Journal of Creation 22(1):85–92, April 2008
Browse our latest digital issue Subscribe
Facilitated variation: a new paradigm emerges in biology
Facilitated variation is the first comprehensive theory of how life works at the molecular level, published in 2005 by systems biologists Marc Kirschner and John Gerhart in their book The Plausibility of Life: Resolving Darwin’s Dilemma. It is a very powerful theory, is supported by a great deal of evidence, and the authors have made it easy to understand. It identifies two basic components of heredity: (a) conserved core processes of cellular structure, function and body plan organization; and (b) modular regulatory mechanisms that are built in special ways that allow them to be easily rearranged (like ® Lego blocks) into new combinations to generate variable offspring. Evolvability is thus built-in, and the pre-existing molecular machinery facilitates the incorporation of new DNA sequence changes that occur via recombinations and mutations. The question of origin becomes especially acute under this new theory because the conserved core processes and the modular regulatory mechanisms have to already be in place before any evolution can occur. The new molecular evidence shows virtually all the main components of neo-Darwinian theory are wrong.
Scientific literature is currently drowning in information about the molecular mechanisms of life, but most people are unable to appreciate what it all means—so vast is the amount, so highly specialized in each reported study, and so obscured by the necessary but incomprehensible jargon. The publication in 2005 of the first comprehensive and easily readable theory of how it all works—Marc Kirschner and John Gerhart’s The Plausibility of Life: Resolving Darwin’s Dilemma1—thus marks a great milestone in the history of biology. Kirschner is Professor of Systems Biology at Harvard Medical School and Gerhart is Professor of Systems Biology at UC Berkley Medical School.2
In this article, I shall show how Kirschner and Gerhart’s theory signals the emergence of a new paradigm in biology by contrasting it with origin-of-life experiments and neo-Darwinian theory, and will augment it with some more recent research findings.
Life and non-life
To appreciate what life looks like at the molecular level we need some background understanding of the gap between life and non-life, and how originating events may have filled that gap. According to neo-Darwinian theory, life evolves in small steps. Genes produce organisms, and mutations in genes produce changes in organisms. Those changes that survive the ‘sieve’ of natural selection provide the required small steps that turn one kind of life into another. Population biology experiments are claimed to have validated this theory for many different kinds of genetic traits.
Extrapolating this theory backwards, life must have also arisen in small steps via natural chemical events in the environment. Nobel Prize winning biochemist Christian de Duve has clearly summarized most of the necessary events in his book, Singularities: Landmarks on the Pathways of Life.3 There is, yet, no experimental evidence for a stepwise neo-Darwinian originating mechanism, so de Duve’s singularities are what we might colloquially call ‘brick walls’.
Living organisms have two main components: (a) enzyme-mediated biochemistry and (b) information-based regulatory processes. Which came first? De Duve favours an ‘enzymes first’ model because the information-based systems are so optimal and specialized that he believes some process of selection was needed to separate out the spectacularly clean (100% purity) components from the ‘dirty gemisch’ (impure mixture) of the environment.
However, physicist Hubert Yockey has studied information in biology for 50 years and persuasively argues that because life has no reverse code for transferring information from proteins to RNA or DNA then it is impossible for life to have arisen in a ‘proteins first’ scenario. The information must have come first. The simplest code would have been a binary (two-letter) alphabet but all life works upon a more complex four-letter alphabet, so Yockey concludes that the question of origin is undecidable.4 This is not a necessary conclusion however, and appears to be no more than a ruse to avoid the uncomfortable conclusion that life may have been intelligently designed.
Life in molecular detail: the new paradigm
Against this background, we can now look to the summary model of how life works as given by Kirschner and Gerhart (I shall refer to it as the KG model). They identify two major components:
- conserved core processes of cell structure, function, and body plans;
- core processes are regulated in modular ways (like ® Lego blocks) that can be easily rearranged into new combinations, to be used in new times, places and amounts to generate variable offspring.
Evolvability is thus built-in. The existing modular structure and its regulatory systems facilitates the incorporation of changes in DNA sequences (produced by recombinations and mutations) into functionally viable offspring that can adapt to new environments. KG theory is claimed to be a largely complete molecular explanation for how natural variation and natural selection produce all the variety of life on Earth—Darwin’s theory, according to the authors, is now a validated whole.
