Journal of Creation 34(3):87–94, December 2020
Browse our latest digital issue Subscribe
The surprisingly complex tRNA subsystem: part 2—biochemical modifications
Scores of enzymes are necessary to chemically modify tRNAs since thermodynamic calculations on unmodified sequences rarely predict proper folding into the cloverleaf structure as the ground state. Other modifications are indispensable to permit translation of synonymous codons, translational fidelity of codon–anticodon interaction, maintenance of reading frame at ribosomes, and other cellular processes. Many very complex enzymes are necessary. The yW modification requires 5 polypeptides; t6A modification of adenine found at position 37 requires four proteins; and around 25 individual proteins are involved in wobble uridine modifications. The modifications are dynamic, tissue specific, respond to environmental changes, and tRNA concentrations are used as a regulatory signal in several ways to regulate amount of translation.

The cellular genetic system is irreducibly complex, being composed of several collaborating subsystems based on DNA, RNA, and proteins. Genetic code translation cannot occur without tRNAs. In part 11 we saw, however, that multiple protein-based molecular machines are also needed to transcribe the tRNA genes, remove 5’-end leaders, 3’-end trailers, remove introns and in many cases add a trinucleotide pattern ‘CCA’ so that aminoacylation can occur. We also discussed how the CCACCA pattern is added to defective tRNAs which need to be degraded.
Here in part 2 we will see that dozens of biochemical modifications are necessary for tRNAs to function, each of which require a specific protein-based enzyme. As we progress through this series it will become increasingly apparent that the tRNA subsystem must have been planned upfront and many parts introduced simultaneously.
Post-transcriptional modification. Introduction and overview
Genomes encode scores of enzymes responsible for catalyzing various chemical modifications on tRNAs2 (figure 1). These modifications can occur on tRNA precursors or fully processed tRNAs.3 Nucleotide insertions or substitutions are often necessary to ensure base pairing within the tRNA in the three kingdoms of life and that the modifications occur in a stepwise fashion.4,5
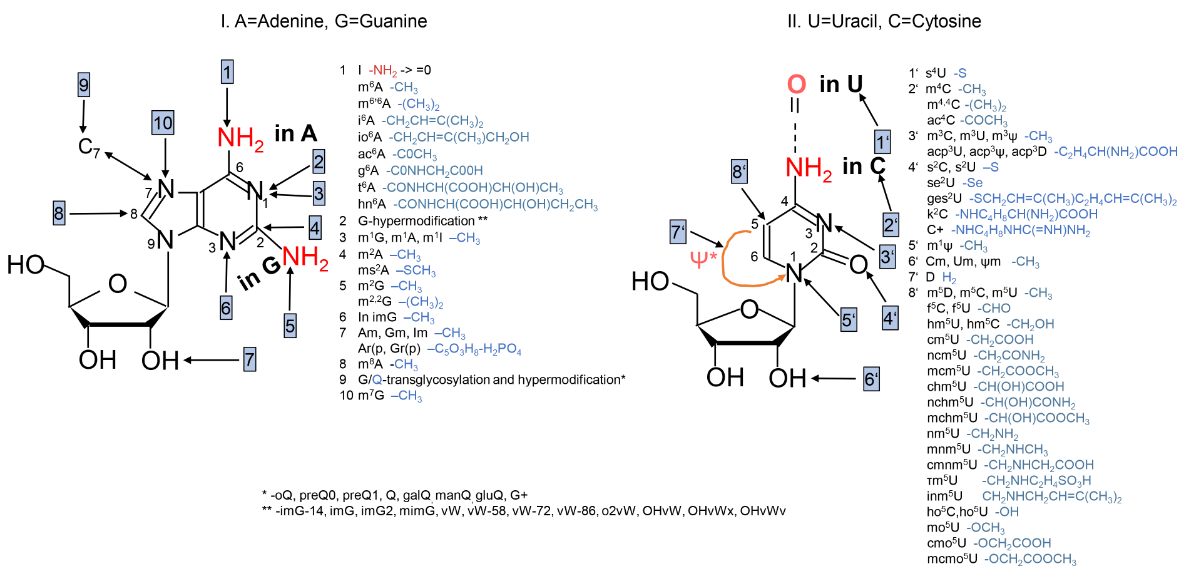
Some enzymes modify different types of RNA (‘dual substrate specificity’). However, most tRNA modification enzymes have unique specificity for tRNA.7
Figure 1 illustrates a vitally important insight concerning enzymes, namely their exquisite specificity. Notice that in many cases a methyl group (-CH3) is added to different, specific parts of a nucleotide. In cells, thousands of different enzymes are needed with extreme specificity to avoid damaging the much greater number of possible substrates and positions which should not be modified.
