Feedback archive → Feedback 2022
Can nucleobases and self-replication arise from non-living chemicals?
More overstated chemical evolution claims

Life arising from non-living chemicals has proven so intractable that many evolutionists dodge the issue, claiming that it is not really part of evolution but is instead ‘abiogenesis’. However, the General Theory of Evolution “is the theory that all the living forms in the world have arisen from a single source which itself came from an inorganic form.” Furthermore, a common academic term for abiogenesis is chemical evolution. And regardless of terminology, atheists must believe it.
It is thus no accident that papers purporting to support chemical evolution keep coming. This week’s feedback comes from Stephen R. of the USA, with a reply from Ph.D. chemist Jonathan Sarfati
[This paper1] talks about forming nucleobases in a Miller–Urey type atmosphere along with the amino acids.I have never seen anything on nucleobases forming with this atmosphere or experiment type before and then it talks about how our atmosphere switched from reducing to oxidizing.

Dear Mr R.
Thank you for writing to CMI.
This paper1 doesn’t seem to have made the rounds for good reason. It argues for a reducing atmosphere, and at best produces parts-per-million of nucleobases, which decompose readily under those same conditions. Later that year, another paper in the same journal obliquely admitted problems with losses of nucleobases:
[T]he rapid losses of nucleobases to pond seepage during wet periods, and to UV photodissociation during dry periods, mean that the synthesis of nucleotides and their polymerization into RNA occurred in just one to a few wet–dry cycles.2
This is very optimistic spin. Note:
- Fact: the nucleobases will either seep out or decompose under UV radiation.
- Dogma: RNA must have formed naturally—well, we are here, aren’t we? (Don’t you dare mention creation as a possibility, because it’s not science even if all the evidence supports it!)
- Therefore, the RNA must have formed very rapidly before the nucleotides were lost or destroyed.3
Of course, this is begging the question: assuming what needs to be proved. As for the transition from an alleged early reducing atmosphere, the data points look to be all over the place. But again, the paper2 is dogma-driven not evidence-driven:
As to the sources of nucleobases, early Earth’s atmosphere was likely dominated by CO2, N2, SO2, and H2O. In such a weakly reducing atmosphere, Miller–Urey-type reactions are not very efficient at producing organics. One solution is that the nucleobases were delivered by interplanetary dust particles (IDPs) and meteorites.
Translation: Earth’s early atmosphere was not very good at producing biomolecules. But we know biomolecules formed naturally, because we are here, so they probably came from outer space.
Earlier, I had discussed some revisions of Miller–Urey, including the atmosphere.
Finally, a colleague wrote a helpful article about handling chemical evolutionary claims in general: Reading ‘origin of life’ research: How to read the secular literature on chemical evolution (i.e. ‘abiogenesis’) critically.
Later, Stephen R. wrote another email about chemical evolutionary claims:
This [paper4] is a fresh new story on supposedly creating replicating RNA.To me it looks like they’re using existing RNA and not making it from scratch but still wouldn’t RNA hydrolyze in water? I would expect it to break apart. Also, it’s interesting that mutations would occur or that it could replicate like this.Could you help me out on this one?