A new view of heredity
Neo-Darwinists view heredity as being all about genetics. For example, the official journal of the Genetics Society is called Heredity. But genetics is all about change and we have discovered so many ways in which organisms can change that we are now faced with a huge unanswered question: how do they manage to stay (approximately) the same, generation after generation? As the late Stephen Jay Gould maintained throughout his career in paleontology—stasis, not change, is the major feature of natural history.5
Neo-Darwinism has no answer to this challenge for two reasons: (a) genes and chromosomes can be mutated at any and every position so there is no limit to the potential for change, and (b) the agents of change (mutations and environment) are beyond the organism’s control.
But KG theory does give us an answer—the conserved core processes remain the same during reproduction. When a mother passes on an egg cell to its offspring, that cell contains everything required by the offspring in its architecture and machinery. The DNA sequences provide for the manufacture of more raw materials for the embryo to go through its development process, but the actual architecture and machinery itself is provided by the mother. The new outer membrane of the embryo is just that of the mother’s cell extended with more of the same material. The new cytoskeleton is just the mother’s cytoskeleton extended with new material. The new organelles are the mother’s organelles that replicate independently of the chromosomes. The new membranes are the mother’s membranes extended with more of the same material.
During the early stages of embryogenesis, the new chromosome set is entirely shut down and all the groundwork of the embryo is laid by thousands of different RNA types supplied by the mother. Only after this groundwork is laid does the new chromosome set become active and the mother’s RNAs are degraded and recycled.
The variability that is built-in to this heredity process is the modular gene regulation and signaling networks. A suitable analogy might be a house and its network of power, plumbing and communications channels and interfaces. The wiring and piping are built into the house structure, but there are numerous interface points to which a wide variety of household appliances can be attached, detached and rearranged. It is the combination of devices plugged into this network that provides the variation, and the house with its plumbing and wiring system that provides the stasis. To what extent the ‘house’ itself can be varied is yet to be determined.
Conserved core processes
Chapter 7 of Kirschner and Gerhart’s book summarizes this subject so I will simply quote selectively from it. My additions or summaries are in square brackets:
‘Conserved core processes [typically consist of] several protein components [on average about 5, maximum probably about 300], conserved in their [amino acid] sequence. Their function is to generate the phenotype from the genotype. These processes arose historically in a few intermittent waves of innovation.
‘On the lineage towards humans, these innovations include:
- the processes in the first bacteria [all the machinery in a bacterial cell],
- [the processes in] the first eukaryotes [all the machinery in a eukaryote cell],
- [the processes in] the first multi-cellular organisms [cooperation between cells, specialization of structure and function of different cells, and integration of specialized cell complexes into functional organs and organisms],
- [the processes in] large bilateral body plans in metazoans (including chordates and vertebrates),
- [the processes in] neural crest cells in vertebrates [which allow diversification of the head region],
- [the processes in] limbs in the first land animals,
- [the processes in] the neocortex [the key region of brain development].
‘Most evolutionary change in the metazoa [multi-celled animals] since the Cambrian has come not from changes of the core processes themselves or from new processes, but from regulatory changes affecting the deployment of the core processes. These regulatory changes alter the time, place, circumstance and amount of gene expression …
‘The core processes are built in special ways to allow them to be easily linked together in new combinations … these special properties include:
(a) Weak linkage, a property particularly of signal transduction [detection and response] and transcription [copying]. … the response is maximally prepared and ready to be triggered [by a GO or STOP signal].
(b) Exploratory behavior, a property of [cellular processes and populations of organisms] … the capacity to generate an unlimited number of outcome states [which are] built to be receptive to the [selective] agent [that will serve] as a stabilizing force, selecting one state among the large number of states generated.
(c) Compartmentation, a property of embryonic spatial organization and cell type control. [Compartmentation has] facilitated a large increase in the complexity of anatomy and physiology without a corresponding increase in the complexity of the conserved core processes.
‘Generation of variation is facilitated principally by:
- reducing the lethality of mutations,
- reducing the number of mutations needed to produce novelty, and
- increasing the genetic diversity in the population by suppressing lethality [and thus allowing the mutations to be stored and inherited].
‘Robustness [is] at the centre of our theory … the conserved core processes are built [robustly] to give sufficient outputs despite altered conditions and inputs. [The properties] of robustness, flexibility and versatility are [needed] to enable the core processes to work together … the organism as a whole is … a poised response system … It responds to mutation by making changes it is largely prepared in advance to make. … Genetic variation or mutation does not have to be creative; it only needs to trigger the creativity built into the conserved mechanisms.