Enzymes process individual biomolecules one after the other, carefully rejecting the wrong substrates. This is why they are called molecular machines. The technical challenge to design de novo enzymes with no knowledge of what is already found in nature, taking the entire cellular context into account, lies considerably beyond the skills of the best chemists. Contrast typical biological enzymes, which consist of tens of thousands of atoms in the correct positions in three dimensions with the kinds of reagents used by chemists in laboratories. For example, secondary alcohols are oxidized to ketones using reagents like chromium trioxide, CrO3 (Jones oxidation). This is merely a four-atom reagent which is consumed for each molecule oxidized, with little discrimination among secondary alcohols.
It is now appreciated that tRNA modifications serve many functions including: tRNA discrimination, translational fidelity via codon–anticodon interaction, maintenance of reading frame, tRNA stability,7 lipid aminoacylation,8 and bacterial conjugation.9
Human nuclear-encoded tRNAs contain on average 13 modifications per molecule, but the number varies considerably.10 For example, tRNATyr from placenta has 17 modifications, whereas tRNASec from HeLa cells has only 3. Mitochondrial tRNAs contain about five modifications per molecule.10 As of today, <50% of nuclear-encoded human tRNAs have their modifications mapped, and in just one cell type.10
Over 140 nucleotide modifications have been identified so far, some of which are conserved throughout all domains of life.11,12 Nucleotide modifications include base or sugar methylations, base deaminations, base isomerizations, and exotic moiety additions to bases.7 The anticodon loops of nearly all tRNAs are heavily modified, predominantly at positions 34 and 37.11
Budding yeast has 25 known tRNA modifications, creating tRNAs in which ~15% of the residues are nucleotides other than A, G, U, or C, with an average of ~12 nucleotide modified per tRNA. These modified nucleotides serve important functions including tRNA discrimination, translation fidelity, and tRNA quality control.7 Many genes are needed to perform these modifications.7,13
Approximately 15%–25% of all nucleotides in eukaryotic tRNA contain modifications.14 These modifications are dynamic, respond to environmental change,15 and have been proposed to serve various purposes. An important example is discrimination between initiator tRNAMet from elongator tRNAMet through ribosylation at position A64.16
Fully modified tRNAs are invariably more stable than unmodified transcripts.15 In fact, thermodynamic folding of unmodified sequences only rarely predicts the cloverleaf as a ground state. The chicken-and-egg dilemma is ubiquitous. Most tRNAs will not fold reliably into the correct structure unless properly modified chemically, but the modification protein enzymes depend on correctly functioning tRNAs.
Research continues to reveal ever more differences across kinds of organism. Histidyl tRNAs contain an extra 5’-G nucleotide missing in all other tRNAs. In E. coli, the G is derived transcriptionally, and RNaseP does not remove the extra G when it processes the 5’ end. In yeast, however, the 5’-most G in the histidyl tRNA is added by a histidyl tRNA guanylyltransferase. The 5’-G of spinach chloroplast histidyl tRNA is derived from the gene, whereas that of animal mitochondria is added post-transcriptionally.4
As one would expect from designs which rely on such important modifications, mutations in tRNA and tRNA-processing genes have been linked to many diseases.16
Most modification enzymes are individually unessential. Despite the important roles played and the fact that many modification enzymes are very similar across broad classes of organisms, most of the yeast genes encoding tRNA modification enzymes are unessential individually.17 In yeast only those responsible for adenosine A34 to inosine I34 deamination (TAD2 and TAD3) and methylation of adenosine m1A58 (TRM6 and TRM61) are known to be absolutely essential.7 However, severe phenotype deficiencies occur when two or more modification genes of cells are missing or damaged. For example, PUS1 and PUS4 are individually unessential genes but the cell is severely defective if both genes are non-functional.7
Note that natural selection would not have produced nor fine-tuned an unessential new gene. What is not immediately needed and poses relative selective disadvantages tends to be eliminated from genomes, especially for small, rapidly reproducing organisms. RNA modifications are costly, requiring significant energy for the cell. For example, one biochemical modification, RNA methylation, uses S-adenosylmethionine (SAM) as the methyl donor, and to produce one SAM molecule requires the energy equal to hydrolyzing 12 to 13 ATP molecules.16
Complex enzymes are required
Some tRNA nucleotide modifications rely on large, sophisticated modification complexes13 to cleave, trim, splice, add terminal nucleotides, and modify specific nucleotide residues through special processing pathways.18 Most of the modifiers are dynamically regulated in response to environmental cues, often with integrated feedback between many modifications and their pathways.19
Many tRNA modification enzymes are composed of a single subunit whereas others require multiple proteins. In the latter case different enzymes often modify chemical groups in a sequential fashion (figure 2). A synthetic natural products organic chemist can best appreciate the brilliance of pathways such as this one, and the absurdity of any claim that the necessary enzymes arose through an unplanned process.
