creation. The existence and distribution of RPs does not make sense, except in the light of creation.
Materials and methods
Figures 2 and 3 were drawn in Excel. Figure 4 was made using the RiboVision software (Bernier et al.61). Figure 5 was made using the Venn diagram software at bioinformatics.psb.ugent.be/webtools/Venn/. The multiple alignment of all 80 eukaryotic RPs were downloaded from the Ribosomal Protein Gene Database at ribosome.med.miyazaki-u.ac.jp/. A perl script was used to calculate the degree of conservation in each of the proteins. A position in the multiple alignment was taken to be conserved if any given amino acid was present in at least 80% of the sequences. Only those positions were counted where at least 90% of the characters were not a gap character (“-”). The data for table 1 was taken from Narla et al.32 and Wang et al.33 Deletion information was taken from the Saccharomyces Genome Database36. Ribosomal protein annotation for Chlamydomonas reinhardtii was taken in part from the Joint Genome Institute (JGI) at genome.jgi.doe.gov/Chlre3/Chlre3.download.ftp.html.
References and notes
- Black, J.G., Microbiology, 8th edn, John Wiley & Sons, Singapore, pp. 167–172, 2013; Woese, C.R., Interpreting the universal phylogenetic tree, Proceedings of the National Academy of Sciences USA (PNAS) 97(15):8392–8396, 2000. Return to text.
- Root-Bernstein, M. and Root-Berstein, R., The ribosome as a missing link in the evolution of life, J. Theoretical Biology 367:130–158, 2015. Return to text.
- Barbieri, M., Codepoiesis—the deep logic of life, Biosemiotics 5:297–299, 2012. Return to text.
- Barbieri, M., Evolution of the genetic code: the ribosome-oriented model, Biological Theory 10:301–210, 2015. Return to text.
- Wang, X., NMR study of the Escherichia coli 70S ribosome particles using residual dipolar couplings, PhD. thesis, University of California, LA, 2012. Return to text.
- Khatter, H., Myasnikov, A.G., Natchiar, S.K., and Klaholz. B.P., Structure of the human 80S ribosome, Nature 520(7549):640–645, 2015. Return to text.
- Yamaguchi, K. and Subramanian, A.R., The plastid ribosomal proteins. Identification of all the proteins in the 50 S subunit of an organelle ribosome (chloroplast), J. Biological Chemistry 275(37):28466–28482, 2000. Return to text.
- O’Brien, T.W., Properties of human mitochondrial ribosomes, IUBMB Life 55(9):505–13, 2003. Return to text.
- Koonin, E.V. and Novozhilov, A.S., Origin and evolution of the genetic code: the universal enigma, IUBMB Life 61(2):99–111, 2009. Return to text.
- Woese, C.R., Order in the genetic code, PNAS 54(1):71–75, 1965. Return to text.
- Gamow, G., Possible relation between deoxyribonucleic acid and protein structures, Nature 173(4398):318, 1954. Return to text.
- Wong, J.T., A co-evolution theory of the genetic code, PNAS 72:1909–1912, 1975. Return to text.
- Petrov, A.S., Gulen, B., Norris, A.M., Kovacs, N.A., Bernier, C.R. et al., History of the ribosome and the origin of translation, PNAS 112(50):15396–401, 2015. Return to text.
- Carter, C.W. and Wolfenden, R., tRNA acceptor stem and anticodon bases form independent codes related to protein folding, PNAS 112(24):7489–7494, 2015 Return to text.
- Harish, A. and Caetano-Anolles, G., Ribosomal history reveals origins of modern protein synthesis, Plos One, 12 March 2012 | doi:org/10.1371/journal.pone.0032776. Return to text.
- Freeland, S.J., Wu, T., and Keulmann, N., The case for an error minimizing standard genetic code, Origins of Live and Evolution of the Biosphere 33:457–477, 2003. Return to text.
- Vetsigian, K., Woese, C., and Goldenfeld, N., Collective evolution and the genetic code, PNAS 103(28):10696–10701, 2006. Return to text.
- Saxinger, C., Ponnamperuma, C., and Woese, C., Evidence for the interaction of nucleotides with immobilized amino-acids and its significance for the origin of the genetic code, Nature New Biology 234(49):172–174, 1971. Return to text.
- Yarus, M., Caporaso, J.G., and Knight, R., Origins of the genetic code: the escaped triplet theory, Annual Review of Biochemistry 74:179–198, 2005. Return to text.
- Wong, J.T., Evolution of the genetic code, Microbiology 5:174–181, 1988. Return to text.
- Amirnovin, R., An analysis of the metabolic theory of the origin of the genetic code, J. Molecular Evolution 44:473–476, 1997. Return to text.