‘All the special properties of the conserved core processes had to evolve before regulatory evolution could escalate, for if the components of different processes were to interfere with one another in the new combinations, such expression would afford no benefit.
‘Facilitated variation assumes the availability of [the conserved core processes]. The evolution of these processes and properties would seem to be the primary events of evolution, requiring high novelty. … Once the conserved processes were available, though, the possibility of variation by regulatory shuffling and gating of these processes was unleashed, and shuffling and gating were much simpler than inventing the processes.
‘The main accomplishment of the theory of facilitated variation is to see the organism as playing a central role in determining the nature and degree of variation … We think the organism is so constituted that its own random genetic variation can evoke complex phenotypic change.’
Further relevant comments from Chapter 8 include:
‘ … evolvability … is the greatest adaptation of all … Variation is facilitated largely because so much novelty is available in what is already possessed by the organism’ (pp. 252, 273).
‘The theory of facilitated variation opens up a new set of questions about the origins of the conserved core processes … [they] may have emerged together as a suite, for we know of no organism today that lacks any part of the suite. … The most obscure origination of a core process is the creation of the first prokaryotic cell. The novelty and complexity of the cell is so far beyond anything inanimate in the world of today that we are left baffled by how it was achieved’ (pp. 253, 256).
Invisible anatomy
Kirschner and Gerhart coined the term ‘invisible anatomy’ to describe the regulatory circuits that produce the visible anatomy. To construct an adult from a zygote, the zygote must first build a phylotypic embryo—a mass of cells with highly conserved form, which is the same right across its phylum. This philotypic stage is divided into numerous, largely independent, 3-dimensional compartments within which different gene switching networks are wired up in different ways appropriate for the unique developmental cascade that will subsequently occur in each compartment.
But the signal network is not instructive, it is permissive—it does not tell the circuits what to do, it merely releases or represses the already built-in abilities of cells to do whatever needs to be done. Humans have about 300 compartments in their phylotypic embryo. That means there must be least 300 different circuits—developmental programs for body segments—that can be activated or repressed in every cell.
Switching networks
The main difference between neo-Darwinian and KG theory is that the former views genes as having a continual effect on organisms, whereas the molecular reality is that genes only work when they are switched ON. This is a profound difference. Everything in KG theory flows from this fact. Evolution occurs not primarily by changing DNA sequences, as neo-Darwinists assume, but by rearrangement of switching circuits.
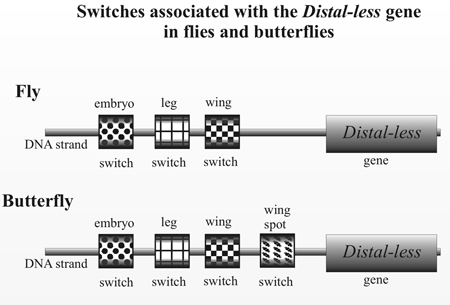
Gene switches are sections of DNA on the chromosome usually near to where the gene is situated (figure 1). One gene may be involved in ten or more stages in development and it will have a separate switch for each stage. Sean Carroll, a leading researcher in this field, says, ‘animal bodies [are] built—piece by piece, stripe by stripe, bone by bone—by constellations of switches distributed all over the genome [emphasis added].’6 Evolution occurs primarily by adding or deleting switches, for this is the only way to change the organism while leaving the gene itself undamaged by mutation so that it can continue to function normally in its many other roles. Carroll considers this concept to be ‘perhaps the most important, most fundamental insight from evolutionary developmental biology.’7
Figure 1 illustrates evolution-by-switch-addition by showing how butterfly wing spots are produced by adding a new wing-spot switch to an existing gene Distal-less that is already involved in development of the insect embryo, leg and wing.8
Gene switches are very complex devices. Carroll compares them to a Global Positioning System (GPS) satellite-navigating device that integrates information from many different satellites to calculate the correct output in a given situation. Gene switches likewise give ‘exquisite geographic specificity [from the built-in logic] … the makeup of every switch is different [and] the physical integrity of switches is very important to normal development. If a switch is disrupted or broken by mutation, then the proper inputs are not integrated.’9
The reason why genes only work by being either fully ON or OFF is very easy to understand—because a part-formed transcript would become useless junk in a crowded cell. Only fully formed transcripts are useable, and when they are not wanted, the gene needs to be turned OFF so that it will not clog up the heavily crowded cell with unwanted transcripts.