yW modification requires 5 polypeptides7 (figure 2).

Biosynthesis of mcm5s2U34, mcm5U34 and derivatives requires >25 polypeptides.7
t6A biosynthesis found at position 37 requires four proteins: YrdC, YgjD, YeaZ, and YjeE in bacteria such as E. coli.11 However, of these four proteins only members of the YrdC and YgjD families are used by all organisms studied so far. The absence of both YeaZ and YjeE in eukarya and archaea means new genes need to be discovered.11 This modification prevents formation of a U33-A37 pairing interaction, while allowing cross-strand stacking of A38 and t6A37 with the first position of the codon. This modification of A (Adenine) is necessary to permit translation of all codons with A in the first position.11 We see once again the chicken-and-egg problem. Without the prerequisite enzyme about a fourth of the codons could not be translated.
Methylations at different positions affect multiple properties, including folding details, thermostability, protection from cleavage, and priming for subsequent modifications. tRNA methyltransferases catalyze these reactions.13
Pseudouridylation Ψ (C11–C5) is one of the most common alterations. The carbon-nitrogen glycosidic bond (C1–N1) is replaced with a carbon-carbon bond (C1–C5) between the ribose and uracil. This isomerization is catalyzed by pseudouridine synthases (PUS).13
i6A37 modifications consist of the addition of an isopentenyl group to N6 of adenosine-37 for certain tRNAs, catalyzed by isopentenyl transferases (IPTases).13
Carboxymethylation (cm5). In eukaryotes the Elongator complex converts 11 out of 13 yeast tRNAs carrying a U34 in the wobble position. The core Elongator complex (molecular weight of more than 850 kDa) includes two copies of six subunits and is regulated by about nine regulatory factors.13 After the necessary cm5 modification on U34 other enzymatic cascades can then guide additional modifications to produce mcm5U, ncm5U, mcm5s2U, mcm5Um, or mchm5U. Overall, around 25 individual proteins are involved in wobble uridine modifications, which are very similar among eukaryotes and also have similar counterparts in bacteria and archaea.13
Nucleotide modifications necessary for translation to be possible
Each GC base pair in codon-anticodon interactions is held together by three hydrogen bonds (G≡C), whereas AU base pairs use only two (A=T or A=U). This leads to a total of 6 to 9 H-bonds per codon. The difference in free energy due to number of H-bonds or their spatial orientation alone is not enough to ensure sufficient discrimination between correct and wrong codon identification. Cells do not rely on only these H-bond interactions to hold tRNA and mRNA together but rely also on van der Waals forces, steric complementarity, and shape acceptance features.20
As an analogy, the need for exact recognition also exists in mRNA regulation by miRNAs (microRNAs). miRNA:mRNA seed regions use 6–8 perfect base-pair matches (unlike only three for codon:anticodon interactions) but even here additional machinery like the RISC complex (RNA-induced silencing complex) is used to ensure reliable interactions. Pairing of 8 perfect matches are indeed more likely to be bona fide miRNA targets. Nevertheless, in many cases even perfect 8-nt seed pairing between miRNA:mRNA does not down-regulate the bound mRNA.21 In a 2019 study, almost twice as many mRNA transcripts having a suitable seed sequence were shown not to be true targets.22 Presumably those interactions could be transiently recognized but then dissociate.