I had also seen articles like that on the same experiments, and answered several other queries about it. The article you cited excitedly claims:
Researchers at the University of Tokyo have for the first time been able to create an RNA molecule that replicates, diversifies, and develops complexity, following Darwinian evolution.
This article cites lead researcher Ryo Mizuuchi:
The simplicity of our molecular replication system, compared with biological organisms, allows us to examine evolutionary phenomena with unprecedented resolution. The evolution of complexity seen in our experiment is just the beginning. Many more events should occur towards the emergence of living systems.
What struck me, just as it did you, was how they had to presuppose so much chemical evolution to get even this far: RNA molecules and protein enzymes. The original paper5 partly gives the game away in the title, including “from a single RNA replicator”.
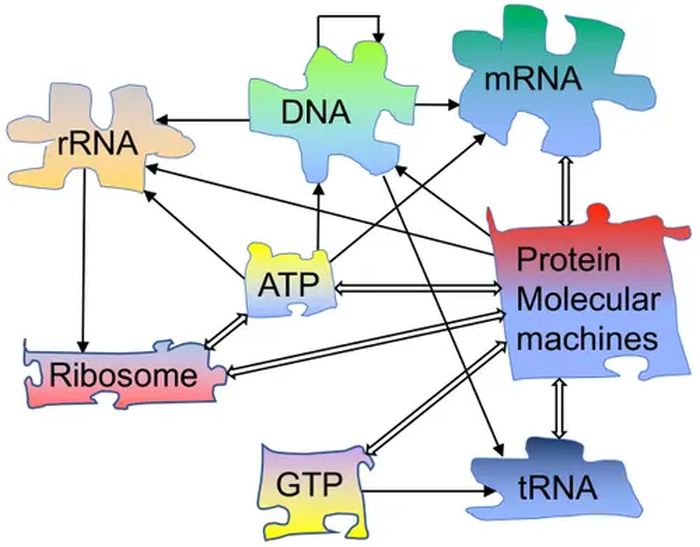
The scientists started with an RNA strand 2125 nucleotides long from a Qb virus. There were also a ribosome, an extremely complex machine, many precisely shaped enzymes, and complex and 46 coordinated t-RNA molecules. The RNA strand codes for Qb replicase, an enzyme that makes new strands of complementary RNA on an RNA strand. So although Dr Mizuuchi refers to “The simplicity of our molecular replication system, compared with biological organisms”, the system is still frightfully complex.
After hundreds of mutation/artificial-selection cycles, many variants were disabled as far as coding for replicase. But some RNA variant strands dominated because they encoded replicases that preferentially transcribed those variants.
But neither the original nor any of the variants could be remotely described as self-replicating. They could neither reproduce nor produce enzymes without the supplied complex machinery. Furthermore, although it is common to refer to such scenarios as ‘directed evolution’, ‘directed selection’ is more accurate.
Furthermore, any real origin-of-life scenario must explain how these homochiral information-rich polymers could arise from a dilute and contaminated primordial soup. As per the previous reply, even the building blocks, the nucleobases, break down extremely quickly.
We now know that DNA is very unstable, but RNA breaks down 100× faster, so again you are right about hydrolysis. See DNA: the best information storage system and reply to Owen H. below the article.
The Intelligent Design website Evolutionnews.org rightly pointed out an alternative summary that fits the facts better:
Researchers further demonstrated the implausibility of life originating through undirected processes. Their experiment reinforces the conclusion that any form of molecular replication requires the highly sophisticated machinery that only exists in living cells. And the origin of any cellular component requires externally imparted information. The study also further discredits the claim that Darwinian evolution could have assisted life’s origin by showing that random mutations at best only slightly modify preexistent functions in proteins. Nothing novel ever emerges, and complexity never significantly increases.6
References and notes
- Ferus, M. and 7 others, Formation of nucleobases in a Miller–Urey reducing atmosphere, PNAS 114(17):4306–4311, 10 Apr 2017 | doi:10.1073/pnas.1700010114. Return to text.
- Pearce, B.K.D. and 3 others, Origin of the RNA world: The fate of nucleobases in warm little ponds, PNAS 114(43):11327–11332, 2 Oct 2017 | doi:10.1073/pnas.1710339114. Return to text.
- Compare Luskin, C., Warm little pond? PNAS paper admits difficulties generating RNA on prebiotic earth, evolutionnews.org, 3 Oct 2017. Return to text.
- University of Tokyo, New insight into possible origins of life: For the first time researchers create an RNA molecule that replicates, scitechdaily.com, 18 Mar 2022. Return to text.
- Mizuuchi, R. and 3 others, Evolutionary transition from a single RNA replicator to a multiple replicator network, Nature Communications 13:1460, 18 Mar 2022 | doi: 10.1038/s41467-022-29113-x. Return to text.
- Miller, B., Fact Check: Did University of Tokyo Researchers Explain the Origin of Life? evolutionnews.com, 22 Mar 2022. Return to text.








Readers’ comments
Comments are automatically closed 14 days after publication.