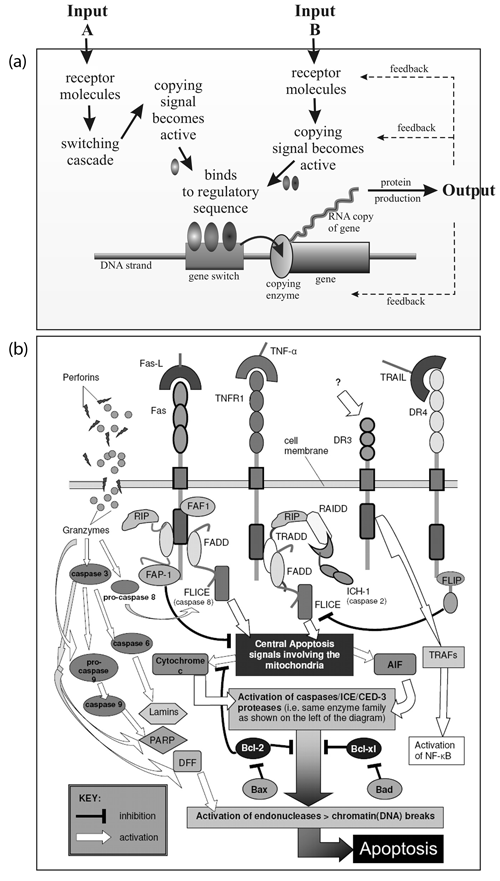
Figure 2 outlines the components of a gene switch that uses negative feedback as its control mechanism. The molecules involved in switches are called ‘transcription factors’ and can be activators (that send a GO message) or repressors (that send a STOP message). If a repressor is repressed then STOP STOP = GO.
Uri Alon at the Weizmann Institute has researched switches and signal networks and found two main types:10
Switches associated with signal reception and response, which act over metabolic time scales of seconds. These include: single factor regulation, negative autoregulation, positive autoregulation, feed-forward loops (FFL) of both positive and negative kind, multi-output FFLs that regulate numerous genes simultaneously, single-input modules, and dense overlapping regulons that can regulate one or hundreds of output genes, and they can have one or hundreds of inputs from various sources.
Switches associated with development over the lifetime of the organism. These include: positive feedback loops, negative feedback loops, diamond networks, multi-layer diamond networks, and feed-forward loops that combine into large networks.
Switches are readily disabled by mutation, so Alon addressed the question of whether systems such as FFLs evolved from duplication of an ancestral FFL. The answer appears to be no, because apparently homologous genes are usually regulated by transcription factors that are so different that they are classed into completely different families. Evolution must have converged independently on the same regulation circuits over and over again.
This is perhaps explained by the fact that
‘ … transcription networks seem to rewire rapidly: it takes only a few mutations to remove the binding site of a regulator in a given promoter, and thereby lose an arrow in a network. Hence, even closely related organisms often have different network motifs to regulate a given gene, provided that they live in different environments … One hypothesis is that the network[s] are selected according to the computations that are required in the environment of each species.’10
This latter finding seems to agree with KG theory, that switching circuit modularity provides the major source of natural variation. Another important confirmation of the concept is the Savageau demand rule. This experimentally observed rule is that frequently needed genes tend to be regulated by activators, while rarely needed genes tend to be regulated by repressors. It has been shown that a strategy in which errors are minimized leads to the Savageau demand rule.11 That is, errors (mutations and imprecise biochemical reactions) are minimized in the search for useful circuit combinations.
Embryonic switching patterns
We are now in a position to illustrate embryogenesis, in broad outline, as a series of switching events. The ‘geography’ or ground-plan for each organism is established during the early divisions of the zygote. Important geographical factors include:
- Inside (endoderm and mesoderm) and outside (ectoderm)
- Head (mouth and brain end) and tail (anal end)
- Left and right (in bilateral animals)
- Front and back (in bilateral animals).
These geographical circuits are positive feedback loops that shunt irreversibly into, for example, ‘tail OFF and head ON’ mode. The comparable circuit in the tail end shunts irreversibly into the ‘tail ON and head OFF’ state. In all descendents of these cells, later instructions will pass through these circuits so that, for example, when the instruction is given to build a limb, the state of the geographical circuits will ensure that a forelimb is produced at the head end and a hind limb is produced at the tail end.