Dozens of features within the 21 to 23 nt-long miRNAs contribute to predict true miRNAs, such as: extreme similarity of 8-nt seed for the same gene across many organisms, GC content of target site, UTR length, free energy of seed binding, and distance to UTR end.22 Clearly there is much more involved in preparing the target for correct recognition than only possessing the seed sequence. The same principle holds also for tRNA anticodon to mRNA codon recognition.
Modification of tRNAs is so important that more genes are devoted to tRNA modification pathways than to the expression of tRNAs themselves. In E. coli, for example, 54 of the 61 sense codons are modified taking only nucleotide positions 34 or 37 into account.22
Modification of anticodon nucleotides to enhance codon–anticodon interactions
Table 1. Revised wobble pairing rules.23 Watson-Crick base pairs are shown in italics. Parentheses denote less favoured bindings.


The first two nucleotides of mRNA codons are most important in the genetic code. The code redundancy occurs at tRNA position 34, the ‘wobble position’, where often different nucleotides represent the same amino acid. This had led to the so-called revised wobble pairing rules (table 1).22,23
Nucleotide 34 undergoes modifications typically required for accurate translation, to permit identification by alternative anticodons. Two examples include modification cmo5U34 in tRNAAla(CGU) and mnm5s2U34 tRNALys(UUU).23 (There are special naming conventions for isoacceptors).24
Another well-studied example of tRNA modification affecting decoding is the deamination of adenosine (A) to inosine (I) at wobble position 34. Since A only base pairs with U, but I base pairs with U, C, and A, tRNAs with I at the wobble position have an extended codon–anticodon interaction capability (figure 3).7,25,16
Given the limited variety of isodecoders, the necessary genes to modify tRNAs at the wobble position were needed from the very beginning. Otherwise translation would have stalled at positions which could not be translated or incorporated the wrong amino acids in the new protein.
Modifications in the anticodon arm to enhance codon–anticodon interactions
Nucleotide 37, the residue immediately preceding the first anticodon nucleotide, is modified in >70% of all tRNAs. For example, the m1G37 modification is present in ~95% of all known sequences of proline tRNAs.10,23 Other examples include a 6-threonylcarbamoyladenosine (t6A) or a 2-methylthio derivative (ms2t6A), necessary to recognize AAA or AAG codons. The t6A and ms2t6A modifications help stabilize codon-anticodon pairing.26,23 Due to the weakness of the A-U hydrogen-bond base, pairings of tRNALys(UUU) with the AAA codon require modifications at both nucleotides 34 (mnm5s2) and 37 (t6A).23
Modifications to prevent frameshifting
Other properties can also be affected through modifications outside the anticodon loop, such as stability, ribosome binding, and various translational aspects, for example to prevent frameshifting.10
On the other hand, modifications in the anticodon loop can also affect the reading frames during translation. Mutations of the genes responsible for (yW) modification of tRNAPhe at position 37 cause increases in -1 frameshifting, useful to compensate for former translation errors.7
Modification to ensure tRNAs fold correctly
Modifications between the acceptor and D-stem help fold tRNAs into their correct structures and also increase tRNA rigidity.10 Several modifications hinder canonical base pairing and may facilitate folding into the typical L-shaped tRNA structure by preventing alternative nucleotide pairings or by producing single-stranded regions.20
Modifications to facilitate tRNA processing on the ribosome
Modifications between the D and anticodon stems increases the stiffness of the modified tRNA, useful to help tRNA on the ribosome adjust to conformational needs on the ribosome during translational elongation.10
The well-studied m1A58 modification on the T loop, present in almost all human tRNAs, is essential for the stability of the initiator tRNAMet. Adjusting the level of this single modification in specific tRNAs can serve as a rheostat to fine-tune translation rates.10
Modifications to permit codon recognition at the P site
Experiments have shown that anticodons including Lys(UUU), Glu(UUC), Gln(UUG), Arg(UCU), and Ala(UGC) in unmodified anticodon stem-loops (ASLs) did not recognize their cognate codons in the ribosomal P site (figure 4) in humans. With the incorporation of s2U34 into the unmodified ASLLys(UUU), the corresponding codons AAA and AAG codons were recognized but not AAC or AAU.27 Clearly additional factors are needed.