- Higgs, P.G. and Pudritz, R.E., From protoplanetary disks to prebiotic amino acids and the origin of the genetic code; in: Pudritz, R.E., Higgs, P.G., and Stone, J. (Eds.), Planetary Systems and the Origins of Life, vol. 3, Cambridge University Press, Cambridge, UK, 2007. Return to text.
- Novozhilov, A.S., Wolf, Y.I., and Koonin, E.V., Evolution of the genetic code: partial optimization of a random code for robustness to translation error in a rugged fitness landscape, Biology Direct 2:24, 2007 | doi:10.1186/1745–6150–2–24. Return to text.
- Schimmel, P., Giege, R., Moras, D., and Yokoyama, S., An operational RNA code for amino acids and possible relationships to genetic code, PNAS 90(19):8763–8768, 1993. Return to text.
- Petrov, A.S., Bernier, C.R., Hsiao, C., Norris, A.M., Kovacs, N.A. et al., Evolution of the ribosome at atomic resolution, PNAS 111(28):10251–10256, 2014. Return to text.
- Fox, G.E., Origin and evolution of the ribosome, Cold Spring Harbor Perspectives in Biology 2(9):a003483, 2010. Return to text.
- Strobel, S.A., Repopulating the RNA world, Nature 411:1003–1006, 2001. Return to text.
- Robertson, M.P. and Joyce, G.G., The origins of the RNA world, Cold Spring Harbor Perspectives in Biology, 28April 2010 | doi:10.1101/cshperspect.a003608. Return to text.
- Salas, E.N., Shu, J., Cserhati, M.F., Weeks, D.P., and Ladunga, I., Pluralistic and stochastic gene regulation: examples, models and consistent theory, Nucleic Acids Research 44(10):4595–609, 2016. Return to text.
- Cserháti, M., Creation aspects of conserved non-coding sequences, J. Creation 21(2):101–108, 2007. Return to text.
- Wolf, Y.I. and Koonin, E.V., On the origin of the translation system and the genetic code in the RNA world by means of natural selection, exaptation, and subfunctionalization, Biology Direct 2:14, 2007 | doi:10.1186/1745–6150–2–14. Return to text.
- Narla, A. and Ebert, B.L., Ribosomopathies: human disorders of ribosome dysfunction, Blood 115( 16):3196–3205, 2010. Return to text.
- Wang, W., Nag, S., Zhang, X., Wang, M.H., Wang, H. et al., Ribosomal proteins and human diseases: pathogenesis, molecular mechanisms, and therapeutic implications, Medical Research Reviews 35(2):225–285, 2015. Return to text.
- Nakao, A., Yoshihama, M., and Kenmochi, N., RPG: the Ribosomal Protein Gene database, Nucleic Acids Research 32(Database issue):D168–70, 2004. Return to text.
- Shoji, S., Dambacher, C.M., Shajani, Z., Williamson, J.R., and Schultz, P.G., Systematic chromosomal deletion of bacterial ribosomal protein genes, J. Molecular Biology 413(4):751–761, 2011. Return to text.
- Cherry, J.M., The Saccharomyces Genome Database: a tool for discovery, Cold Spring Harbor Protocols 2015(12):pdb.top083840 | doi:10.1101/pdb.top083840. Return to text.
- Yutin, N., Puigbò, P., Koonin, E.V., and Wolf, Y.I., Phylogenomics of prokaryotic ribosomal proteins, PLoS One 7(5), e36972, 2012 | doi:org/10.1371/journal.pone.0036972. Return to text.
- Agafonov, D.E., Kolb, V.A., and Spirin, A.S., Proteins on ribosome surface: measurements of protein exposure by hot tritium bombardment technique, PNAS 94(24):12892–12897, 1997. Return to text.
- Mears, J.A., Cannone, J.J., Stagg, S.M., Gutell, R.R., Agrawal, R.K. et al., Modeling a minimal ribosome based on comparative sequence analysis, J. Molecular Biology 321(2):215–34, 2002. Return to text.
- Akanuma, G., Nanamiya, M., Natori, Y., Yano, K. et al., Inactivation of ribosomal genes in Bacillus subtilis reveals importance of each ribosomal protein for cell proliferation and cell differentiation, J. Bacteriology 194(22):6282–6291, 2012 Return to text.
- Baba, T., Ara, T., Hasegawa, M., Takai, Y. et al., Construction of Escherichia coli K-12 in-frame single-gene knockout mutants: the Keio collection, Molecular Systems Biology 2:(1) 2006 | doi:10.1038/msb4100050. Return to text.
- DeLabre, M.L., Kessl, J., Kamanou, S., and Trumpower, B.L., RPL29 codes for a non-essential protein of the 60S ribosomal subunit in Saccharomyces cerevisiae and exhibits synthetic lethality with mutations in genes for proteins required for subunit coupling, Biochimica et Biophysica–Gene Structure and Expression 1574:225–261, 2002. Return to text.