Within our group of bilaterians, the vertebrates, further circuitry is linked up within this three-dimensional ground-plan so that by the ‘phylotypic stage’ all the embryos look remarkably similar (drawings of which Haeckel infamously fudged to make look even more similar than they really are). The similarity is no coincidence, however, because all vertebrate embryos are patterned by exactly the same set of genes, as shown in figure 3. All the genes up to hox6 regulate brain and head development, and those from hox7 to cad regulate spinal cord and body development.
By this stage, the vertebrate embryos consist of about 300 largely independent compartments, and further development occurs according to a separate switching cascade in each compartment. The body-patterning genes shown in figure 3 create these compartments via single-input circuits that have multiple thresholds of interaction with the ground-plan circuits (inside-outside, head-tail, left-right, front-back) and the body differentiating genes (those that produce limbs, ears, ribs, etc.).
Autopoietic control
Life is controlled by coded information. The overall purpose of that information appears to be survival, and in particular, survival via variable reproduction. KG theory says that organisms are built to vary, and it could not be any other way because brittle life, like Paley’s metal watch, would malfunction under the first impact of either internal or external impediment. Rather ‘the organism as a whole is a … poised response system [ready to make] changes it is largely prepared in advance to make’ (KG, p. 226).

But protein-coding information of DNA is clearly not the only information operating in cells. A gene only gives the linear sequence of amino acids in a protein, yet its key function is the result of its 3-dimensional shape, not its linear sequence. Many different amino acids could substitute into the linear sequence without reducing its functionality, but the 3-D shape is very tightly constrained, yet cannot be predicted from its linear sequence. Proteins can fold in numerous different ways, so there must be extra information somewhere else that guides the folding process. Special molecules called chaperones guide the folding process, so there must be folding information built-in to the chaperones. They can also detect and correct mis-folded proteins, and they can detect when a protein is mis-folded beyond repair and have it marked for degradation and recycling.
Autopoietic decision making during embryogenesis is of the ‘if … then … ’ kind familiar to computer programmers. Embryonic cells make decisions based upon three kinds of information: (a) instructions from the mother (mRNAs in the egg cytoplasm), (b) conditions within the cell itself, and (c) information from its immediate neighbours. Thus, if a cell has all its specialization circuits in OFF mode, and it has its polarity circuit in an ON state, and it has only one neighbouring cell, then it concludes that it is in the two-cell state of embryogenesis so it will divide and switch ON its bilateral circuits but keep all its specialization circuits in OFF mode.
At a later stage, if there are no longer any instructions from the mother, and the cell’s specialized liver circuit is ON and all its neighbours are liver cells, and the embryogenesis circuitry is OFF and the fetal circuitry is ON, then the cell will divide and reproduce an identical copy of itself to allow the liver to grow in size until birth stage.
In later life, the autopoietic system will ensure that maintenance and repairs are carried out to keep the cell functioning properly. But when the telomere ‘clock’ says that time has run out, it will trigger a release of cytochrome c from the mitochondria into the cytoplasm which will set the apoptosome into action to dismantle the cell and recycle its contents.12
Evidence supporting the theory

The primary difference between neo-Darwinism and KG theory is that the former puts genes in control of heredity and thus evolution, while the latter puts the cell in control. Figure 4 illustrates this crucial difference.
The molecular evidence is clearly in favour of cell control. A recent intensive study of transcription activity in a 1% sample of the human genome found an astonishing amount of unexpected activity. Virtually the whole genome is transcribed, in both directions (both strands of the DNA double helix), in multiple copies (on average 5 in gene regions and 7 in non-gene regions) that overlap by an average 10 to 50 times the size of a typical gene. The best predictor of where and when this transcription takes place is just one factor—chromatin structure.13 Chromatin is the complex of DNA and protein that super-coils the long thin DNA into short fat chromosomes, and it must be uncoiled in order for transcription to occur.