Modifications to permit codon recognition at the A site
ASLLys(UUU) constructs modified with s2U34, mnm5U34, or t6A37 combined with codon bound AAA individually and with mnm5U34 and t6A37 when present jointly. In addition, ASLLys(UUU)-s2U34 and ASLLys(UUU)-mnm5U34 t6A37 recognized the AAG codons in the ribosomal A site. Therefore, modifications that restored P site ribosomal codon recognition also restore codon recognition to the ribosomal A site.28 These examples further suggest upfront design. The 3-dimensional geometry of the A and P sites must be compatible with the sizes and shapes of all tRNAs, taking also chemical modifications into account.
Modifications so each type of tRNAMet interacts with correct elongator factor
Modifications serve to discriminate tRNAMet species that function in either the initiation or the elongation steps of translation. The initiator and elongator tRNAMet, referred to as tRNAiMet and tRNAeMet respectively, have different primary sequences and structures. Using mutagenesis, researchers uncovered that three G-C base pairs in the acceptor stem of the initiator tRNAMet are required for proper positioning of the mRNA in the correct reading frame.28
Despite these differences, both are aminoacylated by a single methionyl tRNA synthetase, Mes1. They are discriminated during translation via their interactions with translation factors. tRNAiMet interacts with elongator factor 2 (eIF2) and tRNAeMet with elongator factor 1 (eEF1α).7
Modifications to compensate for mutations leading to stop codons
A suppressor mutation, also called synthetic rescue, is a second mutation that restores the mRNA reading frameshift caused by a preceding shift. This works as long as the portion between the two mutations is not critical for protein function. As an example, frameshift-suppressor variants of tRNAPro contain an extra nucleotide in the anticodon loop between those in 37 and 38 (referred to as 37.5), and these tRNAs decode CCC-N codons as proline.23 In the context of tRNASufA6, the inserted G37.5 displaces A38 preventing a U32•A38 pairing. Binding of ASLSufA6 to a cognate CCG or a +1 slippery CCC-U codon is thereby extremely weak.23
Both tRNAPro(CGG) and tRNAPro(GGG) isoacceptors lacking the m1G37 modification undergo +1 frameshifting on CCC-N codons. Otherwise, M1 methylation at position 37 stabilizes the interaction with the codon and maintains the mRNA reading frame.23
Likewise, the absence of pseudouridine (ψ) at tRNATyr position 35 or m5C34 in tRNALeuCAA causes defects in tRNAmediated nonsense suppression.29 Defects of Mod5, which catalyzes isopentylation (i6A37) of a subset of tRNAs also causes a decrease in nonsense suppression of UAA.7
Modifications to ensure tRNA stability
Methylation of m1A58 of tRNAiMet, via Trm6/Trm61 catalysis, is essential for that tRNA’s stability.7
The nucleotide identity of the 32•38 pairing in tRNAs has been found to be predictable on the basis of the strength of the codon-anticodon interaction. For example, the anticodon of E. coli tRNAAla(GGC) is stronger due to the three G•C pairs which provide 3 hydrogen-bond interactions per nucleotide between the codon and the anticodon. In this ‘strong’ case, the 32•38 pairing in tRNA needs to be correspondingly weak. Changing tRNAAla(GGC) from a weak U32•A38 pair to a strong pair such as C32•A38, prevents the ribosome from being able to distinguish correct from incorrect tRNA-mRNA pairs.23
Modifications to produce synonymous codons
Table 2. Universal code, showing codon to amino acid mapping. The same amino acid can be coded for by 1, 2, 3, 4, or 6 different codons. The stop signal is represented by 3 codons.