- Roberts, E., Sethi, A., Montoya, J., Woese C.R., and Luthey-Schulten Z., Molecular signatures of ribosomal evolution, PNAS 105:13953–13958, 2008. Return to text.
- Lecompte, O., Ripp, R., Thierry, J.C., Moras, D., and Poch, O., Comparative analysis of ribosomal proteins in complete genomes: an example of reductive evolution at the domain scale, Nucleic Acids Research 30(24):5382–5390, 2002. Return to text.
- Márquez, V., Fröhlich, T., Armache, J.P., Sohmen, D., Dönhöfer, A. et al., Proteomic characterization of archaeal ribosomes reveals the presence of novel archaeal-specific ribosomal proteins, J. Molecular Biology 405(5):1215–32, 2011. Return to text.
- Rackham, O. and Filipovska, A., Supernumerary proteins of mitochondrial ribosomes, Biochimica et Biophysica Acta 1840(4):1227–1232, 2014. Return to text.
- Sharma, M.R., Booth, T.M., Simpson, L., Maslov, D.A., and Agrawal R,K., Structure of a mitochondrial ribosome with minimal RNA, PNAS 106(24):9637–42, 2009. Return to text.
- Green-Willms, N.S., Fox, T.D., and Costanzo, M.C., Functional interactions between yeast mitochondrial ribosomes and mRNA 5’ untranslated leaders, Molecular and Cellular Biology 18(4):1826–1834, 1998. Return to text.
- Casad, M.E., Abraham, D., Kim, I.M., Frangakis, S., Dong, B. et al., Cardiomyopathy is associated with ribosomal protein gene haploinsufficiency in Drosophila melanogaster, Genetics 189(3):861–870, 2011. Return to text.
- Tselykh, T.V., Roos, C., and Heino, T.I., The mitochondrial ribosome-specific MrpL55 protein is essential in Drosophila and dynamically required during development, Experimental Cell Research 307(2):354–366, 2005. Return to text.
- Manuell, A.L., Quispe, J., and Mayfield, S.P., Structure of the chloroplast ribosome: novel domains for translation regulation, PLoS Biology 5(8):e209, 2007 | doi:10.1371/journal.pbio.0050209; Return to text.
- Yamaguchi, K., Beligni, M.V., Prieto, S., Haynes, P.A., McDonald, W.H. et al., Proteomic characterization of the Chlamydomonas reinhardtii chloroplast ribosome: identification of proteins unique to the 70S ribosome, J. Biological Chemistry 278(36):33774–33785, 2003. Return to text.
- O’Micks, J., Molecular structures shared by prokaryotes and eukaryotes show signs of only analogy and not homology, Answers Research J. 9:284–292, 2016. Return to text.
- Desmond, E., Brochier-Armanet, C., Forterre, P., and Gribaldo, S., On the last common ancestor and early evolution of eukaryotes: reconstructing the history of mitochondrial ribosomes, Research in Microbiology 162(1):53–70, 2011. Return to text.
- Mariscal, C. and Doolittle, W.F., Eukaryotes first: how could that be? Philosophical Transactions of the Royal Society, series B: Biological Sciences, 31 August 2015 | doi:10.1098/rstb.2014.0322. Return to text.
- Lȯpez-García, P. and Moreira, D., Metabolic symbiosis at the origin of eukaryotes, Trends in Biochemical Sciences 24(3):88–93, 1999. Return to text.
- Iwabe, N., Kuma, K-I., Hasegawa, M., Osawa, S., and Miyata, T., Evolutionary relationship of archaebacterial, eubacteria, and eukaryotes inferred from phylogenetic trees of duplicated genes, PNAS 86(23):9355–9359, 1989. Return to text.
- Peyretaillade, E., Biderre, C., Peyret, P., Duffieux, F., Méténier, G. et al., Microsporidian Encephalitozoon cuniculi, a unicellular eukaryote with an unusual chromosomal dispersion of ribosomal genes and a LSU rRNA reduced to the universal core, Nucleic Acids Research 26(15):3513–3520, 1998. Return to text.
- Van de Peer, Y., Ben Ali, A., and Meyer A., Microsporidia: accumulating molecular evidence that a group of amitochondriate and suspectedly primitive eukaryotes are just curious fungi, Gene 246(1–2):1–8, 2000. Return to text.
- Davidovich, C., Belousoff, M., Wekselman, I., Shapira, T. et al., The proto-ribosome: an ancient nano-machine for peptide bond formation, Israel J. Chemistry 50:29–35, 2010. Return to text.
- Bernier, C.R., Petrov, A.S., Waterbury, C.C., Jett, J., Li, F. et al., RiboVision suite for visualization and analysis of ribosomes, Faraday Discussions 169:195–207, 2014. Return to text.
- https://tinyurl.com/y7sqxjlq Return to text.





Readers’ comments
Comments are automatically closed 14 days after publication.