The same conclusion—that chromatin structure lies at the heart of transcription activity—was arrived at via study of the relationship between chromatin and nuclear pores.14 In eukaryotes, chromosomes are housed in the nucleus, and access to and from the nucleus is very closely controlled via special structures called the nuclear pore complex (NPC). Transcription only occurs at the inner opening of these NPCs. The relevant chromosome must be brought to a pore and the transcription site correctly aligned. The DNA is unwound from its scaffold proteins, then the histone coils are twisted around to expose the copy region, the double-helix is unzipped, and the transcription machinery produces an RNA copy of the DNA. The transcript is checked for accuracy and corrected if necessary (or degraded if faulty beyond repair) then the RNA is tagged for export out through the NPC and to its destination in the cell. The DNA is then silenced again by being zipped up and rewound onto its histone and scaffold protein chromatin structures. So DNA is normally in a form analogous to a closed book. When the cell wants some information it opens the book, copies the relevant section, and then closes the book again. DNA does not control this process—it is kept in storage until it is needed. The cell is clearly in control.
The second major difference between KG theory and neo-Darwinism is in the way genes act upon organisms. In the classic case of Darwin’s Galápagos finches, neo-Darwinian theory explains the variation in finch beak size and shape via mutations and natural selection acting repeatedly over a long period of time. Many small changes must occur independently in the upper and lower beaks, in the adjacent skull, and in the head muscles, to coordinate and order them all into the necessarily viable intermediate beaks of the birds that need to survive throughout the period of divergence.
In contrast, recent experimental work suggests that just two regulatory changes are involved. The bone morphology protein BMP4 when expressed earlier or later in embryogenesis causes broad or narrow beak development,15 and more or less of the calcium regulator protein calmodulin produces long or short beaks, respectively.16 Gerhart and Kirschner17 cite this as experimental validation of their theory. The whole craniofacial developmental process is compartmented and coordinated by a modular regulatory system that can be easily rewired ‘with a few regulatory mutations’ (KG, p. 236) to produce new features that are readily integrated into the already-prepared, robust, conserved-core-process-based system. Field observations confirm that such changes take place rapidly across just a few generations.18
More neo-Darwinian errors
The neo-Darwinian genetic theory of heredity assumed that characteristics of organisms are coded on genes with roughly a ‘one-gene-to-one-character’ correspondence. As organisms evolved to greater complexity, more genes were added via gene duplication and subsequent independent mutation of the extra copy into useful new characters.19 More complex organisms were thus expected to carry more genes than less complex ones. Furthermore, lineages that diverged early in the history of life would have mutated at virtually every locus, making them quite unlike at the genetic level. This led Ernst Mayr to state in his 1963 book Animal Species and Evolution ‘the search for homologous genes [derived from the same ancestor] is quite futile except in very close relatives.’20
These predictions have all been dramatic falsified by molecular discoveries:
- There is no one-to-one correspondence between genes and characters. Most genes are pleiotropic—they affect many different parts and stages of life. And all but the most trivial characters are determined by large numbers of genes—50% to 80% of the entire genome is required for many bodily functions in vertebrates.21
- Genetic information structures are not linear, but interleaved, producing multiple overlapping transcripts. Moreover, the exons (DNA segments that directly code for protein segments) in a gene are not specific to that gene but can participate in modular fashion with up to 33 different genes on as many as 14 different chromosomes.22
- There is no correlation between organism complexity and gene number. Rice and crayfish carry more genes than humans.
- Homologous genes occur right across the spectrum of life. About 20% of the human genome is homologous with bacteria, about 50% is homologous with eukaryotes (fungi, plants, animals), about 80% is homologous across the animal kingdom, and about 99% is homologous across all the vertebrates, leaving only about 1% that is uniquely human.23 About 500 genes are ‘immortal’ and have not changed at all in their key functional sequences across the whole history of life.24
One of the most serious errors—that will need a lot of undoing—is the vast amount of molecular taxonomy that has been based upon the idea that ‘junk DNA’ provides us with a record of past mutations and thus acts as a ‘molecular clock.’ We now know that non-protein-coding DNA is more active in the cell than genes. According to KG theory, molecular taxonomy can only work correctly by comparing ‘hidden anatomies’ across taxa, not DNA sequences. To understand hidden anatomy we will have to find the regulatory code. New aspects of gene regulation are being reported daily, but so far, no one has been able to put together the complete code for a whole organism.
Conclusion
Let’s stand back consider the big picture of how life works at the molecular level.
Life consists of conserved core processes and modular regulatory circuits. All the special properties of the conserved processes had to be in place before regulatory evolution could take place. Where did they come from? ‘They may have emerged together as a suite, for we know of no organism today that lacks any part of the suite.’