In the universal genetic code there are five examples of double redundancy, see table 2: Phe vs Leu; His vs Gln; Asn vs Lys; Asp vs Glu; and Ser vs Arg. Looking more closely, we observe that the first amino acid in each pair is specified by a U or C, and the second of each pair by A or G. Experiments described below show that a single design concept was implemented to ensure this pattern of codon with anticodon recognition.28
A key series of experiments in the early 2000s shed light on how a single nucleotide in the wobble position could provide enough binding discrimination. Nucleotide modifications on anticodon position U34 make this possible, specifically chemical modifications s2U34, mnm5U34, and t6A37.27
Modifications to prevent misinterpretation of start and stop codons
Modified methionine is used to identify the start position on mRNAs, and is encoded by AUG. This triplet differs by only one nucleotide at the weak ‘wobble’ position from isoleucine (encoded by AUA, AUU, and AUC). To ensure accurate decoding, in bacteria the isoleucine AUA codon requires a k2C modification at position 34. Conversely, E. coli elongator tRNAMet has an ac4C modification at the same position to prevent misreading of the AUA isoleucine codon.30
Similarly, modification of position 34 with Cm or Q enable tRNATrp or tRNATyr, respectively, to discriminate tryptophan (UGG) or tyrosine codons (UAU and UAC) from stop codons during translation.31
Modifications are responsive to environmental changes
Many tRNA modifications are performed dynamically to enhance survival to environmental changes during the organism’s lifetime. For example, several thermophilic bacteria show an increase in tRNA modification levels with growth temperature.31 In yeast, the tRNA modifications resulting upon exposure to different toxins are considerable and reproducible, revealing an integration with other pathways. In response to oxidative stress induced by hydrogen peroxide, the C34 position of yeast tRNALeu(CAA) is modified by tRNA methyltransferase 4 (Trm4). Furthermore, the amount of m5C modification on tRNAHis rises in response to nutrient depletion and other growth arrest conditions. Another yeast tRNA methyltranferase, Trm9, completes the formation of mcm5U and mcm5s2U at the wobble position U34 of tRNAArg(UCU) and tRNAGlu(UUC); this methylation prevents cell death by promoting translation of DNA damage response genes that are enriched with arginine and glutamic acid codons.16
DNMT2-mediated methylation protects tRNA against stress-induced fragmentation, which is beneficial because tRNA fragments can inhibit the activity of the small RNA processing enzyme Dicer-2 and cause dysfunction of RNA interference.16 This topic will be followed up on in part 431 of this series.
Specialized protein machines known as writers, erasers, and readers of m6A are well known for mRNAs. The writer proteins selectively attach ligands, in particular methyl groups. Demethylases balance the methylation stoichiometry of specific mRNAs, often in a pathway- and cell-type-specific manner. The reader proteins execute the biologically intended activities on specific transcripts.16
tRNA concentration used as a regulatory signal
In E. coli, accumulation of uncharged tRNAs at the ribosome A site due to amino acid limitation activates a protein called RelA. This RelA synthesizes pppGpp (5’-triphosphate3’- diphosphateguanosine), which initiates a complex process to decrease global gene transcription, while simultaneously stimulating a few genes related to the synthesis of amino acids. Limitation of amino acid availability thus leads to lower levels of rRNA and tRNA synthesis which are not needed by the cell at the time.3
In fact, accumulation of uncharged tRNAs serves as an input signal for several pathways that regulate gene expression levels, helping the organism to survive under adverse conditions.3 In eukaryotes, uncharged tRNAs present due to amino acid deficiency interact with a protein kinase called Gcn2p which has a reduced affinity for the charged form of a tRNA.3,15
tRNAArg mediates amino-terminal arginylation as a tag for protein degradation. In prokaryotes leucine is used instead of arginine for tRNA-dependent N-terminal modifications.15
Various Gram-positive bacteria regulate expression of synthetase genes and genes involved in amino acid biosynthes by using uncharged tRNAs as an information signal, an example of cellular autoregulation. The Tbox control system involved is integrated with a 200–300 nucleotide regulatory sequence in the 5’ untranslated regions of the regulated genes.3
Conclusions
Attachments of specific ligands to tRNAs ensure the proper three-dimensional shape is produced, with the necessary rigidity, ability to interact with several ribosomal components, and with many other cellular processes. In many cases the enzymes involved are very complex. Of vital importance, errors must be prevented in decoding the start and stop codons, which differ by only one nucleotide from coding triplets. The necessary discrimination is provided by modifying key nucleotides using protein-based enzymes31 which could not have existed before they were needed by the tRNAs. However, a fundamental principle in biology is that small, rapidly reproducing organisms have a strong drive to streamline their genomes.32,33 But we see that evolution would require a DNA replicator ‘organism’ preceding the genetic code to possess large amounts of superfluous DNA to be used as the genes to process the tRNAs. Natural selection, however, will favour those replications freed of unnecessary genetic ballast.
The exciting challenge in creation science lies in trying to understand the top down planning and design logic used in cells. This leads to reflection on the tradeoffs involved in alternative solutions, confident that brilliant engineering was involved. The evolutionist can only claim ‘these things just happened’ and allege some evolutionary path must have been followed. This kind of vague thinking offers no research guidance, but the opposite helps develop the kinds of logical thinking so important in developing modern technologies, and cures for diseases. In other words, design analytical skills.
Since evolution lacks planning and foresight the naturalist must invoke what we will call the Insanely Improbable Chance Change (IICC) principle. Some biochemical change happened for the first time with an immediate effect of sufficient benefit for natural selection to act on. For example, at some point for the first time an enzyme catalyzed addition of a second CCA to one or more RNAs, and by miraculous good fortune these just happened to be flawed tRNAs which a cell would be better off without, and astonishingly an unrelated enzyme just happened to recognize the CCACCA pattern and degraded it. These kinds of IICC physical changes must have occurred before any natural selection could begin to act, but the chances of the molecular machines being in place and providing the opportunistic starting point are vanishingly small, all the while avoiding deleterious alternatives. We see this evolutionary assumption every time a code or informative signal is used, for example the N-End Rule to specify half-life of proteins.34
References and notes
- Truman, R., The Surprisingly Complex tRNA Subsystem: part 1—generation and maturation, J. Creation 34(3):80–86, 2020. Return to text.
- Hopper, A.K., Pai, D.A., and Engelke, D.R., Cellular dynamics of tRNAs and their genes, FEBS Letters 584:310–317, 2010. Return to text.
- Raina, M. and Ibba, M., tRNAs as regulators of biological processes, Front Genet. 5:171, 2014. Return to text.
- Reiner, R., Ben-Asouli, Y., Krilovetzky, I., and Jarrous, N., A role for the catalytic ribonucleoprotein RNase P in RNA polymerase III transcription, Genes Dev. 20(12):1621–1635, 2006. Return to text.
- Yu, N., Jora, M., Solivio, B., Thakur, P., Acevedo-Rocha, C.G., Randau, L., de Crécy-Lagard, V., Addepalli, B., and Limbach, P.A., tRNA modification profiles and codon-decoding strategies in Methanocaldococcus jannaschii, J. Bacteriology 201(9) e00690-18, April 2019. Return to text.
- Machnicka, M.A. et al., MODOMICS: a database of RNA modification pathways–2013 update, Nucleic Acids Res. 41:D262–D267, 2013. Return to text.
- Hopper, A. K., Transfer RNA post-transcriptional processing, turnover, and subcellular dynamics in the yeast Saccharomyces cerevisiae, Genetics 194:43–67, 2013. Return to text.
- Fields, R.N. and Roy, H., Deciphering the tRNA-dependent lipid aminoacylation systems in bacteria: novel components and structural advances, RNA Biol. 15(4–5):480–491, 2017. Return to text.
- Castillo, A., Tello, M., Ringwald, K. et al., A DNA segment encoding the anticodon stem/loop of tRNA determines the specific recombination of integrative-conjugative elements in Acidithiobacillus species, RNA Biol. 15(4–5):492–499, 2017. Return to text.
- Pan, T., Modifications and functional genomics of human transfer RNA, Cell. Res. 28:395–404, 2018. Return to text.
- Huber, S.M., Leonardi, A., Dedon, P.C., and Begley, T.J., The versatile roles of the tRNA epitranscriptome during cellular responses to toxic exposures and environmental stress, Toxics. 7(1):17, 2019. Return to text.
- Deutsch, C., El Yacoubi, B., de Crécy-Lagard, V., and Iwata-Reuyl, D., Biosynthesis of threonylcarbamoyl adenosine (t6 A), a universal tRNA nucleoside, J. Biological Chemistry 287(17):13666–13673, 2012. Return to text.
- Hopper, A.K. and Nostramo, R.T., tRNA processing and subcellular trafficking proteins multitask in pathways for other RNAs, Front. Genet., 20 February 2019 | doi:10.3389/fgene.2019.00096. Return to text.
- Barciszewska, M. Z., Perrigue, P. M., and Barciszewski J., tRNA—the golden standard in molecular biology, Mol. BioSyst. 12(12):12–17, 2016. Return to text.