‘The novelty and complexity of the cell [the most important conserved core processes that has modular regulatory circuitry built-in] is so far beyond anything inanimate in the world of today that we are left baffled by how it was achieved.’
A living organism is ‘a poised response system [that] responds to mutation by making changes it is largely prepared in advance to make.’ ‘Genetic variation or mutation does not have to be creative; it only needs to trigger the creativity built into the conserved mechanisms.’ It could not be otherwise, because invariable life would soon become extinct.
Who will be game enough to say the words? Only intelligent design can explain such data. There are no naturalistic explanations.
References
- Kirschner, M.W. and Gerhart, J.C., The Plausibility of Life: Resolving Darwin’s Dilemma, Yale University Press, New Haven, CT, 2005. Return to text.
- Lightner, J., Designed to inhabit the earth, a review of The Plausibility of Life by Marc W. Kirschner and John C. Gerhart, Journal of Creation 22(1):33–36, 2008. Return to text.
- De Duve, C., Singularities: Landmarks on the Pathways of Life, Cambridge University Press, New York, 2005. Return to text.
- Yockey, H.P., Information Theory, Evolution and the Origin of Life, Cambridge University Press, MA, 2005, Chs.3, 13. Return to text.
- Gould, S.J., The Structure of Evolutionary Theory, Harvard University Press, 2001, MA, p.749. Return to text.
- Carroll, ref. 8, p. 111. Return to text.
- Carroll, S.B., Evolution at Two Levels: On Genes and Form, PLoS Biology 3(7):1159–1166, 2005; p.1162. Return to text.
- Carroll, S.B., Endless Forms Most Beautiful: The New Science of Evo-Devo, Norton, New York, 2005. Return to text.
- Carroll, ref. 8, pp.114–118, 124. Return to text.
- Alon, U., Network motifs: theory and experimental approaches, Nature Reviews Genetics 8:450–461, 2007. Return to text.
- Shinar, G., Dekel, E., Tlusty, T. and Alon, U., Rules for biological regulation based on error minimization, PNAS 103(11):3999–4004, 2006. Return to text.
- Riedl, S.J. and Salvesen, G.S., The apoptosome: signalling platform of cell death, Nature Reviews Molecular Cell Biology 8:405–413, 2007. Return to text.
- Birney, E. et al., Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project, Nature 447:799–816, 2007. Return to text.
- Akhtar, A. and Gasser, S.M., The nuclear envelope and transcriptional control, Nature Reviews Genetics 8:507–517, 2007. Return to text.
- Abzhanov, A., Protas, M., Grant, B.R., Grant, P.R. and Tabin, C.J., Bmp4 and morphological variation of beaks in Darwin’s finches, Science 305(5689):1462–1465, 2004. Return to text.
- Abzhanov, A., Kuo, W.P., Hartmann, C., Grant, B.R., Grant, P.R. and Tabin, C.J., The calmodulin pathway and evolution of elongated beak morphology in Darwin’s finches, Nature 442:563–567, 2006. Return to text.
- Gerhart, J. and Kirschner, M., The theory of facilitated variation, PNAS 104:8582–8589, 2007. Return to text.
- Grant, P.R., Ecology and Evolution of Darwin’s Finches, Princeton University Press, NJ, 1999. Return to text.
- Ohno, S., Evolution by Gene Duplication. Springer-Verlag, New York, 1970. Return to text.
- Carroll, S.B., The Making of the Fittest: DNA and the ultimate forensic record of evolution, Norton, New York, p. 78, 2006. Return to text.
- Lein, E.S. et al., Genome-wide atlas of gene expression in the adult mouse brain, Nature 445:168–176, 2007. Return to text.
- Kapranov, P., Willingham, A.T. and Gingeras, T.R., Genome-wide transcription and the implications for genomic organization, Nature Reviews Genetics, published online 8 May 2007. Return to text.
- Lewin, B., Molecular Biology, MBIO-3.14 More complex species evolve by adding new gene functions, <www.ergito.com/toc.jsp?bcs=MBIO>, 2 September 2006. Return to text.
- Carroll, ref.20, p.79. Return to text.
- Bell, P., Apoptosis: cell ‘death’ reveals creation, Journal of Creation 16(1):90–102, 2002; <creation.com/apoptosis>. Return to text.




Readers’ comments
Comments are automatically closed 14 days after publication.