- Wang, X. and He, C., Dynamic RNA modifications in posttranscriptional regulation, Mol. Cell 56:5–12, 2014. Return to text.
- Abbott, J.A., Francklyn, C.S., and Robey-Bond, S.M., Transfer RNA and human disease, Front. Genet. 5(158):1–18, 2014. Return to text.
- Hopper, A.K. and Phizicky, E.M., tRNA transfers to the limelight, Genes Dev. 17:162–180, 2003. Return to text.
- Deutscher, M.P., Processing of tRNA in prokaryotes and eukaryote, Critical Reviews in Biochemistry 17:1:45–71, 1984. Return to text.
- Sokolowski, M., Klassenc, R., Bruch, A., Schaffrath, R., and Glatt, S., Cooperativity between different tRNA modifications and their modification pathways, Biochim Biophys Acta 1861:409–418, 2018. Return to text.
- Ou, X., Cao, J., Cheng, A., Peppelenbosch, M.P., and Pan, Q., Errors in translational decoding: tRNA wobbling or misincorporation? PLoS Genet 15(3):e1008017, 2019. Return to text.
- Liu, W. and Wang, X., Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data, Genome Biology 20:18, 2019. Return to text.
- Nguyen, H.A., Hoffer, E.D., and Dunham, C.M., Importance of a tRNA anticodon loop modification and a conserved, noncanonical anticodon stem pairing in tRNACGGPro for decoding, J. Biol. Chem. 294:5281–5291, 2019. Return to text.
- Agris, P.F., Decoding the genome: a modified view, Nucleic Acids Research 32(1):223–238, 2004. Return to text.
- Naming convention for isoacceptors (tRNAs having different anticodons but charged with the same amino acid). Consider Methionine (mRNA codon UAC). Codons in mRNA are read in the 5’→ 3’ direction, anticodons are oriented in the 3’→ 5’ direction. The charged molecule would be reported as tRNAMet(CAU). Return to text.
- O’Donoghue, P., Ling, J., and Söll, D., Transfer RNA function and evolution, RNA Biology 15:(4–5):423–426, 2018. Return to text.
- Agris, P.F., Eruysal, E.R., Narendran, A., Väre, V.Y. P., Vangaveti, S., and Ranganathan, S.V., Celebrating wobble decoding: half a century and still much is new, RNA Biology 15(4–5):537–553, 2018. Return to text.
- Yarian, C., Townsend, H., Czestkowski, W., Sochacka, E., Malkiewicz, A.J., Guenther, R., Miskiewicz, A., and Agris, P.F., Accurate translation of the genetic code depends on tRNA modified nucleosides, J. Biological Chemistry 277(19):16391–16395, 2002. Return to text.
- Roy, B., Liu, Q., Shoji, S. et al., IF2 and unique features of initiator tRNA(fMet) help establish the translational reading frame, RNA Biol. 15(4–5):604–613, 2017. Return to text.
- Mutations or misreading during translation can produce a stop codon. A nonsense suppressor mutated tRNA can suppress this error by binding with a termination codon on the mRNA. Of the three stop codons, cells often use ochre codons (UAA) as the termination signal to a much higher degree, for which nonsense suppressors usually interact less effectively. Return to text.
- Shepherd, J. and Ibba, M., Bacterial transfer RNAs, FEMS Microbiology Reviews 39(3):280–300, 2015. Return to text.
- Truman, R., The surprisingly complex tRNA subsystem: part 4—tRNA fragments regulate processes, J. Creation, submitted 2020. Return to text.
- Truman, R. and Borger, P., Genome truncation vs mutational opportunity: can new genes arise via gene duplication?—part 1, J. Creation 22(1):99–110, 2008. Return to text.
- Truman, R. and Borger, P., Genome truncation vs mutational opportunity: can new genes arise via gene duplication?—part 2, J. Creation 22(1):111–119, 2008. Return to text.
- Truman, R., The ubiquitin protein: chance or design? J Creation 19(3):116–127, 2005. Return to text.
- Perche-Letuvée, P, Molle, T., Forouhar, F., Mulliez, E., and Atta, M., Wybutosine biosynthesis: structural and mechanistic overview, RNA Biology 11(12):1508–1518, 2014. Return to text.







Readers’ comments
Comments are automatically closed 14 days after publication.